Introduction
By 2030, the US Census Bureau projects that one in five people in the US alone will be over the age of 65 [1], a major risk factor for many of the most prevalent, costly, and devastating diseases of today, including cancer, cardiovascular disease, Alzheimer’s disease, and Type II diabetes [2]. To offset the burden of this increase, efforts are underway to develop an anti-aging drug or other geroprotective intervention that could extend healthspan, lower disease rates, and maintain productivity in this age group.
Unfortunately, there are many roadblocks to such an intervention. While many aging mechanisms are now catalogued [3] and hundreds of databased drugs extend lifespan in animal models [4,5], approval and testing of new drugs in humans is slow, expensive, and prone to high failure rates. This is particularly true in longevity research and exacerbated by a lack of reliable aging biomarkers [6,7] other than disease itself [6,8]. Even if successful, to be used preventatively, anti-aging drugs face extraordinarily high safety and efficacy standards for approval [9].
One strategy to hasten the process has been the repurposing of existing, FDA-approved drugs that show off-label anti-cancer and anti-aging potential [10,11], and at the top of that list are metformin and rapamycin, two drugs that mimic caloric restriction [12].
Caloric restriction is a well-known intervention for extending lifespan across species [13] , but has limited practical value in humans [14]. Mimetics of caloric restriction would theoretically exert its beneficial effects without actual reduction in caloric intake. Hallmarks of caloric restriction include reduced levels of circulating glucose and insulin as well as beneficial responses to these reductions in nutrient- and energy-sensing net-works, such as activation of AMP-activated protein kinase (AMPK) and inhibition of mammalian target of rapamycin (mTOR) [15]. The mTOR pathway is a particularly important growth pathway essential for early development but also potentially detrimental in later years if not suppressed, contributing to geroconversion, cellular senescence, disease and decline [16] . Inhibition of the mTOR pathway slows conversion to senescence [16] and extends longevity across species, including Saccharomyces cerevisiae (yeast) [17], Caenorhabditis elegans (nematodes) [18,19], and Mus musculus (mice) [12,20–22] .
Rapamycin and metformin, while distinct in clinical use, are both mTOR inhibitors and exhibit multiple anti-aging, anticancer, and anti-cardiovascular disease benefits [23].
Rapamycin (sirolimus) is an immunosuppressant used following renal transplantation, but also has life-extending properties in multiple animal models, including yeast [24], Drosophila melanogaster (fruit flies) [25], and mice [26,27], though effects can be sex and genotype-dependent [28] . In renal transplant patients, rapamycin has been shown to reduce cancer risk post-surgery [29–34]. It also has significant anti-cancer properties in mice [35–37]. While the extent to which its anticancer properties underlie its anti-aging effects and/or vice versa remains a point of discussion [15,38,39], as an anti-aging agent it has also been reported or theorized to protect against a number of other aging-related diseases in humans: cardio-vascular disease, osteoporosis, obesity, autoimmune disease and arthritis, macular degeneration, diabetes, Alzheimer’s disease, and Parkinson’s disease [16]. While rapamycin interacts with various nutrient signalling-related pathways, it acts primarily as an mTOR inhibitor, via direct inhibition of mTOR complex 1 (mTORC1) [23]. Analogs of rapamycin, or rapalogs (e.g. everolimus), are currently in use as anticancer drugs [40]. Also, mTORins, dual mTOR kinase inhibitors, are in development as anticancer agents, but much remains undetermined, such as proper dosage, toxicity, and adverse effects [15,38].
Like rapamycin, metformin is also an mTOR inhibitor, although indirectly so and via multiple mechanisms [41–45]. Metformin is a biguanide most renowned as the first-line treatment for type II diabetes and meta-bolic syndrome. It corrects hyperglycemia primarily by lowering hepatic gluconeogenesis but also by increasing insulin sensitivity and lowering levels of circulating lipids [9]. Its effects, however, appear to be pleiotropic, with benefits extending to a number of other age-related conditions in humans, including cancer [46,47] and cardiovascular disease [10].
In animal models as well, multiple beneficial effects of metformin have been reported across species with varying anticancer and prolongevity effects, including AMPK-mediated improvements in cutaneous wound healing [48]. Results, however, depend on dosage, sex, and age at onset of treatment [49–53], factors relevant to widescale, prophylactic metformin use in humans [49,50].
Metformin’s mechanisms of action have been extensively studied but are complex and remain only partially understood. Although metformin inhibits mTOR [43-45], its primary mode of action may be inhibition of mitochondrial complex I [54–62]. This action leads, among other things, to beneficial changes in cellular energy status and activation of AMPK [51,59, 62–66], a cellular energy sensor with a broad range of downstream effects on cellular function [ 67]. Through a combination of AMPK-dependent and -independent mechanisms [68], metformin influences a number of signaling pathways, including IGF-1 [69], hepatic sirtuin 1 (SIRT1) [70–73] and mTOR complex 1 (mTORC1) [74], that contribute directly or indirectly to its clinical response and multiple anticancer effects.
Taken together, rapamycin and metformin are promising candidates for life and healthspan extension; however, concerns of adverse side effects have hampered their widescale adoption for this purpose. While short term rapamycin use is considered safe, it has been reported to be associated with more adverse events than cyclosporin A in renal transplant patients, including wound complications, mouth ulcers, diarrhea, hypokalemia, bronchopneumonia, and proteinuria and higher discontinuation rates (28.2% vs 14.9%) [75–77]. In addition, chronic rapamycin use can lead to hepatic gluconeogenesis, insulin resistance, and severe glucose intolerance in rats [78], impaired glucose tolerance in mice [79], and even diabetes in male mice [80]. While rapamycin-induced diabetes is argued to differ from true type II diabetes [81], rapamycin may require pairing with metformin to counter induced hyper-glycemia [40]. Metformin, while relatively safe, is poorly tolerated in one fourth to one half of patients due to gastrointestinal side effects [82], although preliminary findings suggest these can be alleviated in some with an extended-release form of the drug [83]. Metformin also carries a slight risk of lactic acidosis in certain individuals [84–86]. Interestingly, rapamycin lowers lactate production, so may buffer this risk [87]. Metformin and rapamycin in combination may have additional benefits; in vitro they potentiate chemo-therapy with mitotic inhibitors while protecting normal cells [41]. One suggestion has been varying dosage schedules and combinations of rapamycin with metformin and five other anti-aging compounds per individual to reduce side effects [40]. However, the best preventative, widescale intervention would be one for which risk is negligible.
Given the urgency of the present need for anti-aging, disease preventive interventions, it may be beneficial to look to natural alternatives, such as nutraceuticals, that would be safe enough to administer widely with little to no risk of harm and with fewer regulatory hurdles than drugs. Nutraceuticals have received considerable attention in recent years for potential roles in preventing or treating a number of age-related diseases [88].
In this work, we initiate an effort to identify safe, natural alternatives to metformin and rapamycin. Our work is done entirely in silico and entails the use of metformin and rapamycin transcriptional and signaling pathway activation signatures to screen for matches amongst natural compounds. We have shown previously that the transcriptional signature of a given drug response, disease state, or other physiological condition, when mapped to the signalome, can be useful for biomarker development [89–91] and drug screening [7,92,93]. Transcriptional signatures have been suggested by others as well for aiding in biomarker development [94], cancer drug screening [75] and repositioning [11], and diabetes management [95].
The transcriptional signature of metformin is particularly well-suited to this type of analysis, as it includes thousands of AMPK-dependent and AMPK-independent changes in gene expression related to a diverse set of signaling pathways [96]. AMPK itself acts in part by directly and indirectly regulating metabolic gene expression when activated [97]. Metformin’s transcriptional signature also shows considerable similarity to the gene expression signature of long term caloric restriction [98,99,49], which is thought to play a role in mediating its effects on lifespan [100,101].
Gene expression data is in general a highly valuable resource that is still underutilized in drug discovery. With the public banking of data such as the LINCS project resulting in large repositories of cellular signatures of drug responses and disease states, large-scale screening, signalome analysis, and deep learning can be employed at little cost to make new discoveries [102]. Yet due to the size, difficulty in cross-platform analysis, and high dimensionality of microarray datasets, much information remains unparsed.
To overcome and even exploit these challenges, we have developed bioinformatic methods including Oncofinder [92,103], Geroscope [93] , and in silico Pathway Activation Network Decomposition Analysis (iPANDA) [ 104], which extract robust, biologically relevant pathway activation signatures from the data by combining various elements of previous approaches. The iPANDA method in particular was recently shown to outperform other methods in cross-platform micro-array analysis, noise and dimensionality reduction, and production of robust sets of biomarkers and reliable pathway signatures [93]. Illustratively, it was used successfully to identify biomarkers for breast cancer subtypes by stratifying samples by pathway activation [104], however it has many other potential applications, including drug discovery and drug mimicry, as we will demonstrate herein. We are currently using iPANDA in several other applications, including mapping the transcriptional signature of senescence and screening for novel senolytics, drugs that would selectively eliminate senescent cells [8]. We have also previously developed deep learning methods involving training of deep neural networks (DNNs) to recognize transcriptional signatures and pathway activation signatures of drugs or disease states from microarray data or to predict adverse effects [93].
In the present study, we apply these methods to screen for nutraceuticals that mimic metformin and/or rapamycin. Using LINCS perturbation data, we reduce a list of over 800 natural compounds to a shortlist of candidate nutraceuticals that show both similarity to the target drugs and low adverse effects profiles [93]. We then discuss the top candidates in light of shared mechanisms and previously reported anticancer and other health benefits that may deem them particularly promising for future experimental validation.
Results
To screen for potential candidate nutraceuticals, we used gene expression data from the Library of Integrated Network-based Cellular Signatures (LINCS) L1000 dataset to investigate similarity to metformin and/or rapamycin at the gene and pathway level (Figure 1). We employed several complementary approaches, including conventional statistical methods, pathway scoring-based methods, and training of deep neural networks (DNN) for signature recognition. Additionally, to evaluate potential adverse effects of top-scoring natural compounds we utilised a set of deep learned predictors, trained on drug-induced transcriptional response data. One important attribute of natural compounds we also looked closely at was GRAS (Generally Recognized As Safe) status and safety data.
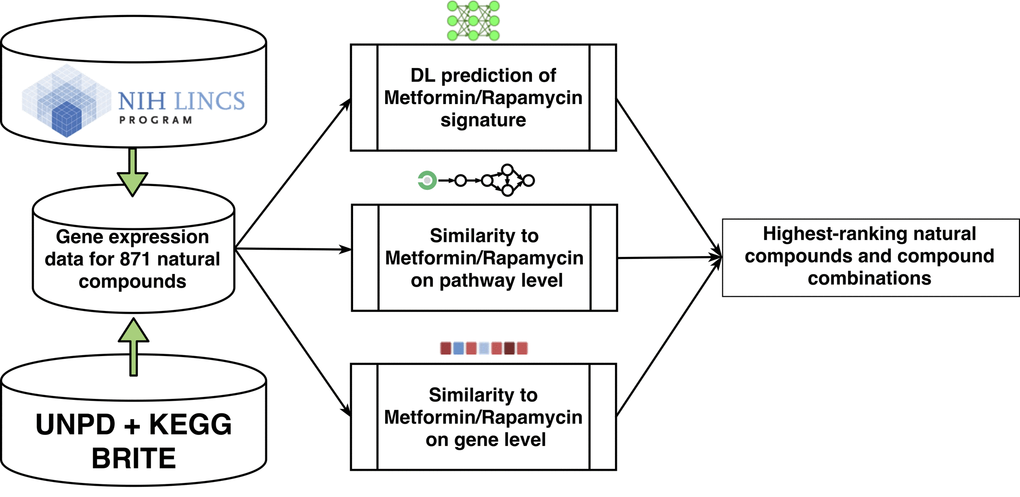
Figure 1. Workflow diagram showing multi-level analysis for screening and ranking nutraceuticals that mimic metformin and rapamycin in transcriptional and pathway activation response. A subset of 871 LINCS compounds were selected from the UNPD and KEGG BRITE databases. Perturbations with those compounds in cancer cell lines were compared with perturbations with metformin and rapamycin to estimate similarity at the gene and pathway level and deep learning techniques were employed to recognize the transcriptional signature of metformin and rapamycin and screen for matches amongst the LINCS compounds
Selection of natural compounds for screening
Prior to analysis, we filtered the LINCS dataset for compounds of natural origin by combining the compound lists from UNPD [105] and KEGG BRITE [106] databases and using the resulting list to select compounds in the LINCS dataset. In total, this resulted in 871 natural compounds with transcriptional response data across various times, concentrations, and cell lines. We utilized all available gene expression profiles for each compound, including metformin and rapamycin.
Deep learning-based scoring of compounds at gene level
For similarity scoring, we first used deep learning to train binary classifiers to recognize perturbations similar to metformin or rapamycin in transcriptional signature. A five fold cross-validation classifier for metformin and rapamycin achieved an F1-score of 0.725 and 0.905 and Matthews correlation of 0.705 and 0.896, respectively. Each sample corresponding to perturbation with a natural compound was run through each DNN classifier and assigned a probability. We used a threshold of 0.5 to determine the significant hits and then performed a Fisher’s exact test to estimate the statistical significance for each compound (Figure 2, Supplementary Table 1 and Supplementary Table 2).
