Introduction
Cerebral ischemic stroke continues to incur high morbidity and mortality rates [1]. Therapeutic protocols have continued to evolve as our understanding of the umbra and salvageable penumbra increases. These advances have been broadly aimed at either 1) resolving the infarct in a timely fashion to limit the continued damage incurred by hypoxic stress and resultant neuronal injury or 2) stimulating neural replacement [2]. To date, the application of molecular understanding of hypoxic neuronal injury has been limited [3, 4]. In previous studies, we have demonstrated that inhibitor of DNA binding/differentiation 2 (Id2) promoted hypoxia/ischemia (H/I) induced neuronal apoptosis [5, 6]. Absence of an Id2 targeted inhibitor, however, has limited its application as a potential therapeutic option to protect brain from ischemic injury. Thus, seeking the upstream regulating molecules of Id2 is of great significance.
Long noncoding RNAs (lncRNAs) are transcripts larger than 200 nucleotides that regulate gene expression at the level of chromatin modification, transcription and post-transcriptional processing [7]. LncRNA H19 (H19), the product of H19 gene, is maternally expressed and paternally imprinted [8], and has been described as an oncogene in various cancers, including bladder, colorectal, gastric and breast cancers [9–12]. Specifically, in the setting of hypoxia, H19 expression level is upregulated in response to increased activity of hypoxia inducible factor (HIF) [13, 14]. Given this relationship, manipulation of hypoxia-HIF-H19 axis has been identified as a promising oncologic target [13, 14]. Recent studies have demonstrated a similar elevated H19 levels following ischemic stroke, however, the exact role of H19 in the ischemic brain remains unclear [15, 16].
Recently, Luo et al. reported that upregulated H19 contributed to bladder cancer cell proliferation by increasing Id2 expression [17]. With bioinformatics analysis, Zhao et al. suggested that H19 regulated Id2 expression through competitive binding of miR-19a/b in acute myelocytic leukemia cells. However, the exact interactive mechanisms between H19, miR-19a/b and Id2 remains to be elucidated [18]. More recently, a new regulation mechanism of miRNA sponging was proposed whereby H19 was thought to function as competing endogenous RNA (ceRNA), thereby modulating the depression of miRNA targets and imposing an additional level of post-transcriptional regulation [19–21]. Taken together, this rationale provided the framework for our investigation into the regulatory mechanisms between H19, miR-19a and Id2. In particular, we sought to investigate the role of the H19-miR-19a-Id2 axis in the process of H/I induced neuronal injury.
Results
H19 level was also elevated in the penumbra area of MCAO/R rats, as well as in OGD neuronal cells
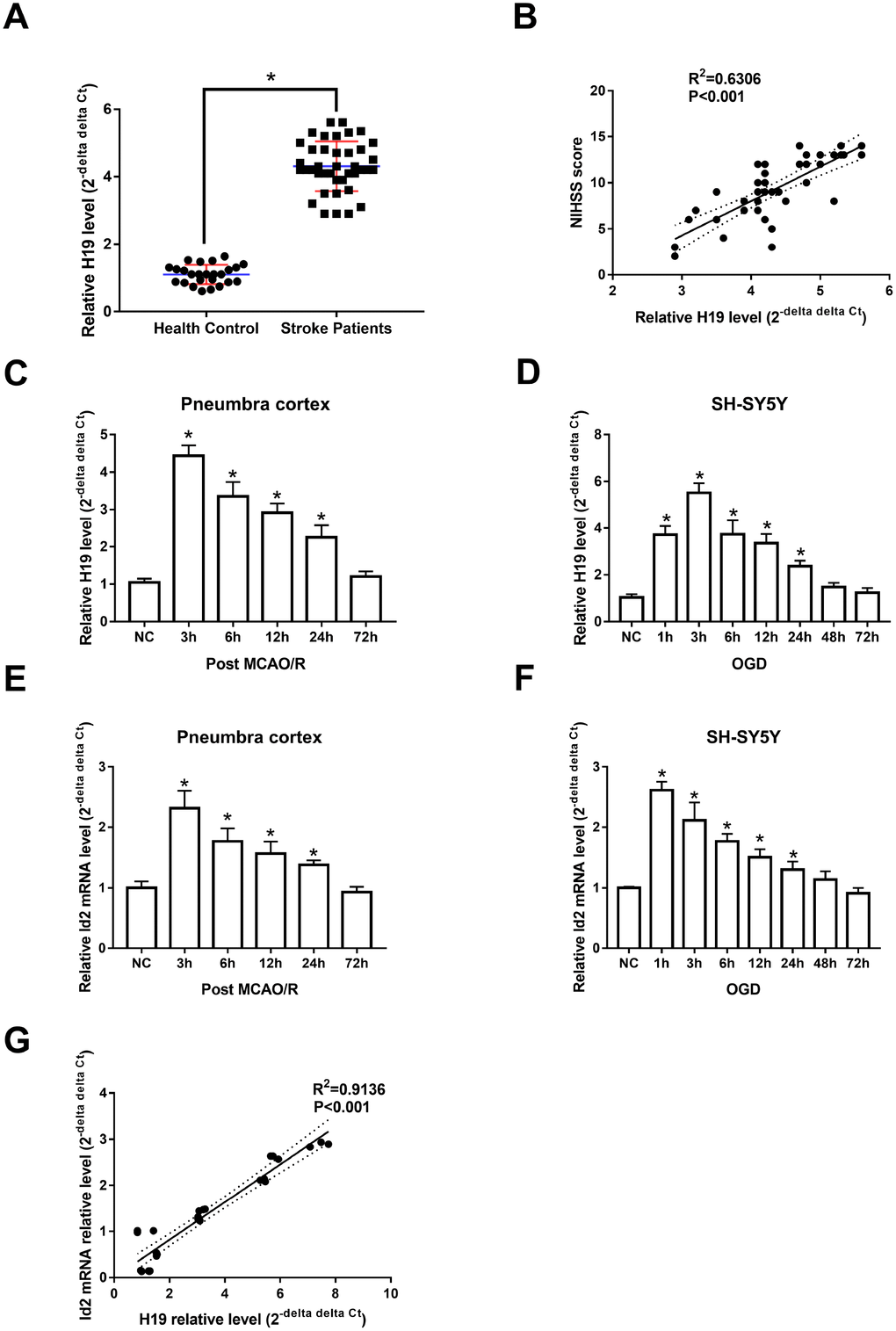
Animal and cell-based models demonstrated similar trends in H19 expression levels as those seen in human subjects. Specifically, H19 levels were significantly elevated in the penumbra cortex of MCAO/R rats and in OGD neuronal cells when compared to controls. H19 levels significantly increased and peaked at 3 h (p<0.05, respectively), and returned to baseline 72 h after exposure of H/I in both in vivo and in vitro experiments (Figure 1C and 1D).
Temporal expression of Id2 mRNA in vivo and in vitro models in the setting of H/I, resembling the tendency of H19 expression
qRT-PCR analysis revealed that Id2 mRNA levels increased significantly and peaked at 3 h after H/I (p<0.05, respectively); however, despite the significant rise, Id2 mRNA levels returned to baseline 72 h after H/I (Figure 1E and 1F). These results suggested that the expression of Id2 mRNA respond to H/I in a specific time dependent manner. Further, since the expression of H19 and Id2 mRNA demonstrated a similar trend, a correlation analysis was completed and the result revealed that Id2 mRNA level was positively correlated with H19 level in OGD neuronal cells (R2=0.9136, p<0.01) (Figure 1G).
