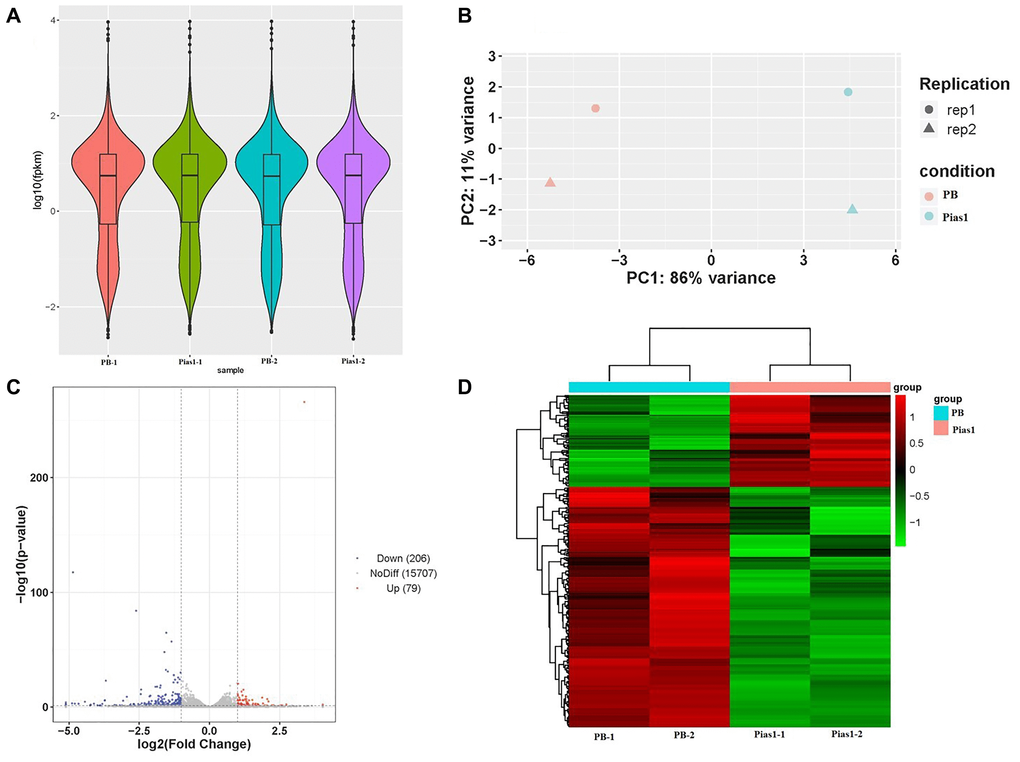
Figure 3.RNA-seq data quality and DEG analysis. (A) The violin plot based on values of log10 (fpkm) showed the whole gene expression distribution of four groups, including PB-1, PB-2, Pias1-1 and Pias1-2. (B) The PCA plot showed how well the four sequencing samples (PB-1, PB-2, Pias1-1 and Pias1-2) are separated from each other. (C) The scatter plot showed the results of DEGs based on the values of log2 (fold change) and -log10 (p value), including 206 downregulated genes, 79 upregulated genes and 15707 genes with no significant differences. (D) The heatmap showed the expression patterns of 285 significantly dysregulated genes with FC greater than 2 and p value less than 0.05, including 79 upregulated genes and 206 downregulated genes, by comparison of the Pias1/+ group to the WT group.