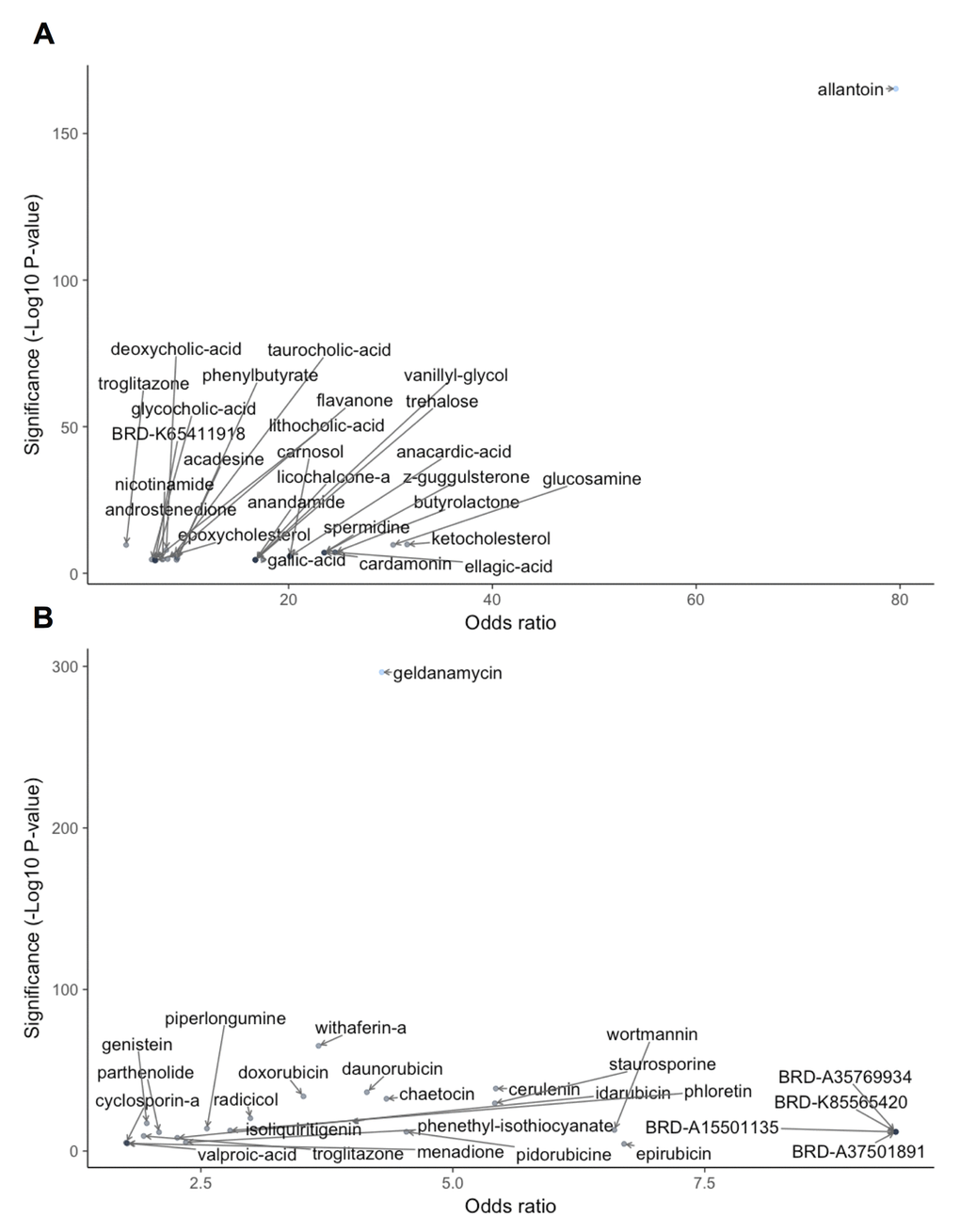
Figure 2. DL-based similarity to metformin (A) and rapamycin (B). Significance of natural compound was determined as the -log10(p-value) and odds ratio for compound according to Fisher's exact test performed on the DNN output for each perturbed sample. Only compounds with -log10(p-value)>4 and odds ratio > 1 are shown.
The compound exhibiting the highest similarity to metformin according to the metformin classifier (Fig. 2A) was allantoin, a key beneficial compound in yam (Dioscorea spp.). Like metformin, allantoin is a guanidinium derivative with anti-hyperglycemic effects [107,108]. It is an important metabolic intermediate of purine metabolism in many species across Eukarya and Bacteria domains [109,110]. Being a guanidinium derivative, allantoin is similar to metformin in structure and has been shown to induce glucose lowering effects via imidazoline I-2 receptors [107,108]. Other top hits for metformin included glucosamine, a compound used in the treatment of osteoarthritis [111,112], and cardamonin, a member of the anti-inflammatory chalcones found in plant-based foods [113], which inhibits mTOR and exhibits antitumor effects in vitro and in vivo [114] .
With the rapamycin classifier, the most significant hit was geldanamycin (Fig. 2B). Geldanamycin is an antibiotic belonging to Ansamycins family and targets the ADP/ATP binding site of heat shock protein 90 (Hsp90). Similarly to rapamycin, it has been shown to suppress the mTOR pathway through inhibition of the interaction between Hsp90 and RAPTOR [115]. Interestingly, the second most significant hit was withaferin A, which aligned with our subsequent results of gene- and pathway-level scoring for metformin and rapamycin, respectively. Other compounds with significant similarity to rapamycin according to the DNN classifier included another Hsp90 inhibitor, radicicol, several members of the anthracyclines antibiotic family used in cancer treatment (daunorubicin, idarubicin, doxorubicin, epirubicin) [116], cerulenin, a fatty acid synthase inhibitor with potential anticancer effects [117], chaetocin, being investigated as a histone lysine methyltransferase inhibitor [118,119], phloretin, an anti-tumor agent found in plant-based foods that shows effectiveness in inducing apoptosis in human lung cancer cells [120] and staurosporine, which also exhibited metformin similarity in subsequent results. The highest odds ratios were observed with four relatively unexplored compounds (BRD-A35769934, BRD-K85565420, BRD-A15501135, BRD-A37501891). Notably, 24 of 24 profiled samples for each of these compounds reached statistical sig-nificance.
Similarity at gene and pathway level
We next determined gene-level similarity of each compound to metformin and rapamycin using conventional statistical methods. First, this involved comparing each distinct time- and concentration-specific compound perturbation measured across various cell lines to corresponding DMSO-treated reference samples. We performed differential gene expression analysis to determine statistically significantly perturbed genes. Then, to screen for compounds with high similarity to metformin or rapamycin in terms of individual gene expression changes, we used Fisher’s Exact Test to directly compare all metformin or rapamycin perturbations to individual perturbations of other natural compounds (Supplementary Table 3).
To determine pathway-level similarity, we applied the iPANDA algorithm [104] to acquire pathway activation profiles for the same set of individual perturbations. For each compound, perturbation pathway activation scores (PAS) were calculated for 378 pathways. Similarity of pathway activation signatures of natural compounds to metformin and rapamycin was evaluated by the number of commonly up- and down-regulated pathways (Supplementary Table 4).
Combined results of gene- and pathway-level analysis are depicted in Figure 3. Gene-level analysis of similarity to metformin (Fig. 3A) showed that the most significant perturbation was associated with withaferin-A, a steroidal lactone that exhibits antidiabetic and anticancer properties [121]. Pathway-level scoring, on the other hand, demonstrated ginsenoside Rc, a compound isolated from ginseng, to be the top hit. Other compounds at the top of the list for significant gene- and pathway-level similarity to metformin included umbelliferone, a coumarin with antihyperglycemic, anti-inflammatory, and antitumor properties [122], coumaric acid, the p- isomer of which shows immunosuppressive, anti-inflammatory, and anti-diabetic properties [123,124], staurosporine, a kinase inhibitor with promising antitumor properties but poor selectivity [123–125], bile acids, which have been shown to have anti-cancer properties and specifically anti-hypoxic tumor effects [126], and ellipticine, a plant-derived compound with significant anticancer effects but issues with toxicity [ 127].
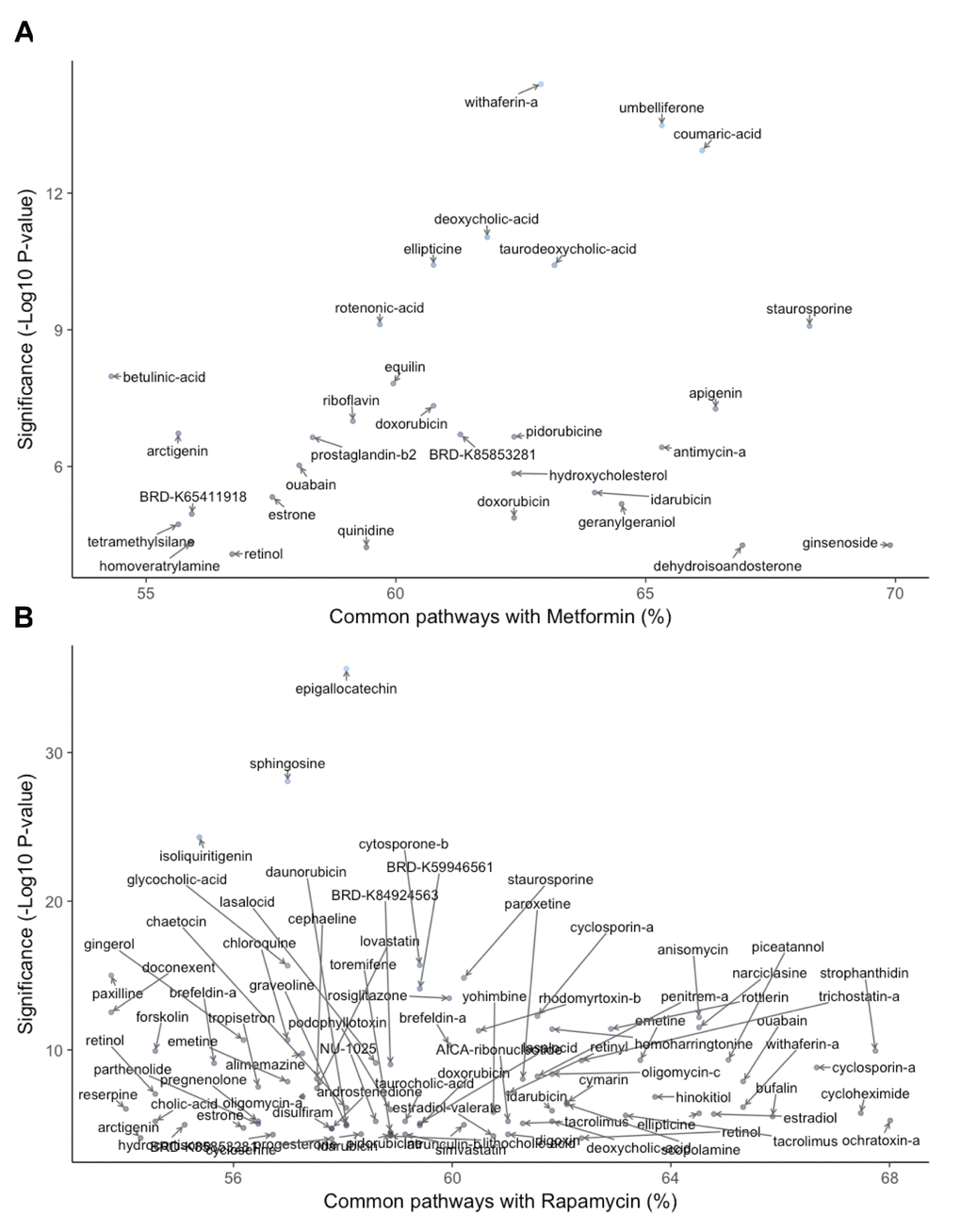
Figure 3. Gene- and pathway-level similarity to metformin (A) and rapamycin (B). Significance of natural compound was determined as the -log10(p-value) of the most significant perturbation of compound according to Fisher's exact test. Percentage of common pathways designates the amount of pathways that have the same direction of the change as Metformin. Only compounds with -log10(p-value)>4 and over 50% of common pathways are shown.
For rapamycin (Fig. 3B), the most significant hits at the gene level were epigallocatechin gallate (EGCG), a compound underlying the aging-related benefits of green tea, including protection against cancer, cardiovascular events, and UV-mediated skin aging [128], sphingosine, the precursor to sphingosine 1-phosphate, a second messenger implicated in several diseases, including multiple sclerosis, sepsis, cancer, and cardiovascular disease [129], and isoliquiritigenin, a compound shown to act as an anticancer, anti-cardio-vascular disease, antioxidant, antimicrobial, hepato-protective, and immunoprotective agent [130]. A number of other compounds were highly similar to rapamycin at the pathway level. These included strophanthidin, a compound recently identified in a similar LINCS screening as being likely to reverse cancer-related gene expression, which was validated in the liver hepatocellular carcinoma (LIHC) cell line [75], cyclosporin A, an immunosuppressant alternative to rapamycin following transplantation [75,76] cyclo-heximide, a highly toxic protein synthesis inhibitor used primarily in basic research, including cancer research [131], ochratoxin A, a potentially carcinogenic myco-toxin found and regulated in a wide variety of foods [111], and, notably due to its gene-level similarity to metformin above, withaferin A.
Effective combinations of natural compounds
Often, natural remedies with proven effectiveness consist of one or several plant species which can account for hundreds of natural compounds [132]. Accumulating evidence suggests that a combination of several compounds targeting multiple pathologic signaling circuits might be more advantageous than single agent treatments [133–137]. Examples of synergistic anti-aging effects of drug combinations with different targets have been reported [37,138]. This is particularly relevant to natural compounds with GRAS status, since the likelihood of serious adverse reactions is low.
For these reasons, we also estimated the similarity of different combinations of natural compounds to metformin. This required us to predict the transcriptional response after perturbation with a given combination of compounds. We chose to do this on the pathway level and to calculate the combinatorial response as the sum of individual PAS values corresponding to individual perturbations. We fully considered that additive effects on a pathway may be limited and other types of effects (e.g. synergistic, competitive, etc.) may be at play. Our rationale for assuming additivity was required for simplification, but we tested the additivity assumption for its predictive value with an external dataset and the results supported the approach. We used external dataset E-MEXP-3192 (http://www.ebi.ac.uk/arrayexpress, Supplementary Figure 1) [139], where the pathway activation signature of two compounds, retinoic acid and lapatinib, was explored, both individually and in combination, to predict their combinatorial drug effects by taking the sum of individual PAS values. Results, at least in the case of these two compounds, showed high similarity between the predicted and actual combinatorial pathway activation signatures, supporting the use of PAS additivity in this context (Supplementary Figure 1).