Inhibition of H19 decreased Id2 expression and attenuated neuronal apoptosis in OGD neuronal cells
To further investigate the effect of H19 on Id2, three different siRNAs were designed against H19 (named as siRNA-1, siRNA-2, and siRNA-3). All three siRNAs were effective in knocking down H19 expression (p<0.05, respectively), however, siRNA-1 demonstrated the best efficiency and was therefore utilized in subsequent knockdown experiments (Supplementary Figure 1). With knockdown of H19, Id2 mRNA (Figure 2A) and protein (Figure 2B) levels were significantly decreased (p<0.05, respectively). Additionally, flow cytometric analysis and immunofluorescence staining demonstrated a significant decrease in the rate of neuronal apoptosis in the H19 siRNA group, compared with the NC group or mis siRNA group (Figure 2C and 2D; p<0.01, respectively).
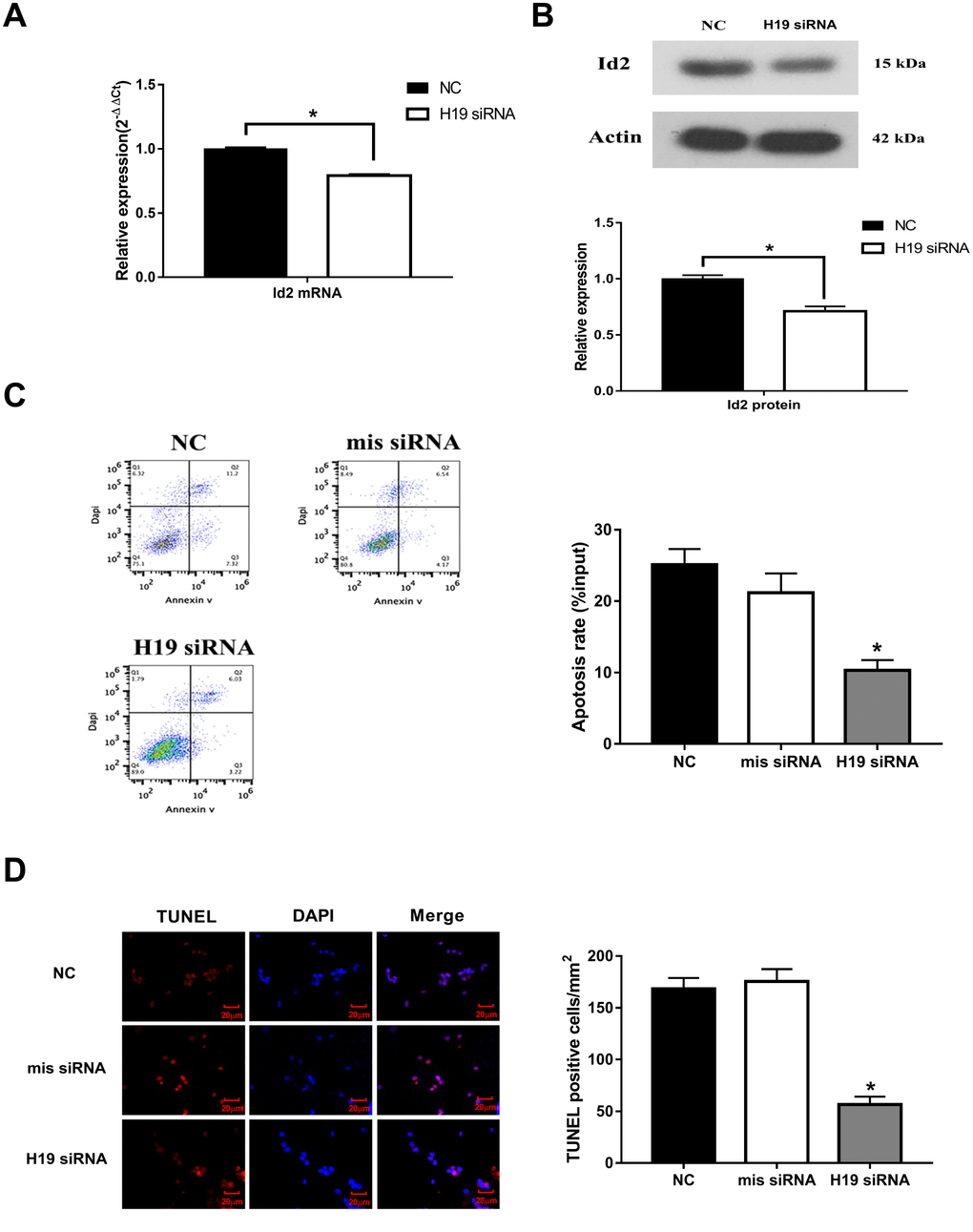
Figure 2. Knockdown of H19 decreased Id2 expression, and attenuated neuronal apoptosis in vitro. (A) After knockdown of H19 with siRNA, Id2 mRNA (A) and protein (B) levels were significantly decreased in OGD neuronal cells. (C) Flow cytometric analysis showed a significant decrease in the rate of neuronal apoptosis in OGD neuronal cells of H19 siRNA group, compared with the NC group or mis siRNA group. (D) Immunofluorescence staining demonstrated a similar decrease rate of neuronal apoptosis in H19 siRNA group.
Inhibition of H19 attenuated neurological deficits and decreased brain infarct volume in MCAO/R rats
Knockdown of H19 by a single intraventricular injection of H19 siRNA offered neuronal protection against ischemic damage following MCA occlusion. Significant reduction in neurological deficit were observed at 3 d and 7 d post MCAO/R when compared to the NS group or mis siRNA group (p<0.01, respectively, Figure 3A). Significant reduction in infarct volumes were also seen in rats injected with H19 siRNA compared with NC group or mis siRNA group (p<0.01, respectively, Figure 3B). Furthermore, to determine whether the lesser extent of ischemic brain injury from H19 knockdown was attributable to the reduced neuronal apoptosis, we assessed the frequency of apoptotic neuronal cells in the penumbra area by TUNEL assay at 72 h post MCAO/R. As shown in Figure 3C, there was a significant decrease in the number of TUNEL-positive cells in the H19 siRNA group compared to NC group or mis siRNA group (p<0.01, respectively).
