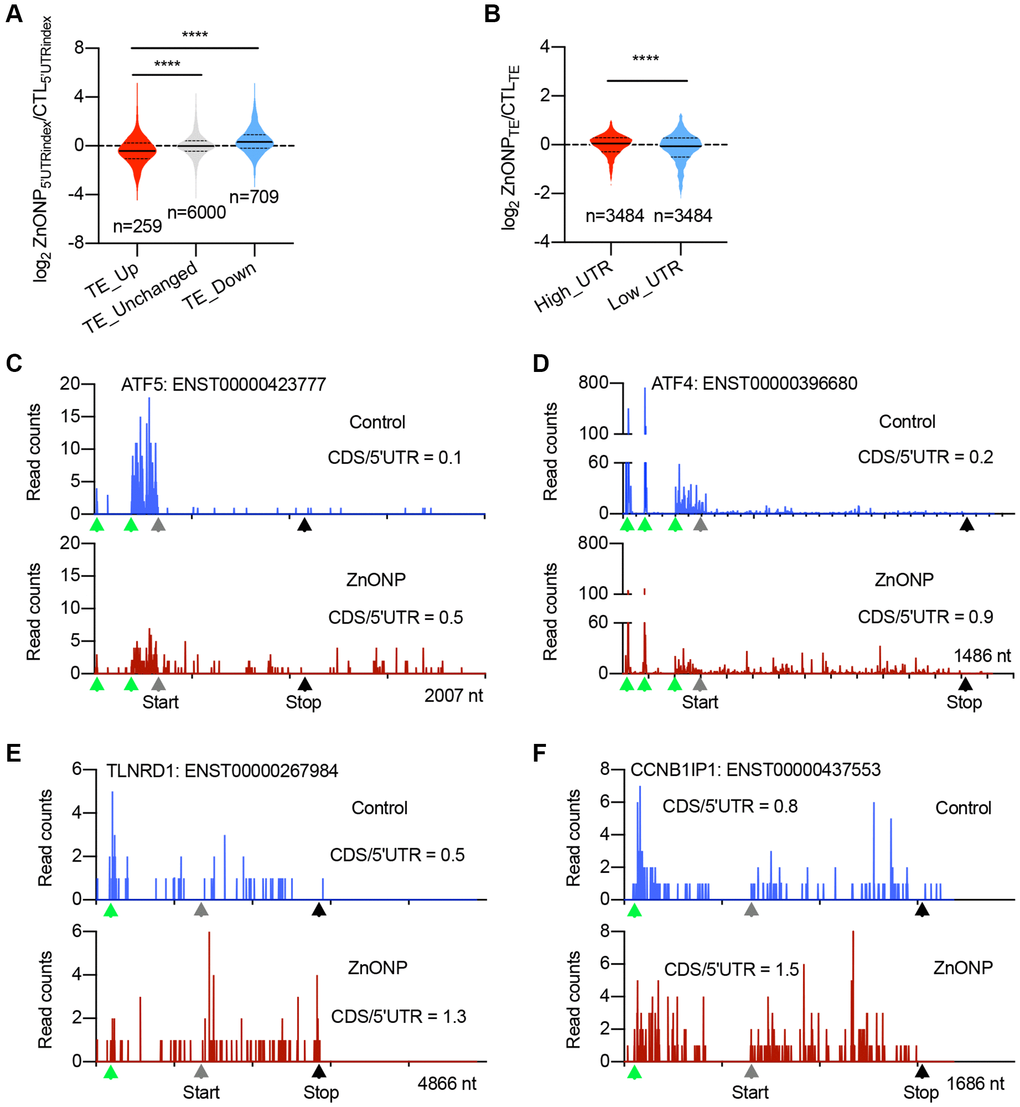
Figure 4.uORFs regulate the expression of upregulated genes. (A) Violin plots showing the RPFs of 5′UTR in translationally (TE) up-regulated, unchanged, and down-regulated genes after ZnO NPs treatment. The upper and lower quartiles and the median are shown for each group. ****P < 0.001, one-way ANOVA. (B) Violin plots showing the translation efficiency change after ZnO NPs treatment between high-UTR and low-UTR genes. The upper and lower quartiles and the median are indicated for each group. ****P < 0.001, t-test. (C–F) The indicated mRNAs whose ribosome densities increase at CDS and decrease at 5′UTR during ZnO NPs treatment. Ribosome density in ATF5 (C), ATF4 (D), TLNRD1 (E), and CCNB1IP1 (F) mRNAs are shown. The ratio of CDS RPFs to 5′UTR RPFs was indicated. The green triangles indicate the predicated start codons of uORF. The grey triangles indicate the start codons of CDS. The black triangles indicate the stop codons of CDS.