To investigate whether any of the natural compound combinations would produce better similarity scores than each compound independently, we selected four compounds with known beneficial effects and good safety profiles: withaferin-A, ginsenoside, apigenin and gamma linolenic acid (GLA).
We used our previously established database of aging-associated pathways and calculated PAS values for each compound (Supplementary Table 5). Then we devised all possible combinations of these compounds and estimated the resulting pathway activation as the sum of PAS values of individual compounds. Each of the combinations was compared to the profile of metformin and Pearson correlation coefficient was used as a similarity metric (Supplementary Table 6). Combinations outperformed the individual compounds, with similarity of the top 10 combinations ranging from 0.73-0.80 (Supplementary Table 6). As an example, we selected a combination of three nutraceuticals with high similarity to metformin, good safety profiles, and/or previously reported anti-aging, anticancer, or anti-disease potential: ginsenoside, GLA, and withaferin A. Overall pathway level similarity between metformin and the top combination of nutraceuticals is depicted in Supplementary Figure 2. Pathways with shared activation between metformin and the combination of these three compounds and each compound individually are shown in Supplementary Figure 2; the most significant of these were upregulation of JNK, cAMP, AKT, MAPK, ERK, and ILK pathways and down-regulation of ubiquitin proteosome signaling. To investigate whether similarity varied among met-activated, met-neutral, and met-inhibited pathways, we also examined correlations between metformin and the nutraceuticals and nutraceutical combination among these groups, with a designated threshold of 1 or -1 to define met-activated or met-inhibited pathways, respectively; results showed the strongest correlations with pathways inhibited by metformin (Supplementary Table 7).
Deep learning-predicted adverse effects
Additionally, to estimate the safety of investigated natural compounds we utilized our deep learned adverse effects prediction approach [93]. For every sample corresponding to perturbation with a natural compound, we ran an ensemble of 305 predictors each corresponding to a particular side effect category. Resulting probabilities were averaged for each side effect of each natural compound. Then, to estimate the overall adverse effects prediction of a compound, we calculated mean probability across all adverse effects and the number of adverse effects with probability >0.5 (Supplementary Table 8).
Interestingly, rapamycin was near the top of the list of compounds with the highest probability of adverse effects, with a maximum mean probability of 0.41 across all potential adverse effects and 134 total effects categories for which probability exceeded 0.5. Of these, the top ten adverse effects categories included cardiac and vascular, lipid, testicular and epididymal, skin, general, immunodeficiency, obstetric and gynecological, eye, neurological, and vascular/hypertensive, all with probabilities >0.9. Metabolic (0.86) and glucose/ diabetic (0.75) effects probabilities were also high for rapamycin. Other compounds with high mean adverse effects probabilities included anthracycline antibiotics, oligomycin-c, tacrolimus, paroxetine, benzethonium, wortmannin and triptolide. The safest compounds, on the other hand, with <3 significant adverse effects categories and mean overall probabilities <0.05, turned out to be the compounds with highest odds ratios for rapamycin similarity scoring (BRD-A35769934, BRD-K85565420, BRD-A15501135, BRD-A37501891) as well as tert-butylhydroquinone, lanatoside-c, syringic acid, morin, niacin and gossypetin (mean probabilities <0.10, 11 or fewer significant adverse effects categories). Metformin was predicted to have relatively few adverse effects, as well, with mean probability 0.2 and 25 significant adverse effects categories.
We then searched the adverse effects table against the list of candidate compounds selected above for metformin and rapamycin similarity to investigate predicted adverse effects. For metformin-like compounds, we found the following mean adverse effects probabilities and number of adverse effects categories: withaferin A (0.14, 52), staurosporine (0.17, 126), ginsenoside (0.25, 29), umbelliferone (0.24, 19), ellipticine (0.17, 69), allantoin (0.14, 28), glucosamine (0.25, 58), cardamonin (0.26, 66). For rapamycin-like compounds, we found similar probabilities and numbers of categories: ECGC (0.25, 31), sphingosine (0.20, 46), isoliquiritigenin (0.23, 88), strophanthidin (0.17, 38), cyclosporin A (0.26, 62), ochratoxin A (0.19, 39), geldanamycin (0.20, 57), radicicol (0.20, 87), cerulenin (0.22, 49), and chaetocin (0.09, 23).
Discussion
In this work, we introduce a rapid, low-cost route to drug mimicry via screening public gene expression datasets for compounds with shared transcriptional and signaling pathway activation signatures. The methods we employ [104] combine and outperform previous methods for pathway activation scoring and capitalize on vast, valuable but underutilized public repositories of microarray data, overcoming significant analytical challenges that have previously hindered their wide-scale use.
In an application of these methods, we focused on mimicry of metformin and rapamycin, seeking nutraceuticals that could preserve their anti-aging and disease-preventive potential while being better suited for wide-scale prophylactic use.
One of the most significant findings was withaferin A, one of only two only compounds topping the list for similarity to both metformin and rapamycin. Withaferin A was the top-scoring compound for gene-level similarity to metformin using the conventional statistical approach and also displayed significant pathway- and gene-level similarity to rapamycin using both the pathway activation approach and the deep learning approach. Withaferin A is a steroidal lactone derived from members of the Solanaceae family (e.g. Acnistus arborescens and Withania somnifera), commonly used in Ayurveda (traditional Indian medicine) for arthritis and menstrual disorders. Mounting evidence in rodent and cell-culture models indicate that it is an anti-diabetic, anti-obesity and anti-cancer agent with potent anti-oxidative, anti-inflammatory, anti-proliferative, apoptosis-inducing and leptin-sensitizing properties [121].
Mice with diet-induced obesity (DIO) have seen 20-25% reductions in body weight as a result of withaferin A treatment [140], as well as a decrease in obesity-associated pathology, e.g. hepatic steatosis. Withaferin A also has beneficial effects on glucose metabolism that are independent of its leptin-sensitizing effect.
Many of its anticancer properties result from its ability to inhibit cell proliferation and decrease glucose utilization, glycolysis and tricarboxylic acid (TCA) cycle intermediates [141]. Additionally, it has been found to be a potent inhibitor of angiogenesis. It inhibits cell proliferation via inhibition of cyclin D1 expression, as well as inhibition of NF-kappa B, which is thought to occur via interference with the ubiquitin-mediated proteasome pathway [142], as suggested by increased levels of polyubiquitinated proteins in cancer cells following treatment with withaferin A. It has also been found to selectively induce cell death in multiple types of tumor cells [143,144] . Its anticancer effects are mediated through modulation of a number of pathways, including inhibition of Notch 1 [145], inhibition of STAT3 activation [146–148], downregulation of the MTOR signalling components pS6K and p4E-BP1 [145], downregulation of the prosurvival pathway Akt/NF-kappaB/Bcl-2 [145], induction of c-Jun-NH(2)-kinase-mediated apoptosis [145], induction of apoptosis via upregulation of Bim, t-Bid, caspase-8, and DR5 [149] , suppression of constitutive and IL-6-induced phosphorylation of STAT3 (on Tyr705) and consequent down-regulation of the STAT3 regulated genes Bcl-xL, Bcl-2, cyclin D1 and survivin [150], inhibition of heat shock protein 90 [151], downregulation of COX-2 and iNOS by blocking NF-κB activity [121], and down-regulation of TNF-a [152].
Withaferin A was one of three compounds we included in the combination explored for metformin similarity. The other two were ginsenoside and GLA, which also demonstrate anti-aging, anticancer, and anti-disease potential in a number of studies.
Ginsenoside was the most similar compound to metformin at the pathway level. Used in traditional Chinese medicine, ginsenosides comprise a group of over 150 naturally occurring compounds isolated from plants of the Panax species (ginseng) [153]. Although the family is relatively diverse in term of chemical structure, most of its members share similar properties. A wide variety of benefits have been reported [153], including anticarcinogenic [154–157], immunomodulatory [157–161], anti‐inflammatory [162], antiallergic [163–165], antiatherosclerotic [166], antihypertensive [167,168], antidiabetic [169], anxiolytic [170,171] and antidepressant effects [172]. Ginsenosides activate AMPK [154,156,169,173] PI3K [169] and SirtI [169,174] pathways, promoting autophagy [154–156] and apoptosis [154–156] in cancer cells.
Another clear standout for metformin similarity was revealed by the DNN classifier, and that was allantoin, one of the active compounds mediating beneficial effects of yam. Yam powder, yam extract, and allantoin have been shown to improve B-cell function in maintaining insulin and glucose in a rat model of Type II diabetes, with antioxidative effects as well, improved lipid profiles, and increased release of glucagon-like peptide 1 (GLP1) [175]. In another study using the same rat model of diabetes, allantoin lowered plasma glucose levels by increasing ß-endorphin secretion, increasing GLUT4 expression, and increasing glucose uptake [108,176,177].
Overall, the most remarkable aspect of the metformin results was that, like metformin, several of the compounds scoring high in similarity exhibit glucose-lowering properties (withaferin A [121], umbelliferone [122] , and allantoin [107,108]) or anti-inflammatory effects (glucosamine [111,112] and cardamonin [113] ) in previous studies, and almost all of the top hits have shown anticancer potential, including withaferin A [141–144], ginsenoside [154–157], umbelliferone [122], staurosporine [123–125], bile acids [126], ellipticine [127], cardamonin [113], and allantoin [177]. This not only lends preliminary support to the validity of our methods, but also adds support to the evidence of metformin-like health benefits with these compounds.
Scoring for rapamycin overall revealed a larger number of significant hits compared to metformin, but more variation in the range of known effects, from beneficial to highly toxic. These also included several unnamed, novel candidates. Four of these relatively unexplored compounds (BRD-A35769934, BRD-K85565420, BRD-A15501135, BRD-A37501891) were the most significant in similarity to rapamycin and were also top-ranking in terms of safety, with extremely low probability of predicted adverse effects. These would be excellent novel candidates for characterization and validation in future work.
Like the metformin DNN classifier, the rapamycin classifier also revealed a clear standout amongst the compounds for rapamycin similarity, geldanamycin. Geldanamycin is an inhibitor of Hsp90 [178], which is an oncogenic target molecule overexpressed in many tumors [115,179]. Geldanamcyin is an inhibitor of mTOR signaling as well [115]. While initially promising as an potent anticancer agent [115,179,180], its hepatotoxicity has precluded its clinical use [180,181]; however, several less toxic derivatives have been developed [182], with 17AEP-GA and 17DMAG recently demonstrating growth suppression of multiple myeloma cells similar to geldanamycin [181]. Geldanamycin analog development is still an active area of research [182–185], with other analogs being recently shown to be effective against breast cancer cells [182,185]. In addition to geldanamycin, at least two of the other rapamycin hits in this study, radicicol and EGCG are also Hsp90 inhibitors [183,184]. Recently, a radicicol derivative, NW457, was shown to be effective against colorectal cancer both in vitro and in vivo [186].
Many of the other top hits for rapamycin show anticancer effects, including anthracyclines [116], cerulenin [117], isoliquiritigenin [130], strophanthidin [75], ECGC [128], phloretin [120], staurosporine [123–125], and withaferin A [141–144]. Several of the rapamycin-like compounds identified are known to modulate mTOR signaling. These include geldanamycin, which suppresses mTOR phosphorylation of downstream protein regulators [115], phloretin, a common flavonoid capable of inducing cell cycle arrest and apoptosis in part via suppression of AKT/PI3K/mTOR cascades [187], and isoliquiritigenin, another flavonoid that induces autophagic and apoptotic cell death in cancer cells via mTOR signaling [188]. Thus, like metformin, many of the compounds identified as being similar to rapamycin in transcriptional signature have been previously shown to have rapamycin-like properties. Other rapamycin-like compounds identified have mTOR-independent mechanisms, such as chae-tocin, a histone H3K9 methyltransferase inhibitor [119].
Finally, rapamycin had a remarkably high number of predicted adverse effects with our methods and significant similarity to at least two compounds known to be toxic, ochratoxin A and cycloheximide, although these toxic compounds were not predicted to have a wide variety of adverse effects (cycloheximide did score particularly high (0.86) in the toxicity category, however, as did strophanthidin (0.93)). This under-scores the need to look for rapamycin alternatives, and also raises interesting questions about the common (and distinct) mechanisms between rapamycin and the wide variety of rapamycin-like compounds, both beneficial and toxic.
The adverse effects prediction also enabled us to have a closer look at overall predicted safety of compounds of interest and likelihoods of specific adverse effects. None of the compounds discussed as similar to metformin or rapamycin stood out as extremely likely or unlikely to cause a wide variety of adverse effects; most scored in the low-moderate range, although this does not fully reflect the severity or importance of any one given adverse effects category for a given compound. Literature-based assessments of safety were also helpful; while several compounds are known to be toxic as noted, others are known to be relatively safe compounds found in plant-based foods, such as cardamonin and ECGC, or used in traditional medicine, such as withaferin A and ginsenosides. Safety in a preventative, chronic use context for each compound would have to be independently evaluated. Also, while there were no standout metformin-like candidates with an absence of gastrointestinal adverse effects, there were several rapamycin-like candidates with low likelihood of glucose/metabolic adverse effects, including withaferin A and ECGC. Perhaps the most notable compounds were the four unnamed compounds with similarity to rapamycin; their novelty, extremely low number of predicted adverse effects, including glucose/metabolic effects, and extremely high odds ratios for rapamycin similarity make them particularly intriguing candidates.