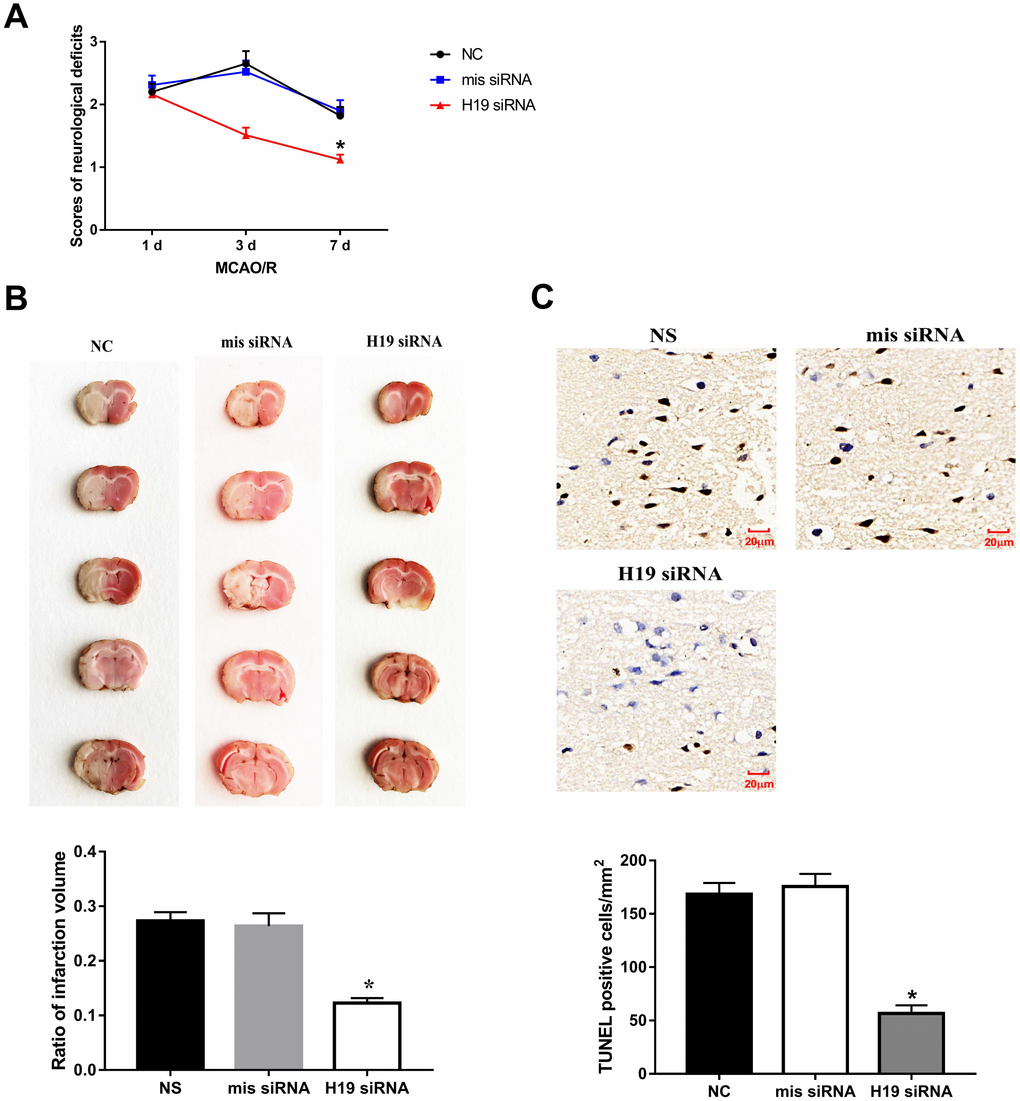
Figure 3. Knockdown of H19 reduced neuronal injury and neuronal apoptosis in vivo. (A) After a single intracerebroventricular injection of H19 siRNA in MCAO/R rats, significant reductions in neurological deficit were observed in H19 siRNA group at 3 d and 7 d post MCAO/R, when compared to the NS group or mis siRNA group. (B) Significant reductions in infarct volumes were also seen in rats of H19 siRNA group, compared with NC group or mis siRNA group. (C) There was a significant decrease in the number of TUNEL-positive cells in the H19 siRNA group compared to NC group or mis siRNA group.
H19 directly interacted with miR-19a
To assess if H19 directly interacted with miR-19a, we first determined the levels of miR-19a in the setting of H/I. qRT-PCR analysis revealed that miR-19a levels increased significantly within the first 72 h after H/I in both MCAO/R and OGD models (p<0.05, respectively) (Figure 4A and 4B). Next, we assessed how miR-19a level varied with knockdown of H19 by siRNA. In the setting of H19 knockdown, miR-19a levels were significantly elevated in OGD neuronal cells (Figure 4C, p<0.05), suggesting H19 decreased miR-19a level after H/I.
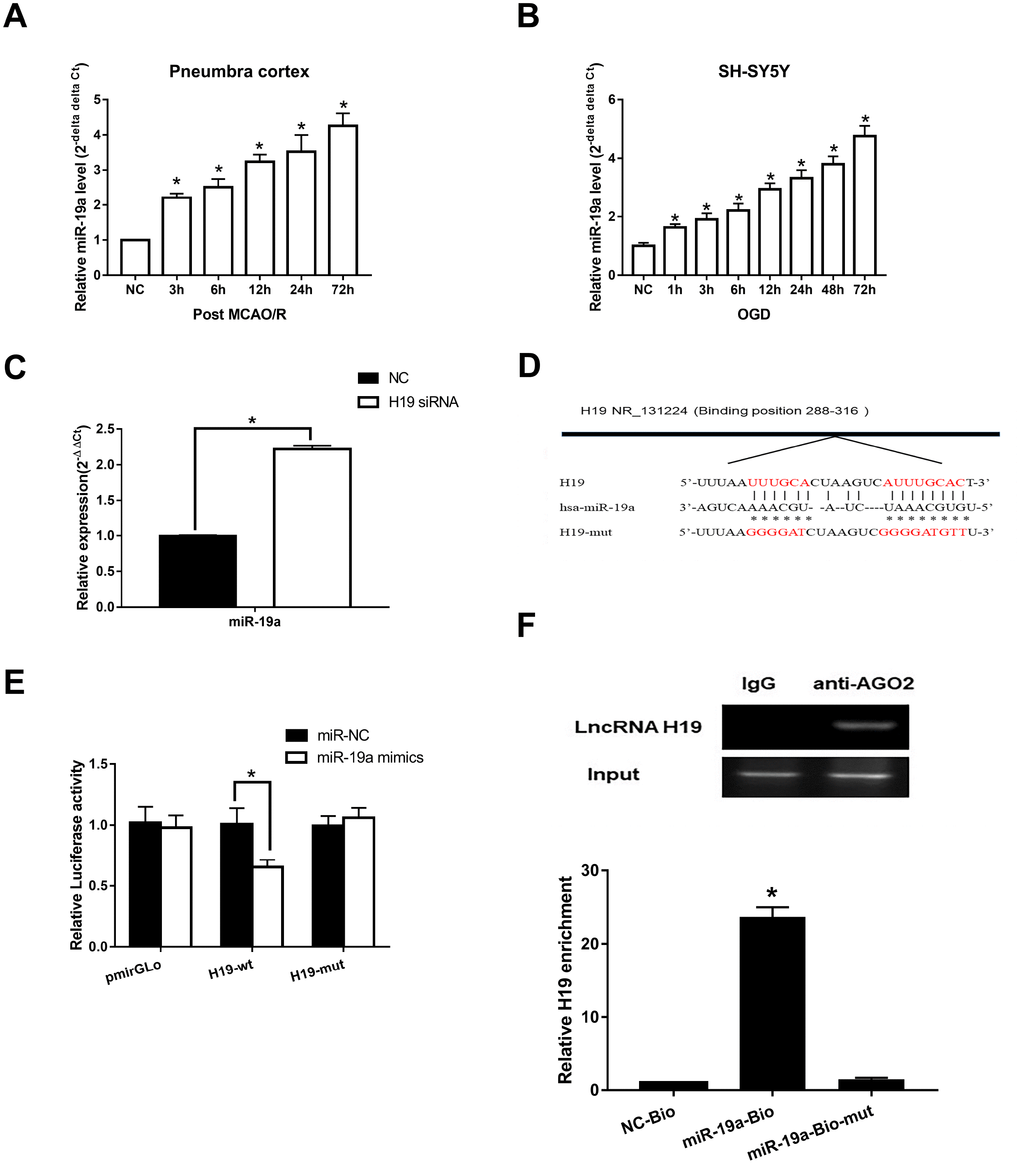
Figure 4. Direct interaction between H19 and miR-19a. (A) qRT-PCR analysis revealed that miR-19a levels in penumbra cortex of rats increased gradually and significantly within 72 h after MCAO/R. (B) Similar results were observed in OGD models. (C) With knockdown of H19 by siRNA, miR-19a level was significantly elevated in OGD neuronal cells. (D) The predicted fragment including miR-19a binding sites on H19 was cloned into a pmirGLO vector (H19-wt), and a mutated vector (H19-mut) was also generated by replacing the binding sites with its complimentary sequence. (E) Dual-luciferase report assay revealed that miR-19a mimic reduced the luciferase activity of H19-wt, but not of H19-mut. (F) RIP indicated that H19 was preferentially enriched in Ago2-containing bead compared to those harboring control immunoglobulin G (IgG) antibody. Furthermore, H19 was pulled down by biotin-labeled miR-19a oligos, but not the mutated oligos.
