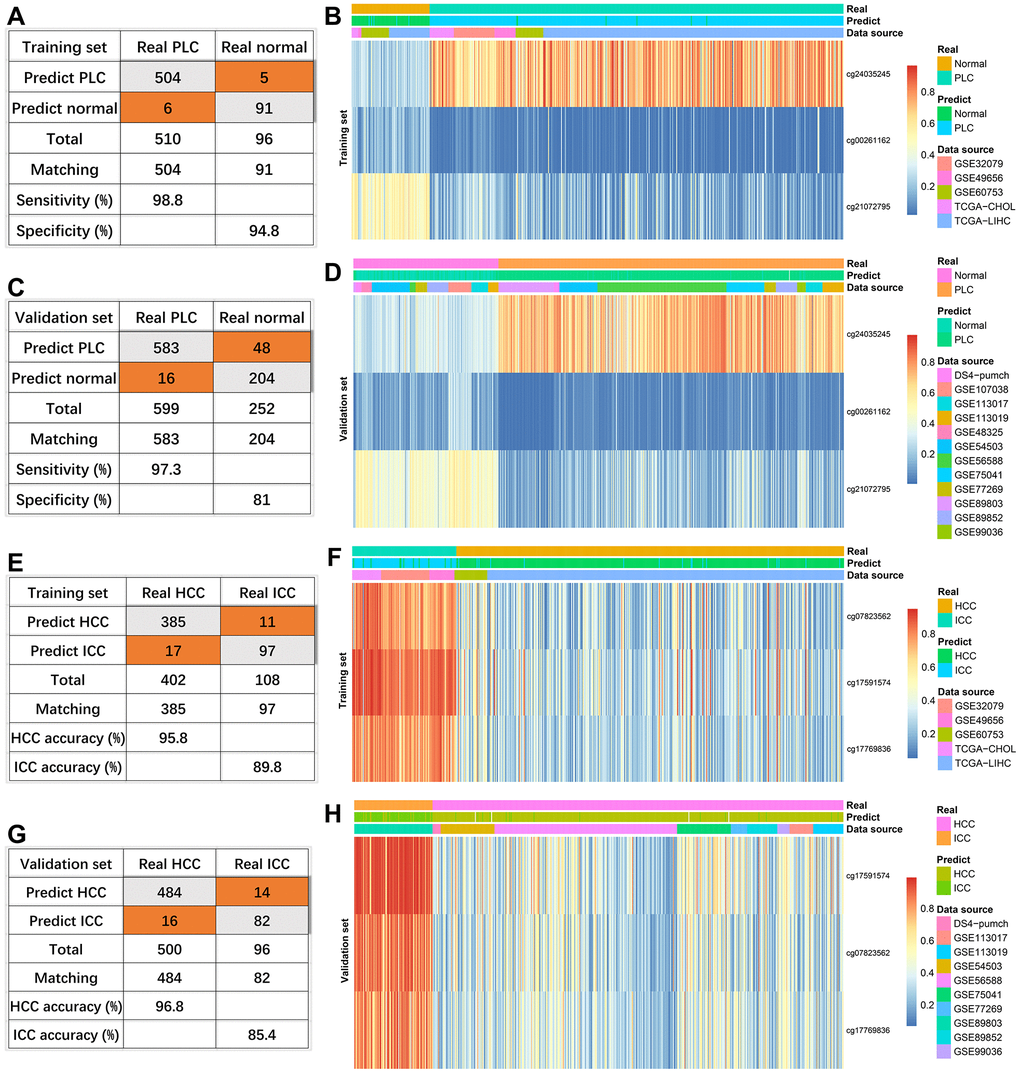
Figure 3.Effectiveness of diagnostic models. The diagnosis efficiency of the model for distinguishing between PLC and normal in the training set (A) and validation set (C). The heatmaps (PLC versus Normal) including real status, predict status, data source, and methylation values of indicated sites in the training set (B) and validation set (D). The diagnosis efficiency of the model for distinguishing between HCC and ICC in the training set (E) and validation set (G). The heatmaps (HCC versus ICC) including real status, predict status, data source, and methylation values of indicated sites in the training set (F) and validation set (H).