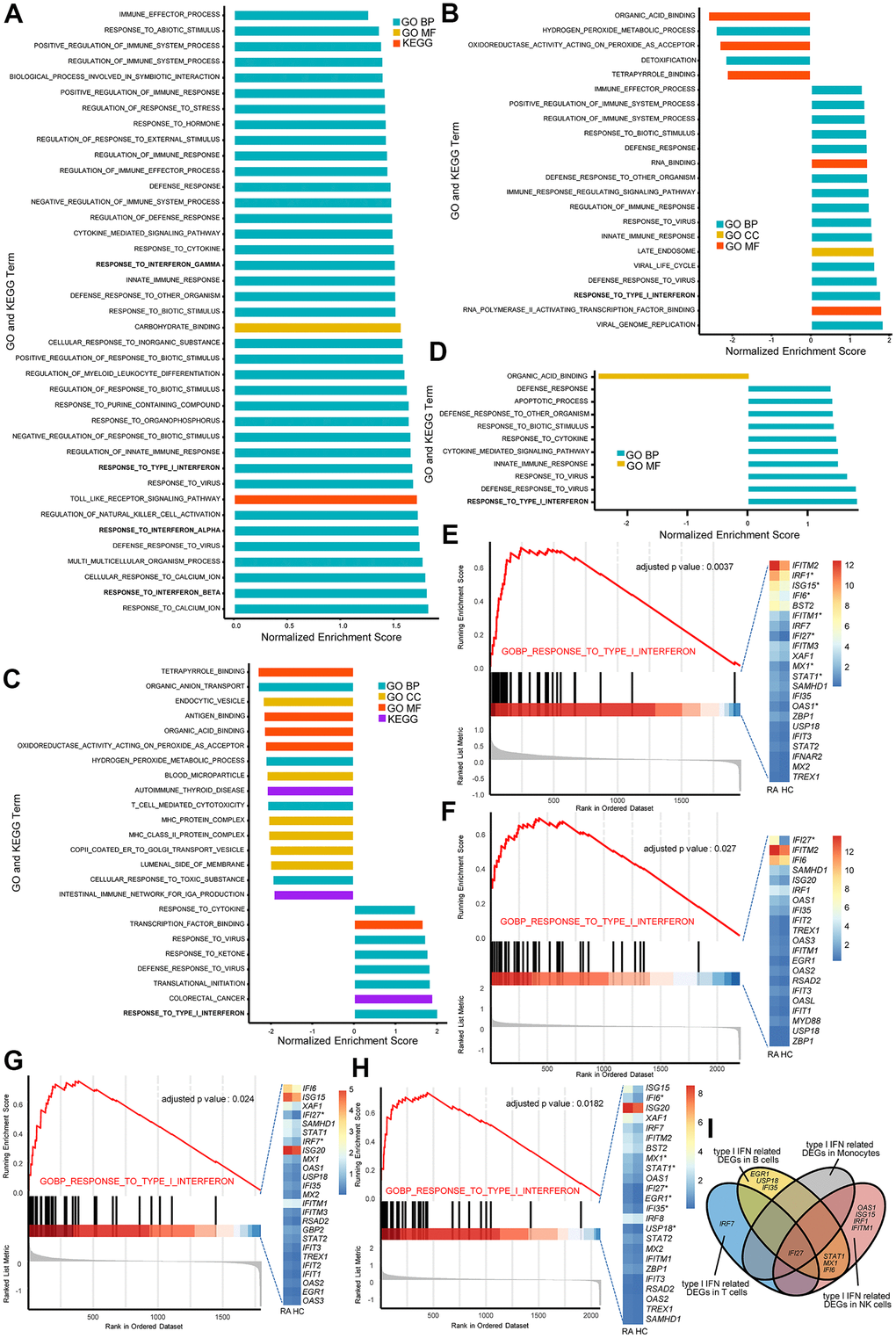
Figure 2.The type I interferon (IFN) signaling pathway is activated in natural killer (NK) cells, monocytes, and T and B cells in rheumatoid arthritis (RA). All GSEA bar plots of NK cells (A), monocytes (B), T cells (C) and B cells (D) detected by single-cell RNA (scRNA) sequencing data. GSEA plots and Het maps showing the activated type I IFN signaling pathway and genes in NK cells (E), monocytes (F), T cells (G) and B cells (H) detected by scRNA sequencing data. Upregulated genes in RA are marked with an asterisk (*). All upregulated genes satisfied log2 (fold change)>0.25 and adjusted p-value<0.05. (I) Venn diagram of upregulated type I IFN-stimulated genes in each immune cell type.