The in silico approach, while time- and cost-saving, does require several considerations in light of its role as a first-pass screening tool. First, and most importantly, the health-extending and adverse effects of all candidate nutraceuticals or other compounds will still require investigation and validation in vitro and in other cell lines, followed by validation in vivo in humans. This is particularly important in the case of nutraceuticals, as wide variation in their bioavailability and metabolism is a significant factor influencing the degree to which the predicted effects actually manifest in vivo [189]. Secondly, our approach hinges entirely upon the biological relevance of the short term (<48 hours) transcription-level response to a drug, and as such does not account for post-transcriptional and post-translational effects on a given pathway or long term changes, which may be biologically or clinically more important [190]. That said, numerous studies have demonstrated the value in using such expression signatures in the characterization of drug response [191].
Thus, while it cannot be overstated that our results will require validation, this work reduces a list of over 800 natural compounds to a manageable shortlist of a few strong candidates for metformin and rapamycin mimicry, substantiated by their similarity to the target drugs in transcriptional response. Several of these compounds are unnamed, novel candidates. Many of the others have known anticancer or other beneficial effects and now are demonstrated to share common cellular signatures with two known anticancer, anti-aging drugs, thus supporting previous findings and further investigation into their potential benefits. That so many compounds with anticancer and other health benefits share common transcriptional signatures raises interesting questions about what pathways are shared and distinct and which shared pathways are most critical to their beneficial effects. This has not only direct practical value in a narrow sense with the search for metformin and rapamycin mimetics, but has broader usefulness for any number of applications in drug discovery. If widely adopted, our approaches have the potential to significantly expedite drug development timelines, reducing cost by offering a viable and biologically-relevant means of screening and ranking compounds prior to in vitro studies and, since screening is based on human data, possibly in place of animal models. Improving our ability to predict the actions of a nutraceutical or drug in humans will give in silico-based approaches enormous utility in streamlining drug discovery, repurposing and development in the years to come.
Methods
Transcriptomic data
To obtain transcriptomic and signaling pathway activation signatures, we utilized transcriptional response data provided by LINCS Project (http://www.lincsproject.org/). To obtain a list of natural compounds present in the LINCS data set we used the UNPD database of natural compounds [105] in combination with 3 compound classification categories derived from KEGG BRITE Database [106]: “Phytochemical compounds”, “Phytochemicals used as drugs” and “Natural toxins”. The natural compound list was then compared to the list from the LINCS data set and 871 compounds were identified.
For each of these compounds, we extracted the level 3 (Q2NORM) gene expression data for each available cell line perturbed with each concentration of compound independently for all available timepoints. In the pathway-level analysis, for each case sample group perturbed with a compound, we generated a reference group consisting of samples perturbed with DMSO that came from the same RNA plate as samples from the case group. We analysed transcriptional response to perturbation with metformin, rapamycin, and a number of nutraceuticals as assayed in various cancer cell lines.
Differential expression
For transcriptome data, a limma test of differential gene expression was used. Each set of differentially expressed genes was ordered according to the following measure, which takes into account both the magnitude and statistical significance of the effect: FC * max(0, -log(p-value)), where FC is fold-change of gene expression between groups and p-value represents the result of limma test.
Gene level similarity to metformin/rapamycin
A statistically motivated score estimating the similarity of a compound was designed. Significantly up- or down-regulated genes were defined as those with an FDR-adjusted p-value of <0.01. A Fisher’s exact test was used to measure the association between two characteristics of each gene: being significantly down-regulated following metformin/rapamycin treatment and being significantly downregulated following treatment with each investigated compound in our compound library. The same test was performed for upregulated genes. The best of p-values of those two tests were taken as a score for the given drug or compound. A multiple testing correction of the obtained p-values for the amount of compound perturbations under study was performed.
Pathway-level similarity analysis
Pathway activation analysis is a powerful tool for extracting biologically-relevant properties from large transcriptomic datasets, enabling the generation of novel results prior to or in place of in vitro and in vivo experimentation. We have recently reported on a novel deep learning-based algorithm, the in silico Pathway Activation Network Decomposition Analysis (iPANDA), which we applied to large-scale trans-criptomic datasets as a tool for biomarker identification [104]. In contrast to other methods of pathway activation analysis, iPANDA generates pathway activation scores (PAS) by using precalculated gene coexpression data in combination with gene importance factors quantified according to the degree of differential gene expression and pathway topology decomposition. Here, we applied the same general approach to the task of drug mimicry, ranking existing nutraceutical compounds and compound combinations according to their transcriptomic signature’s degree of similarity to the known transcriptomic signature of metformin and rapamycin.
For pathway-level similarity analysis we chose gene expression samples of drug induced transcriptional response from A549 cell line. Signaling pathway activation scores for 378 total pathways from SABiosciences collection (http://www.sabiosciences.com/pathwaycentral.php) were calculated for each perturbation of 871 natural compounds, including Metformin and Rapamycin, using iPANDA algorithm [104]. Similarity of two perturbations was measured as percent of commonly up- or down-regulated pathways between them.
Combination scoring. Additivity hypothesis was checked on the dataset E-MEXP-3192 (http://www.ebi.ac.uk/arrayexpress). Preprocessed gene expression data corresponding to samples that underwent 12 hour treatment with 100nm retinoic acid, 100 nm lapatinib and their combination was analysed with iPANDA algorithm. For several selected compounds pathway analysis was done for 97 age-related pathways [ 93 ]. PAS values for withaferin-A, ginsenoside, apigenin and GLA were measured in PC3 cells perturbed for 24 hours with 10μM of the compound with the exception of To estimate the combinatorial effect of 5 selected natural compounds with GRAS status PAS scores were summed for each combination of two or more compounds. Then we used Pearson correlation coef-ficient between metformin and the combination to estimate the similarity.
Deep learning prediction of metformin/rapamycin signature and adverse effects
Deep neural networks (DNNs) were trained with transcriptional response data from the LINCS L1000 dataset. All available perturbations from MCF7, PC3, VCAP, A549, A375 and HT29 cell lines related to Metformin (perturbation id: BRD-K79602928) and Rapamycin (perturbation ids: BRD-A23770159, BRD-A50287119, BRD-A79768653, BRD-K84937637, BRD-K89626439, BRD-K99369265) were independently used and contributed to two training sets. Training and test sets were split at 80/20 ratio. For the Metformin prediction we used 67309 samples as the training set (98 samples are labeled as positive class) and 15788 samples as the test set (24 samples are labeled as positive class). For the Rapamycin prediction we used 68421 samples as the training set (517 samples are labeled as positive class) and 14677 samples as the test set (114 samples are labeled as positive class). The DNN was built by adjusting its hyperparameters (e.g. number of layers, activation function, etc.) on the training set and subsequently measuring the performance of the trained neural network on the test set. All experiments were conducted using an NVIDIA Titan X.
We used multilayer feed-forward neural networks as deep models (i.e. having more than 3 layers). Gridsearch algorithm was used for multiple hyper-parameters optimization in order to achieve the greatest predictive accuracy. We minimized the binary cross entropy loss function using a backpropagation algorithm. We used the Leaky ReLU activation function [192] in each layer, ADAM as optimizer of the cost function [193], dropout with 25% probability after each layer for the purposes of regularization [194] and additional L1 penalty on layer parameters.
Adverse effects for drugs were derived from SIDER database [83]. Side effect categories were mapped onto 321 preferred term from MedDRA v16.0 ontology [84]. An ensemble of class-specific DNNs with binary output was trained in a similar way to the methodology describ-ed previously [85]. All probabilities related to in a single side effect and perturbation id of the drug were aggregated.
Supplementary Materials
Acknowledgements
We would like to thank NVIDIA for assistance with the GPU equipment.
Conflicts of Interest
Andrew G. Swick and Darla Karpinsky-Semper are employed by Life Extension, and Aliper A, Zhavoronkov A, and Artemov A are employed by Insilico Medicine. Life Extension and Insilico Medicine are collaborating on product and biomarker development.
Funding
This study was supported in part with a research grant from the Life Extension Foundation 2016-LEF-AA-INSIL. Insilico Medicine is grateful to Nvidia Corporation for providing Tesla K80 GPUs and early access to the NVIDIA DevBox used in this study.
References
- 1. Colby SL, Ortman JM. Projections of the size and composition of the US population: 2014 to 2060. US Census Bureau. 2015; 9.
- 2. Niccoli T, Partridge L. Ageing as a risk factor for disease. Curr Biol. 2012; 22:R741–52. https://doi.org/10.1016/j.cub.2012.07.024 [PubMed]
- 3. Moskalev A, Zhikrivetskaya S, Shaposhnikov M, Dobrovolskaya E, Gurinovich R, Kuryan O, Pashuk A, Jellen LC, Aliper A, Peregudov A, Zhavoronkov A. Aging Chart: a community resource for rapid exploratory pathway analysis of age-related processes. Nucleic Acids Res. 2016; 44:D894–99. https://doi.org/10.1093/nar/gkv1287 [PubMed]
- 4. Barardo D, Thornton D, Thoppil H, Walsh M, Sharifi S, Ferreira S, Anžič A, Fernandes M, Monteiro P, Grum T, Cordeiro R, De-Souza EA, Budovsky A, et al. The DrugAge database of aging-related drugs. Aging Cell. 2017; 16:594–97. https://doi.org/10.1111/acel.12585 [PubMed]
- 5. Moskalev A, Chernyagina E, de Magalhães JP, Barardo D, Thoppil H, Shaposhnikov M, Budovsky A, Fraifeld VE, Garazha A, Tsvetkov V, Bronovitsky E, Bogomolov V, Scerbacov A, et al. Geroprotectors.org: a new, structured and curated database of current therapeutic interventions in aging and age-related disease. Aging (Albany NY). 2015; 7:616–28. https://doi.org/10.18632/aging.100799 [PubMed]