Further, Starbase 2.0 (http://starbase.sysu.edu.cn) was used to investigate complementary binding patterns between miR-19a and H19. Predicted fragment including binding sites (Figure 4D) were cloned into two vectors: a pmirGLO vector (H19-wt), and a mutated vector (H19-mut) with replacement of binding sites with predicted complimentary sequences. Dual-luciferase report assay revealed that miR-19a mimic reduced the luciferase activity of H19-wt, but not of H19-mut (Figure 4E, p<0.05). RIP results indicated that H19 was preferentially enriched in Ago2-containing beads compared to those harboring control immunoglobulin G (IgG) antibody (Figure 4F). Lastly, we performed pull down experiments using biotin-labeled miR-19a oligos. H19 was pulled down by biotin-labeled miR-19a oligos, but not the mutated oligos (binding sites were mutated to the complement sequences) (Figure 4F; p<0.05). Taken together, these results suggested that miR-19a directly bound to H19.
Id2 mRNA is a direct target of miR-19a
We subsequently investigated the interaction between miR-19a and Id2 mRNA. We used miRanda software (http://www.microrna.org/microrna/home.do) to find miR-19a that could potentially bind to 3′-UTR of Id2 mRNA. The predicted fragment including binding sites were cloned into the wild-type (Id2-wt) and mutated fragment (Id2-mut) (Figure 5A). When treated with miR-19a mimic, the expressions of Id2 mRNA and protein in OGD neuronal cells were inhibited. In contrast, when treated with miR-19a inhibitor, Id2 mRNA and protein expressions increased (Figure 5B and 5C; p<0.05, respectively). Furthermore, dual-luciferase report assay demonstrated that miR-19a mimic reduced the luciferase activity of Id2-wt, but not of Id2-mut (Figure 5D; p<0.05). These results suggested Id2 mRNA is a direct target of miR-19a, which exerted a direct inhibitory effect on Id2 expression via binding 3′-UTR of Id2 mRNA.
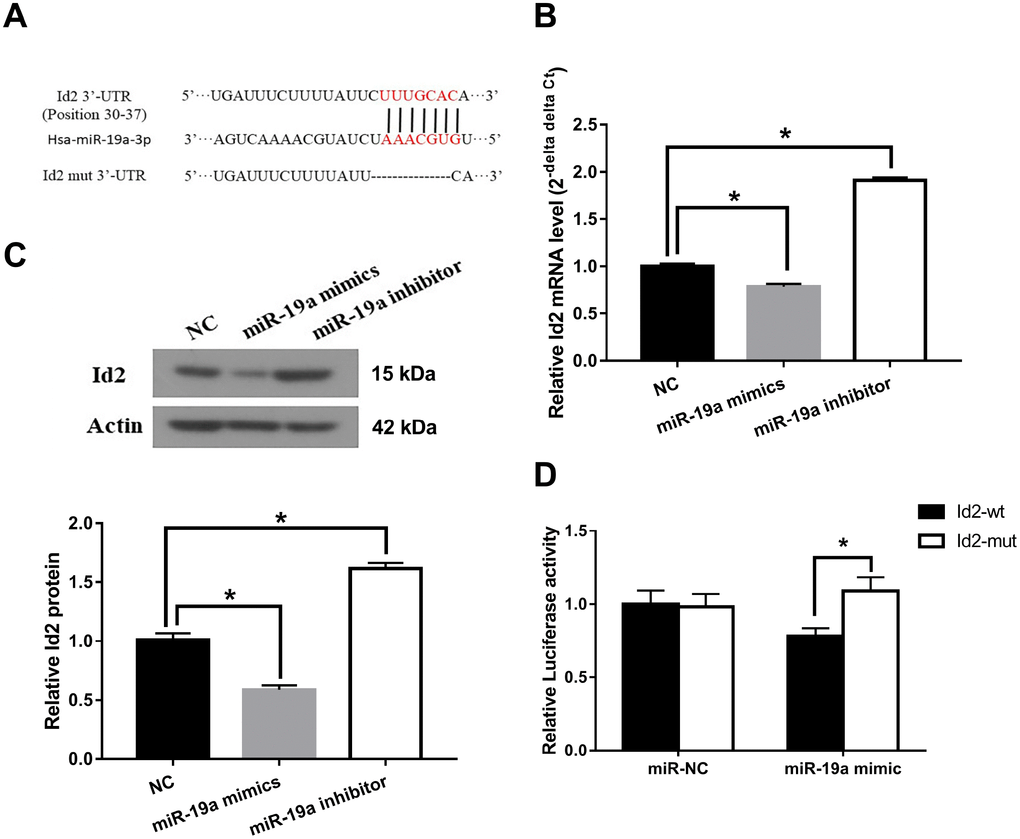
Figure 5. Id2 mRNA is a direct target of miR-19a. (A) The predicted fragment including binding sites were cloned into the wild-type (Id2-wt) and mutated fragment (Id2-mut). (B) miR-19a mimic inhibited the expression of Id2 mRNA in OGD neuronal cells, while miR-19a inhibitor increased its expression. (C) miR-19a mimic inhibited the expression of Id2 protein in OGD neuronal cells, while miR-19a inhibitor increased its expression. (D) Dual-luciferase report assay demonstrated that miR-19a mimic reduced the luciferase activity of Id2-wt, but not of Id2-mut.
