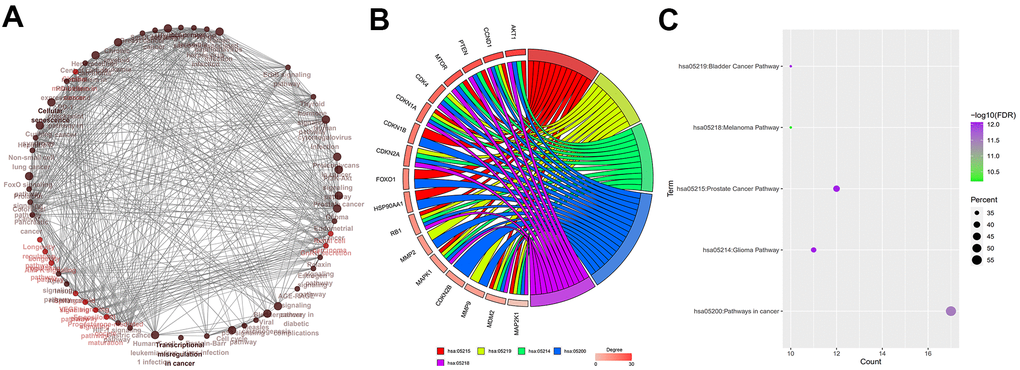
Figure 4.Identification of hub genes. (A) Relationship and interactions between baicalein-targeted genes in the top 5 enriched KEGG pathways. (B) Functional enrichment analysis results of baicalein-targeted genes. CCND1, CDKN1A, RB1, MAPK1, MDM2, and MAP2K1 were common to all the top five shared KEGG pathways and were designated as hub genes. The top five genes based on degree connectivity were AKT1, CCND1, PTEN, MTOR, and CDK4. (C) The FDR values, gene numbers, and rich factor values (ratio of the number of enriched DEGs in the KEGG pathway category compared to the total number of genes in that category) of the top five shared KEGG pathways.