- 6. Blagosklonny MV. Validation of anti-aging drugs by treating age-related diseases. Aging (Albany NY). 2009; 1:281–88. https://doi.org/10.18632/aging.100034 [PubMed]
- 7. Zhavoronkov A, Buzdin AA, Garazha AV, Borisov NM, Moskalev AA. Signaling pathway cloud regulation for in silico screening and ranking of the potential geroprotective drugs. Front Genet. 2014; 5:49. https://doi.org/10.3389/fgene.2014.00049 [PubMed]
- 8. Moskalev A, Anisimov V, Aliper A, Artemov A, Asadullah K, Belsky D, Baranova A, de Grey A, Dixit VD, Debonneuil E, Dobrovolskaya E, Fedichev P, Fedintsev A, et al. A review of the biomedical innovations for healthy longevity. Aging (Albany NY). 2017; 9:7–25. https://doi.org/10.18632/aging.101163 [PubMed]
- 9. Moskalev A, Chernyagina E, Tsvetkov V, Fedintsev A, Shaposhnikov M, Krut’ko V, Zhavoronkov A, Kennedy BK. Developing criteria for evaluation of geroprotectors as a key stage toward translation to the clinic. Aging Cell. 2016; 15:407–15. https://doi.org/10.1111/acel.12463 [PubMed]
- 10. Pryor R, Cabreiro F. Repurposing metformin: an old drug with new tricks in its binding pockets. Biochem J. 2015; 471:307–22. https://doi.org/10.1042/BJ20150497 [PubMed]
- 11. Cardone L. Biocomputing drug repurposing toward targeted therapies. Aging (Albany NY). 2016; 8:2609–10. https://doi.org/10.18632/aging.101135 [PubMed]
- 12. Ingram DK, Anson RM, de Cabo R, Mamczarz J, Zhu M, Mattison J, Lane MA, Roth GS. Development of calorie restriction mimetics as a prolongevity strategy. Ann N Y Acad Sci. 2004; 1019:412–23. https://doi.org/10.1196/annals.1297.074 [PubMed]
- 13. Mattison JA, Colman RJ, Beasley TM, Allison DB, Kemnitz JW, Roth GS, Ingram DK, Weindruch R, de Cabo R, Anderson RM. Caloric restriction improves health and survival of rhesus monkeys. Nat Commun. 2017; 8:14063. https://doi.org/10.1038/ncomms14063 [PubMed]
- 14. Phelan JP, Rose MR. Why dietary restriction substantially increases longevity in animal models but won’t in humans. Ageing Res Rev. 2005; 4:339–50. https://doi.org/10.1016/j.arr.2005.06.001 [PubMed]
- 15. Zhavoronkov A. Inhibitors of mTOR in aging and cancer. Oncotarget. 2015; 6:45010–11. https://doi.org/10.18632/oncotarget.6878 [PubMed]
- 16. Blagosklonny MV. Rejuvenating immunity: “anti-aging drug today” eight years later. Oncotarget. 2015; 6:19405–12. https://doi.org/10.18632/oncotarget.3740 [PubMed]
- 17. Fabrizio P, Pozza F, Pletcher SD, Gendron CM, Longo VD. Regulation of longevity and stress resistance by Sch9 in yeast. Science. 2001; 292:288–90. https://doi.org/10.1126/science.1059497 [PubMed]
- 18. Jia K, Chen D, Riddle DL. The TOR pathway interacts with the insulin signaling pathway to regulate C. elegans larval development, metabolism and life span. Development. 2004; 131:3897–906. https://doi.org/10.1242/dev.01255 [PubMed]
- 19. Vellai T, Takacs-Vellai K, Zhang Y, Kovacs AL, Orosz L, Müller F. Genetics: influence of TOR kinase on lifespan in C. elegans. Nature. 2003; 426:620–620. https://doi.org/10.1038/426620a [PubMed]
- 20. Wu JJ, Liu J, Chen EB, Wang JJ, Cao L, Narayan N, Fergusson MM, Rovira II, Allen M, Springer DA, Lago CU, Zhang S, DuBois W, et al. Increased mammalian lifespan and a segmental and tissue-specific slowing of aging after genetic reduction of mTOR expression. Cell Reports. 2013; 4:913–20. https://doi.org/10.1016/j.celrep.2013.07.030 [PubMed]
- 21. Selman C, Tullet JM, Wieser D, Irvine E, Lingard SJ, Choudhury AI, Claret M, Al-Qassab H, Carmignac D, Ramadani F, Woods A, Robinson IC, Schuster E, et al. Ribosomal protein S6 kinase 1 signaling regulates mammalian life span. Science. 2009; 326:140–44. https://doi.org/10.1126/science.1177221 [PubMed]
- 22. Harrison DE, Strong R, Sharp ZD, Nelson JF, Astle CM, Flurkey K, Nadon NL, Wilkinson JE, Frenkel K, Carter CS, Pahor M, Javors MA, Fernandez E, Miller RA. Rapamycin fed late in life extends lifespan in genetically heterogeneous mice. Nature. 2009; 460:392–95. https://doi.org/10.1038/nature08221 [PubMed]
- 23. Roth GS, Ingram DK. Manipulation of health span and function by dietary caloric restriction mimetics. Ann N Y Acad Sci. 2016; 1363:5–10. https://doi.org/10.1111/nyas.12834 [PubMed]
- 24. Alvers AL, Wood MS, Hu D, Kaywell AC, Dunn WA
Jr , Aris JP. Autophagy is required for extension of yeast chronological life span by rapamycin. Autophagy. 2009; 5:847–49. https://doi.org/10.4161/auto.8824 [PubMed] - 25. Bjedov I, Toivonen JM, Kerr F, Slack C, Jacobson J, Foley A, Partridge L. Mechanisms of life span extension by rapamycin in the fruit fly Drosophila melanogaster. Cell Metab. 2010; 11:35–46. https://doi.org/10.1016/j.cmet.2009.11.010 [PubMed]
- 26. Arriola Apelo SI, Pumper CP, Baar EL, Cummings NE, Lamming DW. Intermittent Administration of Rapamycin Extends the Life Span of Female C57BL/6J Mice. J Gerontol A Biol Sci Med Sci. 2016; 71:876–81. https://doi.org/10.1093/gerona/glw064 [PubMed]
- 27. Miller RA, Harrison DE, Astle CM, Baur JA, Boyd AR, de Cabo R, Fernandez E, Flurkey K, Javors MA, Nelson JF, Orihuela CJ, Pletcher S, Sharp ZD, et al. Rapamycin, but not resveratrol or simvastatin, extends life span of genetically heterogeneous mice. J Gerontol A Biol Sci Med Sci. 2011; 66:191–201. https://doi.org/10.1093/gerona/glq178 [PubMed]
- 28. Swindell WR. Meta-Analysis of 29 Experiments Evaluating the Effects of Rapamycin on Life Span in the Laboratory Mouse. J Gerontol A Biol Sci Med Sci. 2017; 72:1024–32. https://doi.org/10.1093/gerona/glw153 [PubMed]
- 29. Yakupoglu YK, Buell JF, Woodle S, Kahan BD. Individualization of immunosuppressive therapy. III. Sirolimus associated with a reduced incidence of malignancy. Transplant Proc. 2006; 38:358–61. https://doi.org/10.1016/j.transproceed.2006.01.019 [PubMed]
- 30. Campistol JM, Eris J, Oberbauer R, Friend P, Hutchison B, Morales JM, Claesson K, Stallone G, Russ G, Rostaing L, Kreis H, Burke JT, Brault Y, et al. Sirolimus therapy after early cyclosporine withdrawal reduces the risk for cancer in adult renal transplantation. J Am Soc Nephrol. 2006; 17:581–89. https://doi.org/10.1681/ASN.2005090993 [PubMed]
- 31. Garrick R. Sirolimus Therapy after Early Cyclosporine Withdrawal Reduces the Risk for Cancer in Adult Renal Transplantation. Year Book Med. 2007; 2007:226–27. https://doi.org/10.1016/S0084-3873(08)70153-6
- 32. Campistol JM, Eris J, Oberbauer R, Friend P, Hutchison B, Morales JM, Claesson K, Stallone G, Russ G, Rostaing L, Kreis H, Burke JT, Brault Y, et al. Sirolimus therapy after early cyclosporine withdrawal reduces the risk for cancer in adult renal transplantation. J Am Soc Nephrol. 2006; 17:581–89. https://doi.org/10.1681/ASN.2005090993 [PubMed]
- 33. Zmonarski SC, Boratyńska M, Bernat B, Kamińska D, Rabczyński J, Klinger M. Regression of Kaposis’s sarcoma in renal graft recirients after conversion to Sirolimus treatment. Transplantation. 2004; 78:500. https://doi.org/10.1097/00007890-200407271-01348
- 34. Cullis B, D’Souza R, McCullagh P, Harries S, Nicholls A, Lee R, Bingham C. Sirolimus-induced remission of posttransplantation lymphoproliferative disorder. Am J Kidney Dis. 2006; 47:e67–72. https://doi.org/10.1053/j.ajkd.2006.01.029 [PubMed]
- 35. Anisimov VN, Zabezhinski MA, Popovich IG, Piskunova TS, Semenchenko AV, Tyndyk ML, Yurova MN, Rosenfeld SV, Blagosklonny MV. Rapamycin increases lifespan and inhibits spontaneous tumorigenesis in inbred female mice. Cell Cycle. 2011; 10:4230–36. https://doi.org/10.4161/cc.10.24.18486 [PubMed]
- 36. Anisimov VN, Zabezhinski MA, Popovich IG, Piskunova TS, Semenchenko AV, Tyndyk ML, Yurova MN, Antoch MP, Blagosklonny MV. Rapamycin extends maximal lifespan in cancer-prone mice. Am J Pathol. 2010; 176:2092–97. https://doi.org/10.2353/ajpath.2010.091050 [PubMed]
- 37. Danilov A, Shaposhnikov M, Plyusnina E, Kogan V, Fedichev P, Moskalev A. Selective anticancer agents suppress aging in Drosophila. Oncotarget. 2013; 4:1507–26. https://doi.org/10.18632/oncotarget.1272 [PubMed]
- 38. Leontieva OV, Demidenko ZN, Blagosklonny MV. Dual mTORC1/C2 inhibitors suppress cellular geroconversion (a senescence program). Oncotarget. 2015; 6:23238–48. https://doi.org/10.18632/oncotarget.4836 [PubMed]
- 39. Blagosklonny MV. Prevention of cancer by inhibiting aging. Cancer Biol Ther. 2008; 7:1520–24. https://doi.org/10.4161/cbt.7.10.6663 [PubMed]
- 40. Blagosklonny MV. From rapalogs to anti-aging formula. Oncotarget. 2017; 8:35492–507. https://doi.org/10.18632/oncotarget.18033 [PubMed]
- 41. Apontes P, Leontieva OV, Demidenko ZN, Li F, Blagosklonny MV. Exploring long-term protection of normal human fibroblasts and epithelial cells from chemotherapy in cell culture. Oncotarget. 2011; 2:222–33. https://doi.org/10.18632/oncotarget.248 [PubMed]
- 42. Castillo-Quan JI, Blackwell TK. Metformin: Restraining Nucleocytoplasmic Shuttling to Fight Cancer and Aging. Cell. 2016; 167:1670–71. https://doi.org/10.1016/j.cell.2016.11.058 [PubMed]
- 43. Nair V, Sreevalsan S, Basha R, Abdelrahim M, Abudayyeh A, Rodrigues Hoffman A, Safe S. Mechanism of metformin-dependent inhibition of mammalian target of rapamycin (mTOR) and Ras activity in pancreatic cancer: role of specificity protein (Sp) transcription factors. J Biol Chem. 2014; 289:27692–701. https://doi.org/10.1074/jbc.M114.592576 [PubMed]
- 44. Pérez-Revuelta BI, Hettich MM, Ciociaro A, Rotermund C, Kahle PJ, Krauss S, Di Monte DA. Metformin lowers Ser-129 phosphorylated α-synuclein levels via mTOR-dependent protein phosphatase 2A activation. Cell Death Dis. 2014; 5:e1209. https://doi.org/10.1038/cddis.2014.175 [PubMed]
- 45. Kickstein E, Krauss S, Thornhill P, Rutschow D, Zeller R, Sharkey J, Williamson R, Fuchs M, Köhler A, Glossmann H, Schneider R, Sutherland C, Schweiger S. Biguanide metformin acts on tau phosphorylation via mTOR/protein phosphatase 2A (PP2A) signaling. Proc Natl Acad Sci USA. 2010; 107:21830–35. https://doi.org/10.1073/pnas.0912793107 [PubMed]
- 46. Evans JM, Donnelly LA, Emslie-Smith AM, Alessi DR, Morris AD. Metformin and reduced risk of cancer in diabetic patients. BMJ. 2005; 330:1304–05. https://doi.org/10.1136/bmj.38415.708634.F7 [PubMed]
- 47. Anisimov VN. Metformin for aging and cancer prevention. Aging (Albany NY). 2010; 2:760–74. https://doi.org/10.18632/aging.100230 [PubMed]
- 48. Zhao P, Sui BD, Liu N, Lv YJ, Zheng CX, Lu YB, Huang WT, Zhou CH, Chen J, Pang DL, Fei DD, Xuan K, Hu CH, Jin Y. Anti-aging pharmacology in cutaneous wound healing: effects of metformin, resveratrol, and rapamycin by local application. Aging Cell. 2017; 16:1083–93. https://doi.org/10.1111/acel.12635 [PubMed]
- 49. Martin-Montalvo A, Mercken EM, Mitchell SJ, Palacios HH, Mote PL, Scheibye-Knudsen M, Gomes AP, Ward TM, Minor RK, Blouin MJ, Schwab M, Pollak M, Zhang Y, et al. Metformin improves healthspan and lifespan in mice. Nat Commun. 2013; 4:2192. https://doi.org/10.1038/ncomms3192 [PubMed]
- 50. Slack C, Foley A, Partridge L. Activation of AMPK by the putative dietary restriction mimetic metformin is insufficient to extend lifespan in Drosophila. PLoS One. 2012; 7:e47699. https://doi.org/10.1371/journal.pone.0047699 [PubMed]
- 51. Onken B, Driscoll M. Metformin induces a dietary restriction-like state and the oxidative stress response to extend C. elegans Healthspan via AMPK, LKB1, and SKN-1. PLoS One. 2010; 5:e8758. https://doi.org/10.1371/journal.pone.0008758 [PubMed]
- 52. Anisimov VN, Piskunova TS, Popovich IG, Zabezhinski MA, Tyndyk ML, Egormin PA, Yurova MV, Rosenfeld SV, Semenchenko AV, Kovalenko IG, Poroshina TE, Berstein LM. Gender differences in metformin effect on aging, life span and spontaneous tumorigenesis in 129/Sv mice. Aging (Albany NY). 2010; 2:945–58. https://doi.org/10.18632/aging.100245 [PubMed]