H19-miR-19a-Id2 axis modulated hypoxia-induced neuronal apoptosis in vitro
Given the aforementioned results, we hypothesized that the H19-miR-19a-Id2 axis modulated hypoxia induced neuronal apoptosis. To test this hypothesis, luciferase reporter plasmids containing the 3′-UTR of Id2 were constructed. As expected, knockdown of H19 decreased luciferase activity in HEK293T cells transfected with Luc-Id2-3′-UTR (Figure 6A; p<0.05); conversely, luciferase activity was rescued with the introduction of miR-19a inhibitor (Figure 6A; p>0.05). Furthermore, in OGD neuronal cells, knockdown of H19 decreased the expression of Id2 mRNA and protein (Figure 6B and 6C; p<0.05); while this inhibition was attenuated by co-transfection of miR-19a inhibitor (Figure 6B and 6C; p>0.05). Lastly, flow cytometric analysis revealed that knockdown of H19 decreased cell apoptosis in OGD neuronal cells (Figure 6D; p<0.05); however, this decrease was relieved by co-transfection of miR-19a inhibitor (Figure 6D; p>0.05). Taken together, these results demonstrated that H19-miR-19a-Id2 axis modulated hypoxia induced neuronal apoptosis, and blocking H19-miR-19a-Id2 axis could alleviate H/I induced neuronal injury.
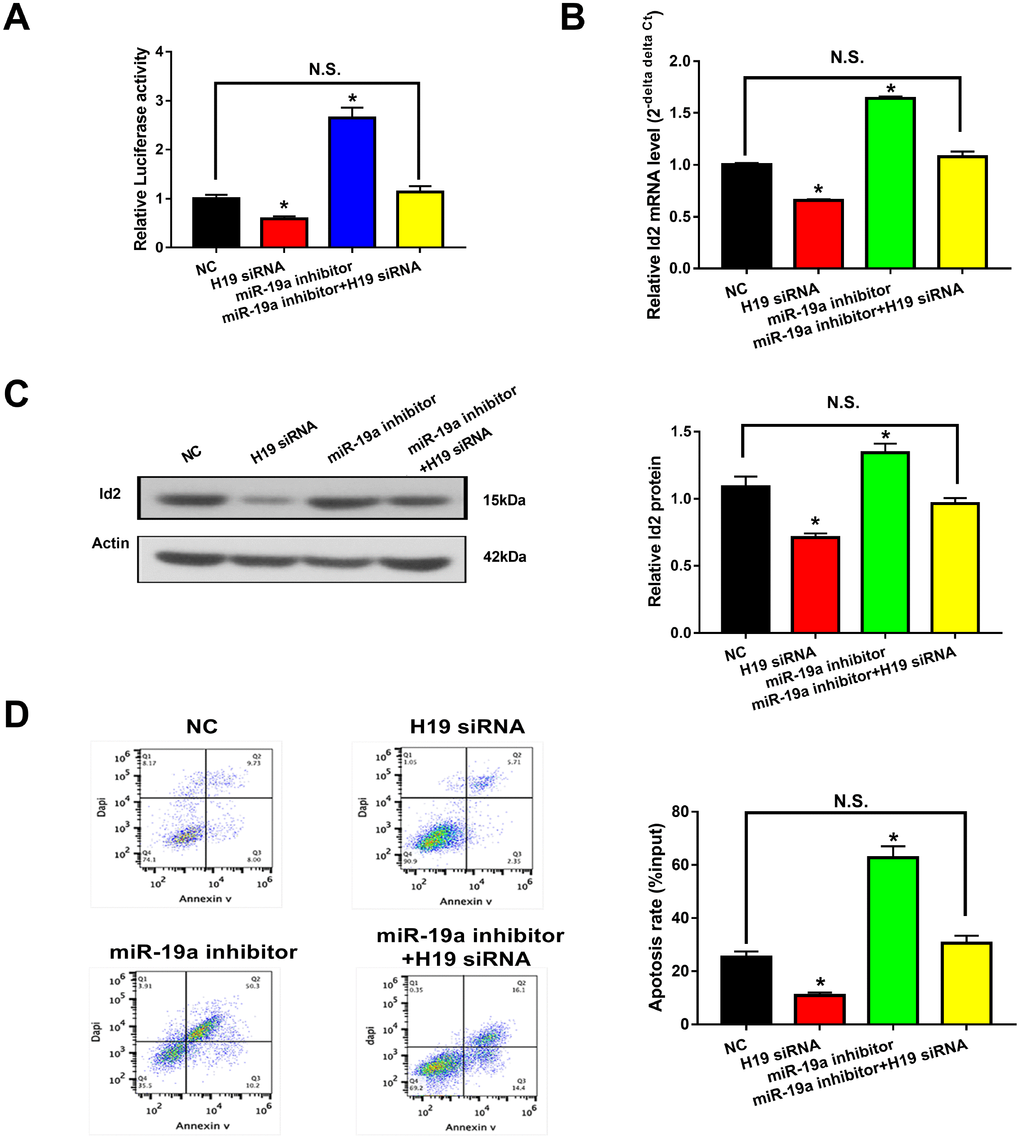
Figure 6. H19-miR-19a-Id2 axis modulated hypoxia-induced neuronal apoptosis. (A) Knockdown of H19 decreased luciferase activity in HEK293T cells transfected with Luc-Id2-3′-UTR; on the other hand, luciferase activity could be rescued by miR-19a inhibitor. In OGD neuronal cells, knockdown of H19 decreased the expression of Id2 mRNA (B) and protein (C); decreased expressions of Id2 mRNA and protein levels were attenuated by co-transfection with miR-19a inhibitor. (D) Flow cytometric analysis revealed that knockdown of H19 decreased cell apoptosis in OGD neuronal cells; however, this decrease was relieved by co-transfection of miR-19a inhibitor.
Discussion
In the present study, we first demonstrated that H19 level was significantly elevated in ischemic stroke patients, in-vivo MCAO/R animal model and in-vitro OGD neuronal cell model. Further, knockdown of H19 with siRNA alleviated cell apoptosis in OGD neuronal cells, and inhibition of H19 in MCAO/R rats significantly attenuated neurological deficit, decreased brain infarct volume and neuronal apoptosis. Lastly, we identified the axis of H19-miR-19a-Id2 modulated hypoxia-induced neuronal apoptosis. Taken together, these results suggested blocking of H19-miR-19a-Id2 axis may serve as a novel therapeutic strategy for ischemic brain injury.
Generally, H19 expression was high in embryonic organs and mostly absent or greatly reduced in adult tissues. Of note, H19 level was demonstrated to be significantly promoted by HIF under hypoxic tumor environments [13, 14]. Similarly, elevated H19 levels have been implicated in the setting of ischemic stroke [15]. In the present study, we confirmed that H19 levels were not only significantly elevated in ischemic stroke patients, in-vivo MCAO/R rats and in-vitro OGD neuronal cells, but also demonstrated circulating H19 levels were positively correlated with patient NIHSS scores within 3 hours from stroke onset. These results indicated H19 level could be a diagnostic and prognostic biomarker for ischemic stroke. However, circulating H19 levels may be a non-specific biomarker, since elevated circulating H19 levels have been implicated in diverse oncological conditions [9–12, 22–26]. Known oncological etiologies would have to be ruled out prior to designating H19 level as a more specific biomarker for cerebra ischemia. Perhaps H19 level in cerebrospinal fluid (CSF) following H/I would render more specific finding. Hence, assessment of H19 level in the CSF and its specificity in determining neuronal injury in the setting of H/I will be the subject of future studies.
