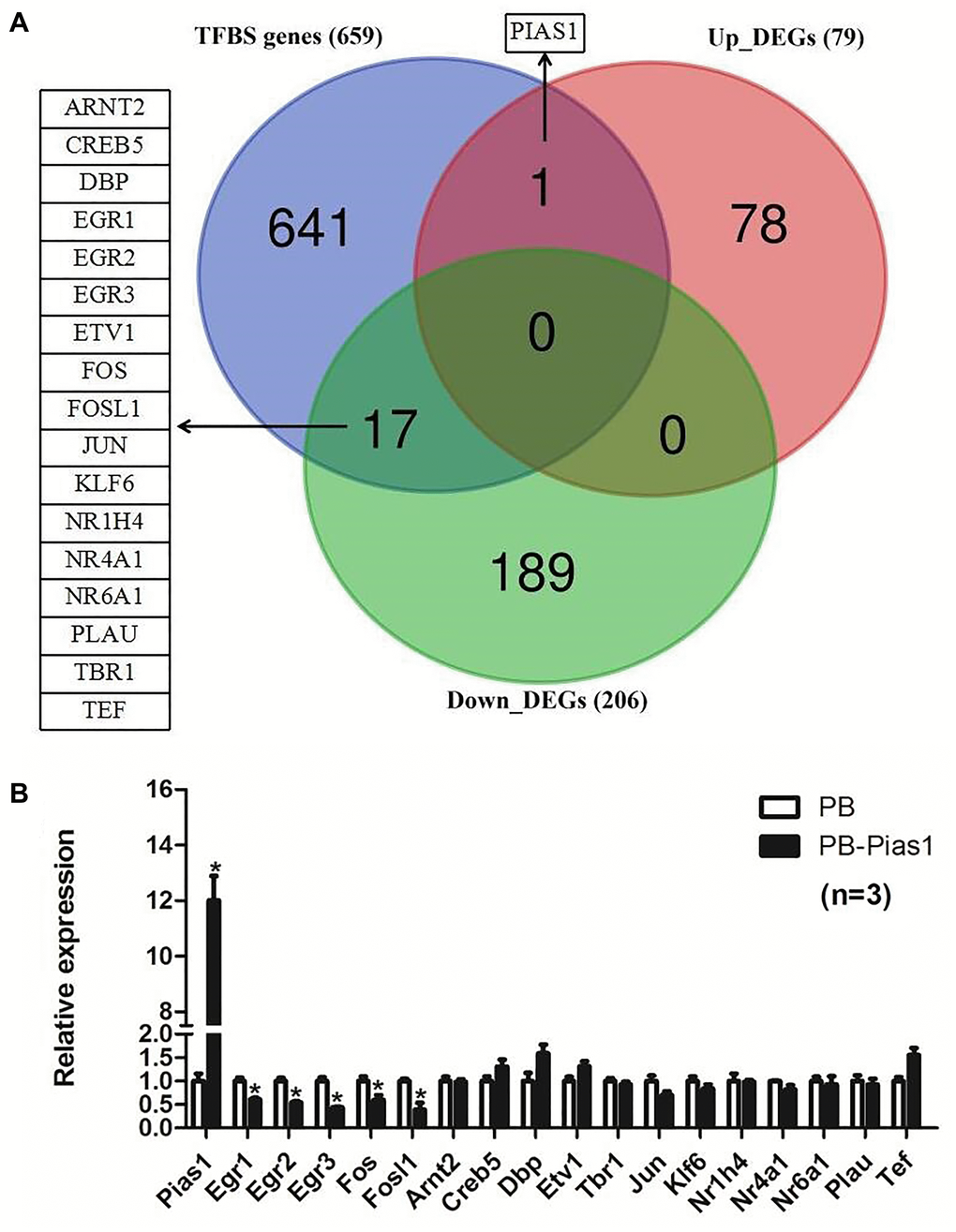
Figure 7.Overlapping targets between transcription factor binding site genes of STAT1 and DEGs of PIAS1. (A) Transcription factor binding site (TFBS) prediction for STAT1 was performed and compared with the differentially expressed genes of RNA-seq results in PIAS1 overexpression cell lines. As a result, 18 overlapping genes, including one upregulated gene PIAS1 and 17 downregulated genes, were identified as the target genes. (B) The regulation of the overlapping genes was evaluated by qPCR. As a result, five genes, including early growth response 1 (Egr1), early growth response 2 (Egr2), early growth response 3 (Egr3), FBJ osteosarcoma oncogene (Fos) and fos-like antigen 1 (Fosl1), were significantly downregulated in PIAS1-overexpressing cells.