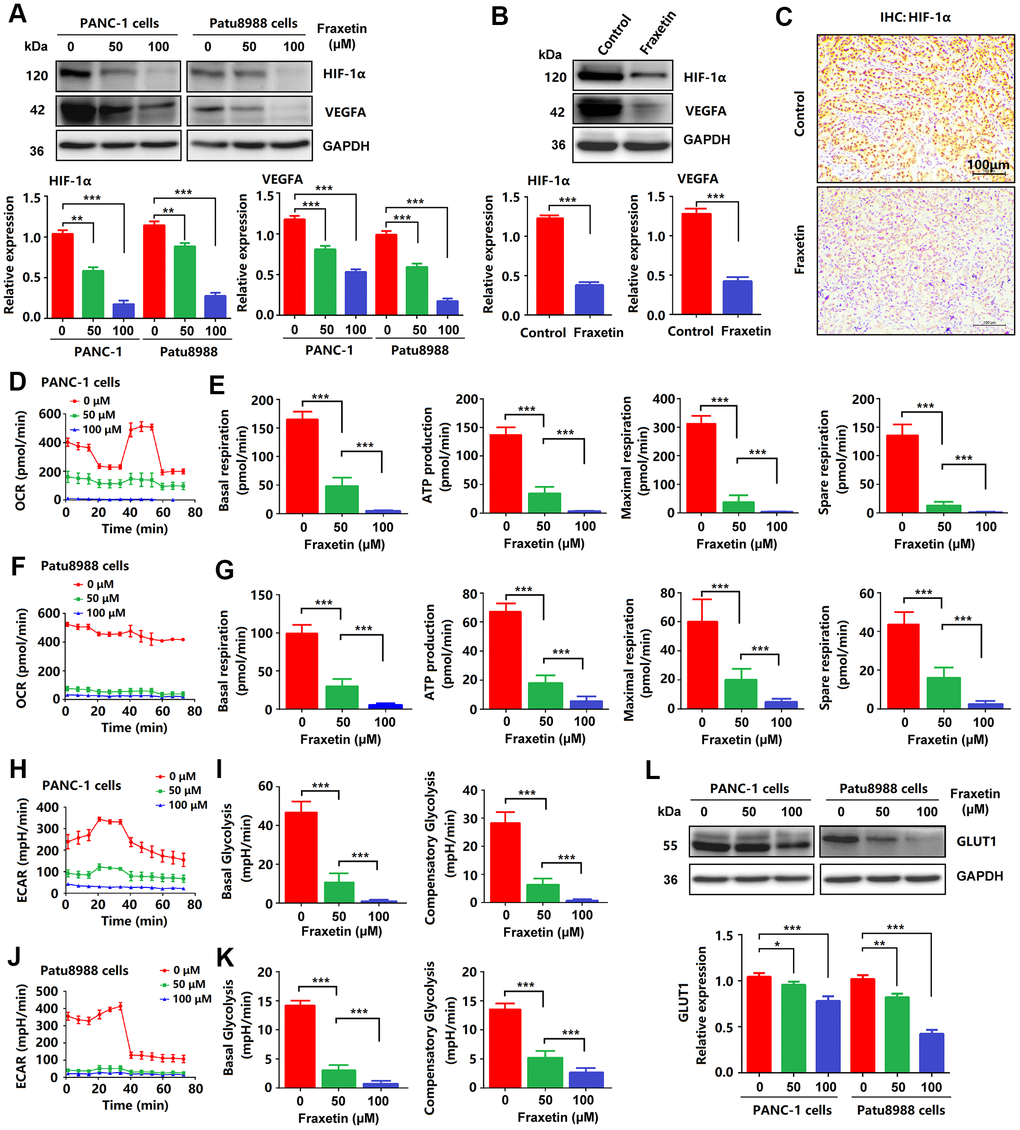
Figure 7.Fraxetin inhibited angiogenesis, glucose metabolism and EMT in PCCs. (A) HIF-1 and VEGFA expression in fraxetin-treated PANC-1 and Patu8988 cells as seen on a Western blot. (B) Western blot analysis of HIF-1 and VEGFA expression in fraxetin-treated animal xenograft models. (C) IHC staining for HIF-1α in fraxetin-treated models. Bar = 100 μm. (D–G) Glucose metabolism assay shows downregulated oxygen consumption rate (OCR), basal respiration, spare respiration, maximal respiration, and ATP production in fraxetin-treated PANC-1 and Patu8988 cells. (H–K) Glucose metabolism assay showing reduced levels of extracellular acidification rate (ECAR), basal glycolysis and compensatory glycolysis in fraxetin-treated PANC-1 and Patu8988 cells. (L) GLUT1 expression in fraxetin-treated PANC-1 and Patu8988 cells as seen on a Western blot. Data were presented as the mean ± standard deviation, and were analyzed by One-way ANOVA with Bonferroni’s post-hoc test and two-sided Student’s t-test. *P < 0.05; **P < 0.01, ***P < 0.001.