Furthermore, we found that inhibition of H19 with siRNA reduced neuronal apoptosis in vitro and in vivo. Recently, Wang et al. reported that H19 could induce cerebral ischemia reperfusion injury via activation of autophagy, and inhibition of H19 could protect neuronal cells from OGD induced death [15]; Zhao et al. reported H19 could induce hippocampal neuronal apoptosis via Wnt signaling, and that inhibition of H19 might serve as a promising novel target for the treatment of cognitive decline in patient with diabetes mellitus [27]. Therefore, consistent with our study, we suggest that H19 plays a critical role in H/I induced neuronal injury.
Interestingly, the elevated H19 level demonstrated a similar trend as Id2 expression after the exposure of H/I in our in-vivo and in-vitro studies. Although a similar relationship between H19 and Id2 has been indicated previously in bladder cancer cells and acute myelocytic leukemia cells, the underlying mechanistic relationship between them have not been well investigated [17, 18]. Inspired by the newly reported regulatory mechanism that H19 functioned as a competing endogenous RNA (ceRNA) via sponging miRNA [19–21, 28–30], we employed bioinformatics analysis to predict that H19 promoted Id2 expression via acting as a ceRNA by sponging miR-19a. With dual luciferase reported assay, RIP and pull-down experiments, we demonstrated the dual competitive interaction of miR-19a with H19 and the 3′-UTR of Id2 mRNA. For the first time, we identified the axis of H19-miR-19a-Id2 in the process of H/I induced neuronal apoptosis, and provide a schematic figure outlining the mechanistic relationships of H19-miR-19a-Id2 axis (Figure 7). Lastly, we demonstrated that H19-miR-19a-Id2 axis modulates H/I-induced neuronal apoptosis and blockage of the axis may have therapeutic benefit in the treatment of ischemic brain injury.
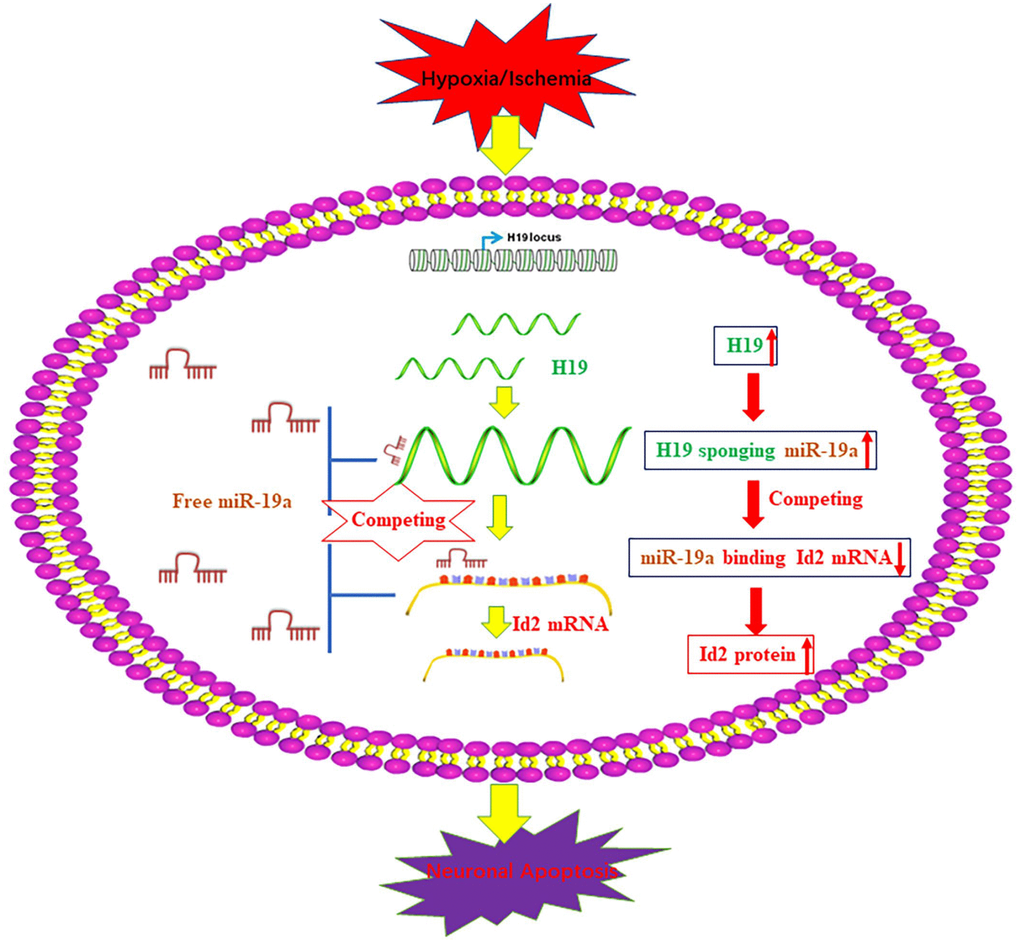
Figure 7. A schematic figure of the axis of H19-miR-19a-Id2. A schematic figure demonstrates the regulatory mechanism of the H19-miR-19a-Id2 axis and its role in modulating H/I-induced neuronal apoptosis.
In conclusion, our findings demonstrated H19 level was elevated in the setting of H/I, and inhibition of H19 significantly reduced H/I induced neuronal apoptosis in vivo and in vitro. Further, we also demonstrated that H19-miR-19a-Id2 axis played a critical role in hypoxia induced neuronal apoptosis. Thus, blocking of H19-miR-19a-Id2 axis could attenuate H/I induced neuronal injury, which sheds new light on the treatment of ischemic brain stroke.
Methods
Study subjects
Forty (40) patients with anterior circulation ischemia and 25 healthy controls were selected from Renji Hospital, School of Medicine, Shanghai Jiaotong University between September 1, 2016, and July 31, 2018. Inclusion criteria were clinical signs and symptoms of focal or global cerebral function loss. Patients who demonstrated new-onset cerebral infarction on magnetic resonance imaging (MRI) within 3 hours from the time of admission were also included in the study. Exclusion criteria included patients with evidence of prior cerebral infarcts and those with the following co-morbidities: diabetes mellitus, coronary artery disease, hypertension, kidney diseases, circulation disorders, or autoimmune diseases. All study subjects underwent routine biochemical tests, cerebral MRIs, and baseline neurological dysfunction assessment using the National Institute of Health Stroke Scale (NIHSS). All study subjects were matched for age (cases, 67.2±9.3; controls, 65.7±10.3) and sex (cases, 25 males to 15 females; controls, 16 males to 9 females). All enrolled patients provided written informed consent, and the study protocol was approved by ethic Committee of Renji Hospital, School of Medicine, Shanghai Jiaotong University.
Cell culture, and OGD model
SH-SY5Y cells (ATCC) were incubated in Dulbecco’s modified Eagles’s medium (DMEM; Hyclone, USA) supplemented with 10% fetal bovine serum (FBS; Gibco, USA) at 37°C in a humidified atmosphere with 5% CO2. HEK-293T cells were cultured in RPMI-1640 medium (Gibco, Grand Island, NY, USA) supplied with 10% fetal bovine serum, 100U/ml penicillin, and 100 μg/ml streptomycin, at 37°C in a 5%CO2 atmosphere.