- 53. Anisimov VN, Berstein LM, Popovich IG, Zabezhinski MA, Egormin PA, Piskunova TS, Semenchenko AV, Tyndyk ML, Yurova MN, Kovalenko IG, Poroshina TE. If started early in life, metformin treatment increases life span and postpones tumors in female SHR mice. Aging (Albany NY). 2011; 3:148–57. https://doi.org/10.18632/aging.100273 [PubMed]
- 54. Batandier C, Guigas B, Detaille D, El-Mir MY, Fontaine E, Rigoulet M, Leverve XM. The ROS production induced by a reverse-electron flux at respiratory-chain complex 1 is hampered by metformin. J Bioenerg Biomembr. 2006; 38:33–42. https://doi.org/10.1007/s10863-006-9003-8 [PubMed]
- 55. Hirsch A, Hahn D, Kempná P, Hofer G, Nuoffer JM, Mullis PE, Flück CE. Metformin inhibits human androgen production by regulating steroidogenic enzymes HSD3B2 and CYP17A1 and complex I activity of the respiratory chain. Endocrinology. 2012; 153:4354–66. https://doi.org/10.1210/en.2012-1145 [PubMed]
- 56. Owen MR, Doran E, Halestrap AP. Evidence that metformin exerts its anti-diabetic effects through inhibition of complex 1 of the mitochondrial respiratory chain. Biochem J. 2000; 348:607–14. https://doi.org/10.1042/bj3480607 [PubMed]
- 57. Fontaine E. Metformin and respiratory chain complex I: the last piece of the puzzle? Biochem J. 2014; 463:e3–5. https://doi.org/10.1042/BJ20141020 [PubMed]
- 58. Bridges HR, Jones AJ, Pollak MN, Hirst J. Effects of metformin and other biguanides on oxidative phosphorylation in mitochondria. Biochem J. 2014; 462:475–87. https://doi.org/10.1042/BJ20140620 [PubMed]
- 59. Zheng Z, Chen H, Li J, Li T, Zheng B, Zheng Y, Jin H, He Y, Gu Q, Xu X. Sirtuin 1-mediated cellular metabolic memory of high glucose via the LKB1/AMPK/ROS pathway and therapeutic effects of metformin. Diabetes. 2012; 61:217–28. https://doi.org/10.2337/db11-0416 [PubMed]
- 60. Owen MR, Doran E, Halestrap AP. Evidence that metformin exerts its anti-diabetic effects through inhibition of complex 1 of the mitochondrial respiratory chain. Biochem J. 2000; 348:607–14. https://doi.org/10.1042/bj3480607 [PubMed]
- 61. El-Mir MY, Nogueira V, Fontaine E, Avéret N, Rigoulet M, Leverve X. Dimethylbiguanide inhibits cell respiration via an indirect effect targeted on the respiratory chain complex I. J Biol Chem. 2000; 275:223–28. https://doi.org/10.1074/jbc.275.1.223 [PubMed]
- 62. Stephenne X, Foretz M, Taleux N, van der Zon GC, Sokal E, Hue L, Viollet B, Guigas B. Metformin activates AMP-activated protein kinase in primary human hepatocytes by decreasing cellular energy status. Diabetologia. 2011; 54:3101–10. https://doi.org/10.1007/s00125-011-2311-5 [PubMed]
- 63. Lien F, Berthier A, Bouchaert E, Gheeraert C, Alexandre J, Porez G, Prawitt J, Dehondt H, Ploton M, Colin S, Lucas A, Patrice A, Pattou F, et al. Metformin interferes with bile acid homeostasis through AMPK-FXR crosstalk. J Clin Invest. 2014; 124:1037–51. https://doi.org/10.1172/JCI68815 [PubMed]
- 64. Lu J, Shi J, Li M, Gui B, Fu R, Yao G, Duan Z, Lv Z, Yang Y, Chen Z, Jia L, Tian L. Activation of AMPK by metformin inhibits TGF-β-induced collagen production in mouse renal fibroblasts. Life Sci. 2015; 127:59–65. https://doi.org/10.1016/j.lfs.2015.01.042 [PubMed]
- 65. Foretz M, Hébrard S, Leclerc J, Zarrinpashneh E, Soty M, Mithieux G, Sakamoto K, Andreelli F, Viollet B. Metformin inhibits hepatic gluconeogenesis in mice independently of the LKB1/AMPK pathway via a decrease in hepatic energy state. J Clin Invest. 2010; 120:2355–69. https://doi.org/10.1172/JCI40671 [PubMed]
- 66. Cho K, Chung JY, Cho SK, Shin HW, Jang IJ, Park JW, Yu KS, Cho JY. Antihyperglycemic mechanism of metformin occurs via the AMPK/LXRα/POMC pathway. Sci Rep. 2015; 5:8145. https://doi.org/10.1038/srep08145 [PubMed]
- 67. Hardie DG. AMPK: a key regulator of energy balance in the single cell and the whole organism. Int J Obes. 2008 (Suppl 4); 32:S7–12. https://doi.org/10.1038/ijo.2008.116 [PubMed]
- 68. Zhou G, Myers R, Li Y, Chen Y, Shen X, Fenyk-Melody J, Wu M, Ventre J, Doebber T, Fujii N, Musi N, Hirshman MF, Goodyear LJ, Moller DE. Role of AMP-activated protein kinase in mechanism of metformin action. J Clin Invest. 2001; 108:1167–74. https://doi.org/10.1172/JCI13505 [PubMed]
- 69. Liu B, Fan Z, Edgerton SM, Yang X, Lind SE, Thor AD. Potent anti-proliferative effects of metformin on trastuzumab-resistant breast cancer cells via inhibition of erbB2/IGF-1 receptor interactions. Cell Cycle. 2011; 10:2959–66. https://doi.org/10.4161/cc.10.17.16359 [PubMed]
- 70. Zhang E, Guo Q, Gao H, Xu R, Teng S, Wu Y. Metformin and Resveratrol Inhibited High Glucose-Induced Metabolic Memory of Endothelial Senescence through SIRT1/p300/p53/p21 Pathway. PLoS One. 2015; 10:e0143814. https://doi.org/10.1371/journal.pone.0143814 [PubMed]
- 71. Arunachalam G, Samuel SM, Marei I, Ding H, Triggle CR. Metformin modulates hyperglycaemia-induced endothelial senescence and apoptosis through SIRT1. Br J Pharmacol. 2014; 171:523–35. https://doi.org/10.1111/bph.12496 [PubMed]
- 72. Bashmakov YK, Petyaev IM. Old Drug Acquires New Target: metformin and Sirt1. J Diabetes Metab. 2011; 2:107e. https://doi.org/10.4172/2155-6156.1000107e
- 73. Chen Q, Yang X, Zhang H, Kong X, Yao L, Cui X, Zou Y, Fang F, Yang J, Chang Y. Metformin impairs systemic bile acid homeostasis through regulating SIRT1 protein levels. Biochim Biophys Acta. 2017; 1864:101–12. https://doi.org/10.1016/j.bbamcr.2016.10.020 [PubMed]
- 74. Howell JJ, Hellberg K, Turner M, Talbott G, Kolar MJ, Ross DS, Hoxhaj G, Saghatelian A, Shaw RJ, Manning BD. Metformin Inhibits Hepatic mTORC1 Signaling via Dose-Dependent Mechanisms Involving AMPK and the TSC Complex. Cell Metab. 2017; 25:463–71. https://doi.org/10.1016/j.cmet.2016.12.009 [PubMed]
- 75. Chen B, Ma L, Paik H, Sirota M, Wei W, Chua MS, So S, Butte AJ. Reversal of cancer gene expression correlates with drug efficacy and reveals therapeutic targets. Nat Commun. 2017; 8:16022. https://doi.org/10.1038/ncomms16022 [PubMed]
- 76. High KP. The antimicrobial activities of cyclosporine, FK506, and rapamycin. Transplantation. 1994; 57:1689–700. https://doi.org/10.1097/00007890-199457120-00001 [PubMed]
- 77. Büchler M, Caillard S, Barbier S, Thervet E, Toupance O, Mazouz H, Hurault de Ligny B, Le Meur Y, Thierry A, Villemain F, Heng AE, Moulin B, Morin MP, et al, and SPIESSER Group. Sirolimus versus cyclosporine in kidney recipients receiving thymoglobulin, mycophenolate mofetil and a 6-month course of steroids. Am J Transplant. 2007; 7:2522–31. https://doi.org/10.1111/j.1600-6143.2007.01976.x [PubMed]
- 78. Houde VP, Brûlé S, Festuccia WT, Blanchard PG, Bellmann K, Deshaies Y, Marette A. Chronic rapamycin treatment causes glucose intolerance and hyperlipidemia by upregulating hepatic gluconeogenesis and impairing lipid deposition in adipose tissue. Diabetes. 2010; 59:1338–48. https://doi.org/10.2337/db09-1324 [PubMed]
- 79. Chang GR, Wu YY, Chiu YS, Chen WY, Liao JW, Hsu HM, Chao TH, Hung SW, Mao FC. Long-term administration of rapamycin reduces adiposity, but impairs glucose tolerance in high-fat diet-fed KK/HlJ mice. Basic Clin Pharmacol Toxicol. 2009; 105:188–98. https://doi.org/10.1111/j.1742-7843.2009.00427.x [PubMed]
- 80. Schindler CE, Partap U, Patchen BK, Swoap SJ. Chronic rapamycin treatment causes diabetes in male mice. Am J Physiol Regul Integr Comp Physiol. 2014; 307:R434–43. https://doi.org/10.1152/ajpregu.00123.2014 [PubMed]
- 81. Blagosklonny MV. Rapamycin-induced glucose intolerance: hunger or starvation diabetes. Cell Cycle. 2011; 10:4217–24. https://doi.org/10.4161/cc.10.24.18595 [PubMed]
- 82. Dujic T, Causevic A, Bego T, Malenica M, Velija-Asimi Z, Pearson ER, Semiz S. Organic cation transporter 1 variants and gastrointestinal side effects of metformin in patients with Type 2 diabetes. Diabet Med. 2016; 33:511–14. https://doi.org/10.1111/dme.13040 [PubMed]
- 83. Blonde L, Dailey GE, Jabbour SA, Reasner CA, Mills DJ. Gastrointestinal tolerability of extended-release metformin tablets compared to immediate-release metformin tablets: results of a retrospective cohort study. Curr Med Res Opin. 2004; 20:565–72. https://doi.org/10.1185/030079904125003278 [PubMed]
- 84. Lalau JD. Lactic acidosis induced by metformin: incidence, management and prevention. Drug Saf. 2010; 33:727–40. https://doi.org/10.2165/11536790-000000000-00000 [PubMed]
- 85. Kim MJ, Han JY, Shin JY, Kim SI, Lee JM, Hong S, Kim SH, Nam MS, Kim YS. Metformin-associated lactic acidosis: predisposing factors and outcome. Endocrinol Metab (Seoul). 2015; 30:78–83. https://doi.org/10.3803/EnM.2015.30.1.78 [PubMed]
- 86. Biradar V, Moran JL, Peake SL, Peter JV. Metformin-associated lactic acidosis (MALA): clinical profile and outcomes in patients admitted to the intensive care unit. Crit Care Resusc. 2010; 12:191–95. [PubMed]
- 87. Leontieva OV, Blagosklonny MV. M(o)TOR of pseudo-hypoxic state in aging: rapamycin to the rescue. Cell Cycle. 2014; 13:509–15. https://doi.org/10.4161/cc.27973 [PubMed]
- 88. Nasri H, Baradaran A, Shirzad H, Rafieian-Kopaei M. New concepts in nutraceuticals as alternative for pharmaceuticals. Int J Prev Med. 2014; 5:1487–99. [PubMed]
- 89. Zhu Q, Izumchenko E, Aliper AM, Makarev E, Paz K, Buzdin AA, Zhavoronkov AA, Sidransky D. Pathway activation strength is a novel independent prognostic biomarker for cetuximab sensitivity in colorectal cancer patients. Hum Genome Var. 2015; 2:15009. https://doi.org/10.1038/hgv.2015.9 [PubMed]
- 90. Borisov NM, Terekhanova NV, Aliper AM, Venkova LS, Smirnov PY, Roumiantsev S, Korzinkin MB, Zhavoronkov AA, Buzdin AA. Signaling pathways activation profiles make better markers of cancer than expression of individual genes. Oncotarget. 2014; 5:10198–205. https://doi.org/10.18632/oncotarget.2548 [PubMed]
- 91. Lezhnina K, Kovalchuk O, Zhavoronkov AA, Korzinkin MB, Zabolotneva AA, Shegay PV, Sokov DG, Gaifullin NM, Rusakov IG, Aliper AM, Roumiantsev SA, Alekseev BY, Borisov NM, Buzdin AA. Novel robust biomarkers for human bladder cancer based on activation of intracellular signaling pathways. Oncotarget. 2014; 5:9022–32. https://doi.org/10.18632/oncotarget.2493 [PubMed]
- 92. Artemov A, Aliper A, Korzinkin M, Lezhnina K, Jellen L, Zhukov N, Roumiantsev S, Gaifullin N, Zhavoronkov A, Borisov N, Buzdin A. A method for predicting target drug efficiency in cancer based on the analysis of signaling pathway activation. Oncotarget. 2015; 6:29347–56. https://doi.org/10.18632/oncotarget.5119 [PubMed]
- 93. Aliper A, Belikov AV, Garazha A, Jellen L, Artemov A, Suntsova M, Ivanova A, Venkova L, Borisov N, Buzdin A, Mamoshina P, Putin E, Swick AG, et al. In search for geroprotectors: in silico screening and in vitro validation of signalome-level mimetics of young healthy state. Aging (Albany NY). 2016; 8:2127–52. https://doi.org/10.18632/aging.101047 [PubMed]
- 94. Wei T, Li S. Development of Genomics-Based Gene Expression Signature Biomarkers in Oncology and Toxicology to Facilitate Drug Discovery and Translational Medicine. Curr Bioinform. 2010; 5:109–17. https://doi.org/10.2174/157489310791268423
- 95. Sithara S, Crowley TM, Walder K, Aston-Mourney K. Gene expression signature: a powerful approach for drug discovery in diabetes. J Endocrinol. 2017; 232:R131–39. https://doi.org/10.1530/JOE-16-0515 [PubMed]
- 96. Luizon MR, Eckalbar WL, Wang Y, Jones SL, Smith RP, Laurance M, Lin L, Gallins PJ, Etheridge AS, Wright F, Zhou Y, Molony C, Innocenti F, et al. Genomic Characterization of Metformin Hepatic Response. PLoS Genet. 2016; 12:e1006449. https://doi.org/10.1371/journal.pgen.1006449 [PubMed]
