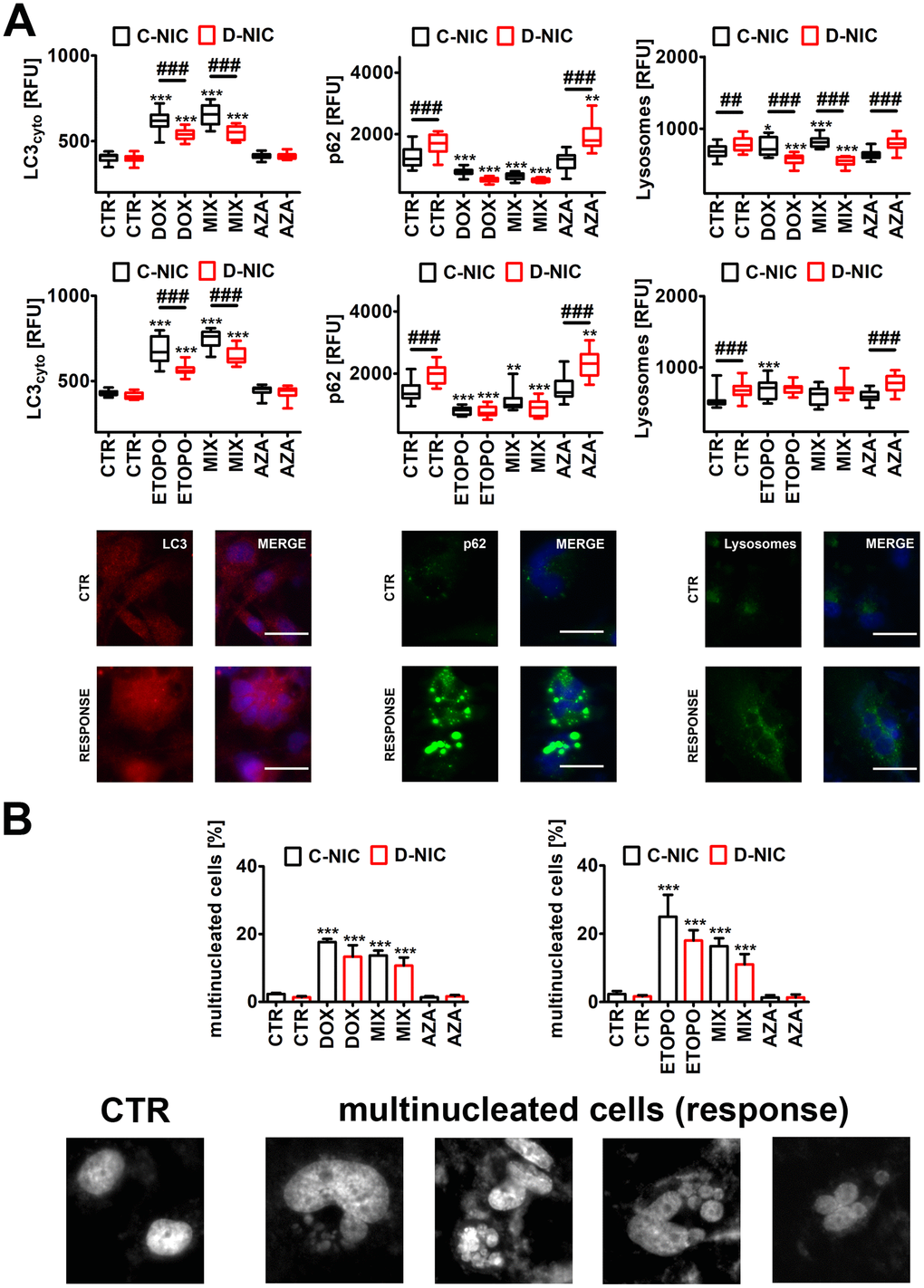
Figure 6.DNMT2/TRDMT1 gene knockout-mediated autophagy (A) and multinucleation (B) in U-251 MG glioblastoma cells treated with DOX or ETOPO. (A) LC3 immunostaining (red) and GFP-based imaging of a lysosomal marker Lamp1 and an autophagy marker p62 (green). The levels of LC3, p62 and Lamp1 are expressed as relative fluorescence units (RFU). Representative microphotographs are shown, objective 20x, nucleus staining (blue), RESPONSE, representative DOX or ETOPO treatment. Box and whisker plots are shown, n = 3, ***p < 0.001, **p < 0.01, *p < 0.05 compared to CTR (ANOVA and Dunnett’s a posteriori test), ###p < 0.001, ##p < 0.01 compared to C-NIC cells at the same culture conditions (ANOVA and Tukey’s a posteriori test). (B) Multinucleation events (%) were analyzed using Hoechst 33342 staining. Representative microphotographs are shown, objective 20x, RESPONSE, representative DOX or ETOPO treatment. Bars indicate SD, n = 3, ***p < 0.001 compared to CTR (ANOVA and Dunnett's a posteriori test). CTR, control conditions; DOX, doxorubicin treatment; ETOPO, etoposide treatment; AZA, azacytidine treatment; MIX, azacytidine post-treatment; C-NIC, control cells with unmodified levels of DNMT2/TRDMT1 containing control plasmid; D-NIC, cells with DNMT2/TRDMT1 gene knockout containing dedicated DNMT2 double nickase plasmid.