For OGD model, the SH-SY5Y cells were incubated in serum and sugar-free artificial cerebrospinal fluid at 37°C in an atmosphere of 1% O2, 94% N2 and 5% CO2 for 4 h and then incubated in DMEM medium in normoxic atmosphere.
In addition, miR-19a mimics, miR-19a inhibitor and their scramble controls were obtained from Guangzhou RiboBio Co., Ltd. (Guangzhou, China). The siRNAs (named as siRNA-1, siRNA-2 and siRNA-3) directed against H19 and negative control H19 scramble were purchased from Invitrogen Life Technologies (Waltham, MA, USA). The sequences of siRNAs were listed in Table 1.
Table 1. Sequences of primers and siRNAs used in this study.
| Names | Sequences |
| H19 siRNA | |
| siRNA-1 | 5′-GCAAGAAGCGGGTCTGTTTCT-3′ |
| siRNA-2 | 5′-GACAAGCAGGACATGACATGG-3′ |
| siRNA-3 | 5′-GCACTACCTGACTCAGGAATC-3′ |
| mis siRNA | 5′-TTGCACGCTAACTTAGGTTCGA-3′ |
| qRT-PCR primers | |
| H19-F (for human samples and SH-SY5Y cells) | 5′-GAAGGCCAAGACGCCAGG-3′ |
| H19-R (for human samples and SH-SY5Y cells) | 5′-TCCTCTGTCCTCGCCGTCAC-3′ |
| H19-F (for rat samples) | 5′-TTCAAAGCCTCCACGACTCT-3′ |
| H19-R (for rat samples) | 5′-GCTCACACTCACGCACACTC-3′ |
| β-actin-F | 5′-GTGGCCGAGGACTTTGATTG-3′ |
| β-actin-R | 5′-CCTGTAACAACGCATCTCATATT -3′ |
| Id2-F | 5′-GGACAGAACCAAACGTCCAG-3′ |
| Id2-R | 5′-TAAGCTCAGAAGGGAATTCAGAC-3′ |
| Plasmid construction | |
| Id2-3′UTR reporter-F | CCGCTCGAGATAAGCGGTGTTCATGAT |
| Id2-3′UTR reporter-R | ATAAGAATGCGGCCGCTTTGACTTCAGACATGTTT |
Transfection
SH-SY5Ycells were transiently transfected with H19 siRNAs using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA). Briefly, after cultured in normal culture medium for 24 h, the culture medium was replaced by normal culture medium plus Lipofectamine 2000 complex, and cells were incubated in a humidified incubator for 24 h.
Intraventricular injection of H19 siRNA
Sprague-Dawley rats weighing 250-300g were obtained from the Experimental Animal Center of Renji Hospital, School of Medicine, Shanghai Jiaotong University. The animal study was approved by the institutional Ethics Committee for Animal Experimentation and conducted in compliance with the Guidelines for Animal Experimentation of this institution. Twenty-four (24) rats were randomly divided into the following 3 groups (n=8 per group): control (intraventricular injection with 0.9% saline), mis siRNA (intraventricular injection with mis siRNA), and H19 siRNA (intraventricular injection H19 siRNA). Intraventricular injections were achieved using the following method : 1) rats were anesthetized with intraperitoneal (i.p) injections of pentobarbital (50mg/kg); rats were then placed in stereotactic frame; 2) a 28-guage stainless steel injection cannula was introduced into the right lateral ventricle to deliver single injection according to group (injections were delivered at a rate of 0.5μl/min using a micro-infusion pump and a 10 μl microsyringe); 3) intraventricular cannulas were withdrawn after 5 mins from infusion time at a rate of 1mm/min.
Rat model of middle cerebral artery occlusion/reperfusion (MCAO/R)
One day after the intraventricular injection of H19 siRNA, all the rats were performed with the operation of MCAO/R (as previously reported [6]). Briefly, a 3-0 monofilament nylon suture (Beijing Shadong Industrial Corp., China) was introduced into the right internal carotid artery via the external carotid artery until a slight resistance was encountered. The suture was left in place for 90 min and removed to facilitate reperfusion.
Assessment of neurological deficits for MCAO/R rats
A standardized 0-4 scales was used to evaluate the neurological behavior at 1 day, 3 day and 7 day following MCAO/R [31, 32]. The neurological deficit testing was performed by two observers who were individually blinded to the assignments of the three animal groups.
Infarct volume measurement
72 h after MCAO/R, the rat brain tissue were collected and the frontal end was sliced and immersed in 2% solution of 2,3,7,-triphenyltetrazolium chloride (TTC) (Sigma Inc., MO, USA) for 30 min at 37°C. Since an infarct tissue does not stain owing to the loss of mitochondrial enzyme activity and a normal tissue will be stained red by TTC, the infarction area that appears white and hemisphere area of each section could be distinguished and quantified by an image analysis system (OlympusImaging ProPlus, Silver Spring, MD, USA). The infarct volume was calculated by dividing infarct area of the ipsilateral hemisphere by total hemisphere.
Cortex tissue collection and sectioning
The rats were anesthetized by i.p. injection of sodium pentobarbital at 100mg/kg. For RNA and protein expression, the rats were perfused transcardially with 4°C normal saline, and the brain regions that corresponded to the ischemic core and penumbra were dissected on ice according to a previous method [6]. For TUNEL staining, the rats were perfused transcardially with 4% paraformaldehyde (PFA, pH7.4) following 4°C normal saline. Upon removal, the brains were post-fixed in 4% PFA and immediately cryoprotected in 20% and 30% sucrose solution sequentially. A series of coronal sections (25 μm) were made between -1.80 and -4.80 mm bregma levels. Every fifth section totaling 24 sections was used for the staining.
TUNEL assay
Apoptosis was evaluated on the sections of the cerebral cortices of penumbra area mounted on slides using the Roche TUNEL staining kit (Penzberg, Germany) following the manufacturer’s instructions. The TUNEL-positive cells were counted in five randomized areas per rat brain and expressed as the number of positively stained cells per square millimeter.
Immunofluorescence
The cells were fixed in 4% formaldehyde for 10 min and were permeabilized in 0.5% Trition X-100 for 10 min and blocked in 1% BSA solution for 45 min. The cells were incubated with the Roche TUNEL staining kit (Penzberg, Germany) following the manufacturer’s instructions. The cells were then counterstained with DAPI (5g/L) for 5 min and observed under fluorescence microscope (Olympus, Japan).
Flow cytometric analysis
Negative control cells or cells transfected with the desired plasmid were plated in six-well plates. After 48 h incubation, the cultures were incubated with propidium iodide for 30 min in the dark. The percentage of cells in G0/G1, S and G2/M phases of the cell cycle were measured using a flow cytometer (FACS Calibur, BD Biosciences, San Jose, CA, USA) after propidium iodide staining. Cultures were also analyzed for cell apoptosis after double staining with FITC-Annexin V and Propidium iodide (PI). The cells were analyzed with a flow cytometer (FACScan, BD Biosciences) equipped with CellQuest software (BD Biosciences).
Dual-luciferase reporter assay
Human HEK293T cells were co-transfected with 150 ng of either empty, pmirGLO-NC, pmirGLO-H19-wt or pmirGLO-H19-mut (Sangon biotech, China). PmirGLO-Id2-wt (Sangon biotech, China) were transfected into HEK293T cells by Lipofectamine 2000 (Invitrogen, USA) -mediated gene transfer, and miR-19a mimics or inhibitor or mis siRNA (NC) were also transfected into HEK293T cells by using the same method. The relative luciferase activity was normalized to Renilla luciferase activity 48 h after transfection. Transfection was repeated in triplicate.