- 97. Leff T. AMP-activated protein kinase regulates gene expression by direct phosphorylation of nuclear proteins. Biochem Soc Trans. 2003; 31:224–27. https://doi.org/10.1042/bst0310224 [PubMed]
- 98. Spindler SR. Use of microarray biomarkers to identify longevity therapeutics. Aging Cell. 2006; 5:39–50. https://doi.org/10.1111/j.1474-9726.2006.00194.x [PubMed]
- 99. Dhahbi JM, Mote PL, Fahy GM, Spindler SR. Identification of potential caloric restriction mimetics by microarray profiling. Physiol Genomics. 2005; 23:343–50. https://doi.org/10.1152/physiolgenomics.00069.2005 [PubMed]
- 100. Cao SX, Dhahbi JM, Mote PL, Spindler SR. Genomic profiling of short- and long-term caloric restriction effects in the liver of aging mice. Proc Natl Acad Sci USA. 2001; 98:10630–35. https://doi.org/10.1073/pnas.191313598 [PubMed]
- 101. Spindler SR. Rapid and reversible induction of the longevity, anticancer and genomic effects of caloric restriction. Mech Ageing Dev. 2005; 126:960–66. https://doi.org/10.1016/j.mad.2005.03.016 [PubMed]
- 102. Aliper A, Plis S, Artemov A, Ulloa A, Mamoshina P, Zhavoronkov A. Deep Learning Applications for Predicting Pharmacological Properties of Drugs and Drug Repurposing Using Transcriptomic Data. Mol Pharm. 2016; 13:2524–30. https://doi.org/10.1021/acs.molpharmaceut.6b00248 [PubMed]
- 103. Buzdin AA, Zhavoronkov AA, Korzinkin MB, Venkova LS, Zenin AA, Smirnov PY, Borisov NM. Oncofinder, a new method for the analysis of intracellular signaling pathway activation using transcriptomic data. Front Genet. 2014; 5:55. https://doi.org/10.3389/fgene.2014.00055 [PubMed]
- 104. Ozerov IV, Lezhnina KV, Izumchenko E, Artemov AV, Medintsev S, Vanhaelen Q, Aliper A, Vijg J, Osipov AN, Labat I, West MD, Buzdin A, Cantor CR, et al. In silico Pathway Activation Network Decomposition Analysis (iPANDA) as a method for biomarker development. Nat Commun. 2016; 7:13427. https://doi.org/10.1038/ncomms13427 [PubMed]
- 105. Gu J, Gui Y, Chen L, Yuan G, Lu HZ, Xu X. Use of natural products as chemical library for drug discovery and network pharmacology. PLoS One. 2013; 8:e62839. https://doi.org/10.1371/journal.pone.0062839 [PubMed]
- 106. Kanehisa M, Sato Y, Kawashima M, Furumichi M, Tanabe M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 2016; 44:D457–62. https://doi.org/10.1093/nar/gkv1070 [PubMed]
- 107. Lin KC, Yeh LR, Chen LJ, Wen YJ, Cheng KC, Cheng JT. Plasma glucose-lowering action of allantoin is induced by activation of imidazoline I-2 receptors in streptozotocin-induced diabetic rats. Horm Metab Res. 2012; 44:41–46. https://doi.org/10.1055/s-0031-1295439 [PubMed]
- 108. Niu CS, Chen W, Wu HT, Cheng KC, Wen YJ, Lin KC, Cheng JT. Decrease of plasma glucose by allantoin, an active principle of yam ( Dioscorea spp.), in streptozotocin-induced diabetic rats. J Agric Food Chem. 2010; 58:12031–35. https://doi.org/10.1021/jf103234d [PubMed]
- 109. Vogels GD, Van der Drift C. Degradation of purines and pyrimidines by microorganisms. Bacteriol Rev. 1976; 40:403–68. [PubMed]
- 110. Kahn K, Serfozo P, Tipton PA. Identification of the True Product of the Urate Oxidase Reaction. J Am Chem Soc. 1997; 119:5435–42. https://doi.org/10.1021/ja970375t
- 111. Malir F, Ostry V, Pfohl-Leszkowicz A, Malir J, Toman J. Ochratoxin A: 50 Years of Research. Toxins (Basel). 2016; 8:191. https://doi.org/10.3390/toxins8070191 [PubMed]
- 112. Zeng C, Wei J, Li H, Wang YL, Xie DX, Yang T, Gao SG, Li YS, Luo W, Lei GH. Effectiveness and safety of Glucosamine, chondroitin, the two in combination, or celecoxib in the treatment of osteoarthritis of the knee. Sci Rep. 2015; 5:16827. https://doi.org/10.1038/srep16827 [PubMed]
- 113. Yadav VR, Prasad S, Sung B, Aggarwal BB. The role of chalcones in suppression of NF-κB-mediated inflammation and cancer. Int Immunopharmacol. 2011; 11:295–309. https://doi.org/10.1016/j.intimp.2010.12.006 [PubMed]
- 114. Niu PG, Zhang YX, Shi DH, Liu Y, Chen YY, Deng J. Cardamonin Inhibits Metastasis of Lewis Lung Carcinoma Cells by Decreasing mTOR Activity. PLoS One. 2015; 10:e0127778. https://doi.org/10.1371/journal.pone.0127778 [PubMed]
- 115. Ohji G, Hidayat S, Nakashima A, Tokunaga C, Oshiro N, Yoshino K, Yokono K, Kikkawa U, Yonezawa K. Suppression of the mTOR-raptor signaling pathway by the inhibitor of heat shock protein 90 geldanamycin. J Biochem. 2006; 139:129–35. https://doi.org/10.1093/jb/mvj008 [PubMed]
- 116. Gianni L, Norton L, Wolmark N, Suter TM, Bonadonna G, Hortobagyi GN. Role of anthracyclines in the treatment of early breast cancer. J Clin Oncol. 2009; 27:4798–808. https://doi.org/10.1200/JCO.2008.21.4791 [PubMed]
- 117. Flavin R, Peluso S, Nguyen PL, Loda M. Fatty acid synthase as a potential therapeutic target in cancer. Future Oncol. 2010; 6:551–62. https://doi.org/10.2217/fon.10.11 [PubMed]
- 118. Cherblanc FL, Chapman KL, Brown R, Fuchter MJ. Chaetocin is a nonspecific inhibitor of histone lysine methyltransferases. Nat Chem Biol. 2013; 9:136–37. https://doi.org/10.1038/nchembio.1187 [PubMed]
- 119. Lee YM, Lim JH, Yoon H, Chun YS, Park JW. Antihepatoma activity of chaetocin due to deregulated splicing of hypoxia-inducible factor 1α pre-mRNA in mice and in vitro. Hepatology. 2011; 53:171–80. https://doi.org/10.1002/hep.24010 [PubMed]
- 120. Min J, Huang K, Tang H, Ding X, Qi C, Qin X, Xu Z. Phloretin induces apoptosis of non-small cell lung carcinoma A549 cells via JNK1/2 and p38 MAPK pathways. Oncol Rep. 2015; 34:2871–79. https://doi.org/10.3892/or.2015.4325 [PubMed]
- 121. Lee IC, Choi BY. Withaferin-A--A Natural Anticancer Agent with Pleitropic Mechanisms of Action. Int J Mol Sci. 2016; 17:290. https://doi.org/10.3390/ijms17030290 [PubMed]
- 122. Mazimba O. Umbelliferone: Sources, chemistry and bioactivities review. Bull Fac Pharm Cairo Univ. 2017; •••. https://doi.org/10.1016/j.bfopcu.2017.05.001
- 123. Amalan V, Vijayakumar N, Indumathi D, Ramakrishnan A. Antidiabetic and antihyperlipidemic activity of p-coumaric acid in diabetic rats, role of pancreatic GLUT 2: in vivo approach. Biomed Pharmacother. 2016; 84:230–36. https://doi.org/10.1016/j.biopha.2016.09.039 [PubMed]
- 124. Pragasam SJ, Venkatesan V, Rasool M. Immunomodulatory and anti-inflammatory effect of p-coumaric acid, a common dietary polyphenol on experimental inflammation in rats. Inflammation. 2013; 36:169–76. https://doi.org/10.1007/s10753-012-9532-8 [PubMed]
- 125. Park BS, Abdel-Azeem AZ, Al-Sanea MM, Yoo KH, Tae JS, Lee SH. Staurosporine analogues from microbial and synthetic sources and their biological activities. Curr Med Chem. 2013; 20:3872–902. https://doi.org/10.2174/09298673113209990176 [PubMed]
- 126. Phelan JP, Reen FJ, Dunphy N, O’Connor R, O’Gara F. Bile acids destabilise HIF-1α and promote anti-tumour phenotypes in cancer cell models. BMC Cancer. 2016; 16:476. https://doi.org/10.1186/s12885-016-2528-2 [PubMed]
- 127. Garbett NC, Graves DE. Extending nature’s leads: the anticancer agent ellipticine. Curr Med Chem Anticancer Agents. 2004; 4:149–72. https://doi.org/10.2174/1568011043482070 [PubMed]
- 128. Nagle DG, Ferreira D, Zhou YD. Epigallocatechin-3-gallate (EGCG): chemical and biomedical perspectives. Phytochemistry. 2006; 67:1849–55. https://doi.org/10.1016/j.phytochem.2006.06.020 [PubMed]
- 129. Chew WS, Wang W, Herr DR. To fingolimod and beyond: the rich pipeline of drug candidates that target S1P signaling. Pharmacol Res. 2016; 113:521–32. https://doi.org/10.1016/j.phrs.2016.09.025 [PubMed]
- 130. Peng F, Du Q, Peng C, Wang N, Tang H, Xie X, Shen J, Chen J. A Review: The Pharmacology of Isoliquiritigenin. Phytother Res. 2015; 29:969–77. https://doi.org/10.1002/ptr.5348 [PubMed]
- 131. Pajak B. Antiapoptotic proteins as targets for bioactive compounds. Pol J Vet Sci. 2007; 10:127–30. [PubMed]
- 132. Peggy Hsu P. Natural Medicines Comprehensive Database, J Med Libr Assoc. Medical Library Association. 2002; 90:114.
- 133. Lehár J, Krueger AS, Avery W, Heilbut AM, Johansen LM, Price ER, Rickles RJ, Short GF
3rd , Staunton JE, Jin X, Lee MS, Zimmermann GR, Borisy AA. Synergistic drug combinations tend to improve therapeutically relevant selectivity. Nat Biotechnol. 2009; 27:659–66. https://doi.org/10.1038/nbt.1549 [PubMed] - 134. Tan X, Hu L, Luquette LJ
3rd , Gao G, Liu Y, Qu H, Xi R, Lu ZJ, Park PJ, Elledge SJ. Systematic identification of synergistic drug pairs targeting HIV. Nat Biotechnol. 2012; 30:1125–30. https://doi.org/10.1038/nbt.2391 [PubMed] - 135. Katouli AA, Komarova NL. Optimizing combination therapies with existing and future CML drugs. PLoS One. 2010; 5:e12300. https://doi.org/10.1371/journal.pone.0012300 [PubMed]
- 136. Liu L, Li H, Guo Z, Ma X, Cao N, Zheng Y, Geng S, Duan Y, Han G, Du G. The Combination of Three Natural Compounds Effectively Prevented Lung Carcinogenesis by Optimal Wound Healing. PLoS One. 2015; 10:e0143438. https://doi.org/10.1371/journal.pone.0143438 [PubMed]
- 137. Bulusu KC, Guha R, Mason DJ, Lewis RP, Muratov E, Kalantar Motamedi Y, Cokol M, Bender A. Modelling of compound combination effects and applications to efficacy and toxicity: state-of-the-art, challenges and perspectives. Drug Discov Today. 2016; 21:225–38. https://doi.org/10.1016/j.drudis.2015.09.003 [PubMed]
- 138. Dessale T, Batchu KC, Barardo D, Ng LF, Lam VY, Xiao L, Wenk MR, Tolwinski NS, Gruber J. Doubling healthy lifespan using drug synergies. bioRxiv. 2017; 153205. https://doi.org/10.1101/153205
- 139. Zanetti A, Affatato R, Centritto F, Fratelli M, Kurosaki M, Barzago MM, Bolis M, Terao M, Garattini E, Paroni G. All-trans-retinoic Acid Modulates the Plasticity and Inhibits the Motility of Breast Cancer Cells: ROLE OF NOTCH1 AND TRANSFORMING GROWTH FACTOR (TGFβ). J Biol Chem. 2015; 290:17690–709. https://doi.org/10.1074/jbc.M115.638510 [PubMed]
- 140. Lee J, Liu J, Feng X, Salazar Hernández MA, Mucka P, Ibi D, Choi JW, Ozcan U. Withaferin A is a leptin sensitizer with strong antidiabetic properties in mice. Nat Med. 2016; 22:1023–32. https://doi.org/10.1038/nm.4145 [PubMed]
- 141. Hahm ER, Lee J, Kim SH, Sehrawat A, Arlotti JA, Shiva SS, Bhargava R, Singh SV. Metabolic alterations in mammary cancer prevention by withaferin A in a clinically relevant mouse model. J Natl Cancer Inst. 2013; 105:1111–22. https://doi.org/10.1093/jnci/djt153 [PubMed]
- 142. Mohan R, Hammers HJ, Bargagna-Mohan P, Zhan XH, Herbstritt CJ, Ruiz A, Zhang L, Hanson AD, Conner BP, Rougas J, Pribluda VS. Withaferin A is a potent inhibitor of angiogenesis. Angiogenesis. 2004; 7:115–22. https://doi.org/10.1007/s10456-004-1026-3 [PubMed]
- 143. Nishikawa Y, Okuzaki D, Fukushima K, Mukai S, Ohno S, Ozaki Y, Yabuta N, Nojima H, Withaferin A, Withaferin A. Withaferin A Induces Cell Death Selectively in Androgen-Independent Prostate Cancer Cells but Not in Normal Fibroblast Cells. PLoS One. 2015; 10:e0134137. https://doi.org/10.1371/journal.pone.0134137 [PubMed]
- 144. Issa ME, Cuendet M. Withaferin A induces cell death and differentiation in multiple myeloma cancer stem cells. MedChemComm. 2017; 8:112–21. https://doi.org/10.1039/C6MD00410E
- 145. Koduru S, Kumar R, Srinivasan S, Evers MB, Damodaran C. Notch-1 inhibition by Withaferin-A: a therapeutic target against colon carcinogenesis. Mol Cancer Ther. 2010; 9:202–10. https://doi.org/10.1158/1535-7163.MCT-09-0771 [PubMed]
- 146. Khazal KF, Samuel T, Hill DL, Grubbs CJ. Effect of an extract of Withania somnifera root on estrogen receptor-positive mammary carcinomas. Anticancer Res. 2013; 33:1519–23. [PubMed]
- 147. Choi BY, Kim BW. Withaferin-A Inhibits Colon Cancer Cell Growth by Blocking STAT3 Transcriptional Activity. J Cancer Prev. 2015; 20:185–92. https://doi.org/10.15430/JCP.2015.20.3.185 [PubMed]
