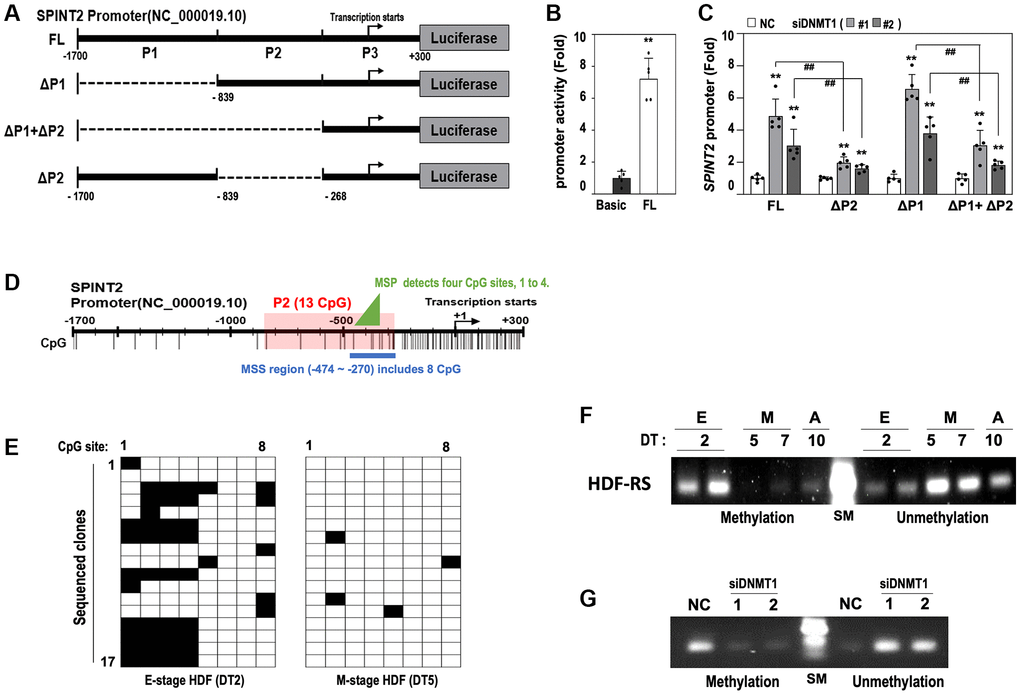
Figure 4.DNMT1 regulates SPINT2 expression via promoter methylation. (A) Schematic representation of SPINT2 promoter constructs used in the luciferase reporter assay, with indicated deletion sites (FL, full-length of −1700 to +300; ΔP1, deletion of −1700 to −839; ΔP1+ΔP2, deletion of −1700 to −268; ΔP2, deletion of −839 to −268). (B) HDFs (DT2) were transfected with pGL4-basic or pGL4-SPINT2 promoter reporter constructs for 4 days, followed by luciferase assay. (C) HDFs (DT2) were transfected with siRNAs targeting negative control (NC) or DNMT1 (siDNMT1) for 24 hours, then transfected with SPINT2 promoter reporter constructs for 2 days. Cell lysates were analyzed for luciferase activity. (D) Schematic representation of CpG islands in the SPINT2 promoter and selected regions used for DNA methylation-specific sequencing (MSS) and methylation-specific PCR (MSP) analyses. (E) MSS analysis of DNA clones from DT2 (early-stage) and DT5 (middle-stage) HDFs. Methylated CpG sites are shown in black, and unmethylated sites in white. Seventeen DNA clones were sequenced per group. (F) Methylation-specific PCR (MSP) analysis of the four CpG methylation hot spots (−445, −366, −363, and −358) identified by MSS. SM indicates DNA size marker. (G) HDFs (DT2) were transfected with siRNAs for negative control (NC) or DNMT1 (siDNMT1) for 3 days and then subjected to MSP analysis against the 4 CpG methylation hot spots.