- 148. Tahara T, Streit U, Pelish HE, Shair MD. STAT3 Inhibitory Activity of Structurally Simplified Withaferin A Analogues. Org Lett. 2017; 19:1538–41. https://doi.org/10.1021/acs.orglett.7b00332 [PubMed]
- 149. Lee HE, Shin JA, Jeong JH, Jeon JG, Lee MH, Cho SD. Anticancer activity of Ashwagandha against human head and neck cancer cell lines. J Oral Pathol Med. 2016; 45:193–201. https://doi.org/10.1111/jop.12353 [PubMed]
- 150. Um HJ, Min KJ, Kim DE, Kwon TK. Withaferin A inhibits JAK/STAT3 signaling and induces apoptosis of human renal carcinoma Caki cells. Biochem Biophys Res Commun. 2012; 427:24–29. https://doi.org/10.1016/j.bbrc.2012.08.133 [PubMed]
- 151. Yu Y, Hamza A, Zhang T, Gu M, Zou P, Newman B, Li Y, Gunatilaka AA, Zhan CG, Sun D. Withaferin A targets heat shock protein 90 in pancreatic cancer cells. Biochem Pharmacol. 2010; 79:542–51. https://doi.org/10.1016/j.bcp.2009.09.017 [PubMed]
- 152. Yan X, Huang G, Liu Q, Zheng J, Chen H, Huang Q, Chen J, Huang H. Withaferin A protects against spinal cord injury by inhibiting apoptosis and inflammation in mice. Pharm Biol. 2017; 55:1171–76. https://doi.org/10.1080/13880209.2017.1288262 [PubMed]
- 153. Christensen LP. Ginsenosides chemistry, biosynthesis, analysis, and potential health effects. Adv Food Nutr Res. 2009; 55:1–99. [PubMed]
- 154. Lim SW, Doh KC, Jin L, Jin J, Piao SG, Heo SB, Chung BH, Yang CW. Ginseng treatment attenuates autophagic cell death in chronic cyclosporine nephropathy. Nephrology (Carlton). 2014; 19:490–99. https://doi.org/10.1111/nep.12273 [PubMed]
- 155. Yoo HS, Kim JM, Jo E, Cho CK, Lee SY, Kang HS, Lee MG, Yang PY, Jang IS. Modified Panax ginseng extract regulates autophagy by AMPK signaling in A549 human lung cancer cells. Oncol Rep. 2017; 37:3287–96. https://doi.org/10.3892/or.2017.5590 [PubMed]
- 156. Kim MJ, Yun H, Kim DH, Kang I, Choe W, Kim SS, Ha J. AMP-activated protein kinase determines apoptotic sensitivity of cancer cells to ginsenoside-Rh2. J Ginseng Res. 2014; 38:16–21. https://doi.org/10.1016/j.jgr.2013.11.010 [PubMed]
- 157. Chen G, Li H, Gao Y, Zhang L, Zhao Y. Flavored black ginseng exhibited antitumor activity via improving immune function and inducing apoptosis. Food Funct. 2017; 8:1880–89. https://doi.org/10.1039/C6FO01870J [PubMed]
- 158. Immunomodulatory Effects of Non-saponin Red Ginseng Components on Innate Immune Cells. J Ginseng Res. 2008; 32:67–72. https://doi.org/10.5142/JGR.2008.32.1.067
- 159. Kang S, Min H. Ginseng, the ‘Immunity Boost’: The Effects of Panax ginseng on Immune System. J Ginseng Res. 2012; 36:354–68. https://doi.org/10.5142/jgr.2012.36.4.354 [PubMed]
- 160. Larsen MW, Moser C, Høiby N, Song Z, Kharazmi A. Ginseng modulates the immune response by induction of interleukin-12 production. APMIS. 2004; 112:369–73. https://doi.org/10.1111/j.1600-0463.2004.apm1120607.x [PubMed]
- 161. Hafez MM, Hamed SS, El-Khadragy MF, Hassan ZK, Al Rejaie SS, Sayed-Ahmed MM, Al-Harbi NO, Al-Hosaini KA, Al-Harbi MM, Alhoshani AR, Al-Shabanah OA, Alsharari SD. Effect of ginseng extract on the TGF-β1 signaling pathway in CCl4-induced liver fibrosis in rats. BMC Complement Altern Med. 2017; 17:45. https://doi.org/10.1186/s12906-016-1507-0 [PubMed]
- 162. Lee YY, Park JS, Lee EJ, Lee SY, Kim DH, Kang JL, Kim HS. Anti-inflammatory mechanism of ginseng saponin metabolite Rh3 in lipopolysaccharide-stimulated microglia: critical role of 5′-adenosine monophosphate-activated protein kinase signaling pathway. J Agric Food Chem. 2015; 63:3472–80. https://doi.org/10.1021/jf506110y [PubMed]
- 163. Inoue K. Korean red ginseng for allergic rhinitis. Immunopharmacol Immunotoxicol. 2013; 35:693. https://doi.org/10.3109/08923973.2013.838254 [PubMed]
- 164. Lee EJ, Song MJ, Kwon HS, Ji GE, Sung MK. Oral administration of fermented red ginseng suppressed ovalbumin-induced allergic responses in female BALB/c mice. Phytomedicine. 2012; 19:896–903. https://doi.org/10.1016/j.phymed.2012.04.008 [PubMed]
- 165. Li LC, Piao HM, Zheng MY, Lin ZH, Choi YH, Yan GH. Ginsenoside Rh2 attenuates allergic airway inflammation by modulating nuclear factor-κB activation in a murine model of asthma. Mol Med Rep. 2015; 12:6946–54. https://doi.org/10.3892/mmr.2015.4272 [PubMed]
- 166. Chan GH, Law BY, Chu JM, Yue KK, Jiang ZH, Lau CW, Huang Y, Chan SW, Ying-Kit Yue P, Wong RN. Ginseng extracts restore high-glucose induced vascular dysfunctions by altering triglyceride metabolism and downregulation of atherosclerosis-related genes. Evid Based Complement Alternat Med. 2013; 2013:797310. https://doi.org/10.1155/2013/797310 [PubMed]
- 167. Hong SY, Kim JY, Ahn HY, Shin JH, Kwon O. Panax ginseng extract rich in ginsenoside protopanaxatriol attenuates blood pressure elevation in spontaneously hypertensive rats by affecting the Akt-dependent phosphorylation of endothelial nitric oxide synthase. J Agric Food Chem. 2012; 60:3086–91. https://doi.org/10.1021/jf204447y [PubMed]
- 168. Wang Y, Dong J, Liu P, Lau CW, Gao Z, Zhou D, Tang J, Ng CF, Huang Y. Ginsenoside Rb3 attenuates oxidative stress and preserves endothelial function in renal arteries from hypertensive rats. Br J Pharmacol. 2014; 171:3171–81. https://doi.org/10.1111/bph.12660 [PubMed]
- 169. Seo YS, Shon MY, Kong R, Kang OH, Zhou T, Kim DY, Kwon DY. Black ginseng extract exerts anti-hyperglycemic effect via modulation of glucose metabolism in liver and muscle. J Ethnopharmacol. 2016; 190:231–40. https://doi.org/10.1016/j.jep.2016.05.060 [PubMed]
- 170. Lee B, Sur B, Cho SG, Yeom M, Shim I, Lee H, Hahm DH. Ginsenoside Rb1 rescues anxiety-like responses in a rat model of post-traumatic stress disorder. J Nat Med. 2016; 70:133–44. https://doi.org/10.1007/s11418-015-0943-3 [PubMed]
- 171. Cha HY, Park JH, Hong JT, Yoo HS, Song S, Hwang BY, Eun JS, Oh KW. Anxiolytic-like effects of ginsenosides on the elevated plus-maze model in mice. Biol Pharm Bull. 2005; 28:1621–25. https://doi.org/10.1248/bpb.28.1621 [PubMed]
- 172. Kim NH, Kim KY, Jeong HJ, Kim HM. Antidepressant-like effect of altered Korean red ginseng in mice. Behav Med. 2011; 37:42–46. https://doi.org/10.1080/08964289.2011.566591 [PubMed]
- 173. Jeong KJ, Kim GW, Chung SH. AMP-activated protein kinase: an emerging target for ginseng. J Ginseng Res. 2014; 38:83–88. https://doi.org/10.1016/j.jgr.2013.11.014 [PubMed]
- 174. Han JY, Lee S, Yang JH, Kim S, Sim J, Kim MG, Jeong TC, Ku SK, Cho IJ, Ki SH. Korean Red Ginseng attenuates ethanol-induced steatosis and oxidative stress via AMPK/Sirt1 activation. J Ginseng Res. 2015; 39:105–15. https://doi.org/10.1016/j.jgr.2014.09.001 [PubMed]
- 175. Go HK, Rahman MM, Kim GB, Na CS, Song CH, Kim JS, Kim SJ, Kang HS. Antidiabetic Effects of Yam (Dioscorea batatas) and Its Active Constituent, Allantoin, in a Rat Model of Streptozotocin-Induced Diabetes. Nutrients. 2015; 7:8532–44. https://doi.org/10.3390/nu7105411 [PubMed]
- 176. Hsu JH, Wu YC, Liu IM, Cheng JT. Dioscorea as the principal herb of Die-Huang-Wan, a widely used herbal mixture in China, for improvement of insulin resistance in fructose-rich chow-fed rats. J Ethnopharmacol. 2007; 112:577–84. https://doi.org/10.1016/j.jep.2007.05.013 [PubMed]
- 177. Liu Y, Li H, Fan Y, Man S, Liu Z, Gao W, Wang T. Antioxidant and Antitumor Activities of the Extracts from Chinese Yam (Dioscorea opposite Thunb.) Flesh and Peel and the Effective Compounds. J Food Sci. 2016; 81:H1553–64. https://doi.org/10.1111/1750-3841.13322 [PubMed]
- 178. Whitesell L, Mimnaugh EG, De Costa B, Myers CE, Neckers LM. Inhibition of heat shock protein HSP90-pp60v-src heteroprotein complex formation by benzoquinone ansamycins: essential role for stress proteins in oncogenic transformation. Proc Natl Acad Sci USA. 1994; 91:8324–28. https://doi.org/10.1073/pnas.91.18.8324 [PubMed]
- 179. Fukuyo Y, Hunt CR, Horikoshi N. Geldanamycin and its anti-cancer activities. Cancer Lett. 2010; 290:24–35. https://doi.org/10.1016/j.canlet.2009.07.010 [PubMed]
- 180. Miyata Y. Hsp90 inhibitor geldanamycin and its derivatives as novel cancer chemotherapeutic agents. Curr Pharm Des. 2005; 11:1131–38. https://doi.org/10.2174/1381612053507585 [PubMed]
- 181. Jurczyszyn A, Zebzda A, Czepiel J, Perucki W, Bazan-Socha S, Cibor D, Owczarek D, Majka M. Geldanamycin and Its Derivatives Inhibit the Growth of Myeloma Cells and Reduce the Expression of the MET Receptor. J Cancer. 2014; 5:480–90. https://doi.org/10.7150/jca.8731 [PubMed]
- 182. Kitson RR, Chang CH, Xiong R, Williams HE, Davis AL, Lewis W, Dehn DL, Siegel D, Roe SM, Prodromou C, Ross D, Moody CJ. Synthesis of 19-substituted geldanamycins with altered conformations and their binding to heat shock protein Hsp90. Nat Chem. 2013; 5:307–14. https://doi.org/10.1038/nchem.1596 [PubMed]
- 183. Khandelwal A, Crowley VM, Blagg BS. Natural Product Inspired N-Terminal Hsp90 Inhibitors: From Bench to Bedside? Med Res Rev. 2016; 36:92–118. https://doi.org/10.1002/med.21351 [PubMed]
- 184. Yin Z, Henry EC, Gasiewicz TA. (-)-Epigallocatechin-3-gallate is a novel Hsp90 inhibitor. Biochemistry. 2009; 48:336–45. https://doi.org/10.1021/bi801637q [PubMed]
- 185. Zhang Z, Li HM, Zhou C, Li Q, Ma L, Zhang Z, Sun Y, Wang L, Zhang X, Zhu B, Hong YS, Wu CZ, Liu H. Non-benzoquinone geldanamycin analogs trigger various forms of death in human breast cancer cells. J Exp Clin Cancer Res. 2016; 35:149. https://doi.org/10.1186/s13046-016-0428-6 [PubMed]
- 186. Kinzel L, Ernst A, Orth M, Albrecht V, Hennel R, Brix N, Frey B, Gaipl US, Zuchtriegel G, Reichel CA, Blutke A, Schilling D, Multhoff G, et al. A novel HSP90 inhibitor with reduced hepatotoxicity synergizes with radiotherapy to induce apoptosis, abrogate clonogenic survival, and improve tumor control in models of colorectal cancer. Oncotarget. 2016; 7:43199–219. https://doi.org/10.18632/oncotarget.9774 [PubMed]
- 187. Liu Y, Fan C, Pu L, Wei C, Jin H, Teng Y, Zhao M, Yu AC, Jiang F, Shu J, Li F, Peng Q, Kong J, et al. Phloretin induces cell cycle arrest and apoptosis of human glioblastoma cells through the generation of reactive oxygen species. J Neurooncol. 2016; 128:217–23. https://doi.org/10.1007/s11060-016-2107-z [PubMed]
- 188. Chen G, Hu X, Zhang W, Xu N, Wang FQ, Jia J, Zhang WF, Sun ZJ, Zhao YF. Mammalian target of rapamycin regulates isoliquiritigenin-induced autophagic and apoptotic cell death in adenoid cystic carcinoma cells. Apoptosis. 2012; 17:90–101. https://doi.org/10.1007/s10495-011-0658-1 [PubMed]
- 189. Espín JC, García-Conesa MT, Tomás-Barberán FA. Nutraceuticals: facts and fiction. Phytochemistry. 2007; 68:2986–3008. https://doi.org/10.1016/j.phytochem.2007.09.014 [PubMed]
- 190. Wu H, Esteve E, Tremaroli V, Khan MT, Caesar R, Mannerås-Holm L, Ståhlman M, Olsson LM, Serino M, Planas-Fèlix M, Xifra G, Mercader JM, Torrents D, et al. Metformin alters the gut microbiome of individuals with treatment-naive type 2 diabetes, contributing to the therapeutic effects of the drug. Nat Med. 2017; 23:850–58. https://doi.org/10.1038/nm.4345 [PubMed]
- 191. Iorio F, Rittman T, Ge H, Menden M, Saez-Rodriguez J. Transcriptional data: a new gateway to drug repositioning? Drug Discov Today. 2013; 18:350–57. https://doi.org/10.1016/j.drudis.2012.07.014 [PubMed]
- 192. Nair V, Hinton GE. Rectified linear units improve restricted boltzmann machines. Proceedings of the 27th international conference on machine learning (ICML-10). 2010;807–14.
- 193. Kingma D, Ba J. Adam: A method for stochastic optimization. arXiv preprint. 2014; arXiv:14126980.
- 194. Srivastava N, Hinton GE, Krizhevsky A, Sutskever I, Salakhutdinov R. Dropout: a simple way to prevent neural networks from overfitting. J Mach Learn Res. 2014; 15:1929–58.