RNA immunoprecipitation (RIP)
RIP assay was performed using the EZ-Magna RIPTM RNA-Binding Protein Immunoprecipitation Kit (Millipore, MA, USA) according to the manufacturer’s instructions. SH-SY5Y cells at 80-90% confluency were scraped off and lysed in complete RIP lysis buffer. Then, 100 μl of whole cell extract were incubated with RIP buffer containing magnetic beads conjugated with human anti-Ago2 antibody (Cell Signaling, USA), negative control normal mouse IgG (Millipore) and positive control SNRNP70 (Millipore). The co-precipitated RNAs were detected by reverse transcription PCR. Total RNAs (input controls) and IgG were assayed simultaneously to test whether the detected signals resulted from RNAs specifically binding to Ago2.
Pulldown assay with biotinylated miRNA
SH-SY5Y cells were transfected with Biotinylated miR-19a or biotinylated mutant miR-19a or biotinylated NC (synthesized by Shanghai Gene Pharma Co. Ltd) using Lipofectamine 2000 according to the manufacturer’s instruction. The final concentration of each biotinylated miRNA was 20 nM. The cell lysates were collected 48 h after transfection and incubated with M-280 streptaviden magnetic beads (Invitrogen, San Diego, CA, USA) as described previously [33]. The bound RNAs were purified using TRIzol reagent (TAKALA) for further qRT-PCR analysis.
Quantitative real-time polymerase chain reaction (qRT-PCR)
To assess H19 levels in ischemic stroke patients, peripheral blood samples were drawn from these stroke patients within 3 hours of stroke onset, as well as the healthy controls. All blood samples underwent qRT-PCR to measure H19 levels in serum.
Total RNA in the ischemic patients’ blood serum or the cerebral cortices of penumbra area in MCAO/R rats or OGD SH-SY5Y cells were extracted using TRIzol reagent (Invitrogen, Carlsbad, CA) and further purified with a RNA Micro Kit (Qiagen, Valencia, CA, USA). The following PCR reaction conditions were used when performed in a lightcycler PCR detection system (Roche Diagnostics Ltd., Shanghai, China): 10 min at 95°C, 40 cycles of 10 s at 95°C, 15 s at 60°C, and 20 s at 72°C. Primers for amplification of H19, Id2, and β-actin were synthesized by Invitrogen Life Technologies, and are listed in Table 1. The β-actin gene was used as a reference gene, and relative mRNA expression levels were calculated using the standard 2−ΔΔCt method.
Western blot analysis
Proteins were extracted from the cerebral cortices of the penumbra area in MCAO/R rats or OGD SH-SY5Y cells with RIPA lysis buffer (KenGen Biotech, Nanjing, China) and were quantified using a BCA protein Assay kit (Thermo Fisher Scientific, Inc., Waltham, MA, USA). A total of 30 mg protein lysates were loaded to SDS-PAGE gel, and transferred to PVDF membrane (Millipore, Billerica, MA, USA). The transferred membrane was probed overnight at 4°C with specific primary antibodies (anti-Id2, 1:200 dilution, Santa Cruz Biotechnology; anti-Bax, 1:1000 dilution, Cell signaling technology; anti-β-actin, 1:50000 dilution, Sigma-Aldrich, St.Louis, MO) followed by horseradish peroxidase (HRP)-conjugated goat anti-rabbit secondary antibodies (1:2500 dilution, Promega, Madison, WI, USA). β-actin was used as a control (1:1000; Cell Signaling Technology, Inc., Danvers, MA, USA).
Statistical analysis
Data was presented as means ± standard deviation and the difference as analyzed by one-way ANOVA followed by Bonferroni’s post hoc test. Pearson’s correlation coefficient was used to analyze the correlation between H19 levels and NIHSS scores, H19 and miR-19a. A P-value less than 0.05 was considered statistically significant.
Ethics approval
The study was conducted under the approval of the Ethic Committee of Renji Hospital, School of Medicine, Shanghai Jiaotong University. All enrolled patients provided written informed consent. All animal experiments in this study conformed to the principles of the management and use of local experimental animals and followed the Guide for the Care and Use of Laboratory Animals published by the National Institute of Health.
Supplementary Materials
Author Contributions
Liemei Guo, Xiaohua Zhang and Yongming Qiu designed the experiments, analyzed data, and wrote the article; Zhipeng Xiao, Yingying Lin and Zezhi Li performed the main experiments; and Rogelio Medina, Sophie Zhuang, Jared S Rosenblum and Jing Cui helped with the experiments and revised the article.
Conflicts of Interest
No conflicts of interest exit in the submission of this manuscript. All the authors listed have read and approved the manuscript that is enclosed.
Funding
This work was supported by grants from the National Natural Science Fund of China (81701201 and 81671203), the Renji Hospital Zhong Zi Research Program (RJZZ16-018), the Renji Hospital Pei Yu Zi You Research Program (PYZY16-009) and Project of Shanghai Science and Technology Commission (16140902900).
References
- 1. Kernan WN, Ovbiagele B, Black HR, Bravata DM, Chimowitz MI, Ezekowitz MD, Fang MC, Fisher M, Furie KL, Heck DV, Johnston SC, Kasner SE, Kittner SJ, et al, and American Heart Association Stroke Council, Council on Cardiovascular and Stroke Nursing, Council on Clinical Cardiology, and Council on Peripheral Vascular Disease. Guidelines for the prevention of stroke in patients with stroke and transient ischemic attack: a guideline for healthcare professionals from the American Heart Association/American Stroke Association. Stroke. 2014; 45:2160–236. https://doi.org/10.1161/STR.0000000000000024 [PubMed]
- 2. George PM, Steinberg GK. Novel Stroke Therapeutics: Unraveling Stroke Pathophysiology and Its Impact on Clinical Treatments. Neuron. 2015; 87:297–309. https://doi.org/10.1016/j.neuron.2015.05.041 [PubMed]
- 3. Deb P, Sharma S, Hassan KM. Pathophysiologic mechanisms of acute ischemic stroke: an overview with emphasis on therapeutic significance beyond thrombolysis. Pathophysiology. 2010; 17:197–218. https://doi.org/10.1016/j.pathophys.2009.12.001 [PubMed]
- 4. Turley KR, Toledo-Pereyra LH, Kothari RU. Molecular mechanisms in the pathogenesis and treatment of acute ischemic stroke. J Invest Surg. 2005; 18:207–18. https://doi.org/10.1080/08941930591004449 [PubMed]
- 5. Guo L, Yang X, Lin X, Lin Y, Shen L, Nie Q, Ren L, Guo Q, Que S, Qiu Y. Silencing of Id2 attenuates hypoxia/ischemia-induced neuronal injury via inhibition of neuronal apoptosis. Behav Brain Res. 2015; 292:528–36. https://doi.org/10.1016/j.bbr.2015.07.018 [PubMed]
- 6. Guo L, Lan J, Lin Y, Guo P, Nie Q, Mao Q, Ren L, Qiu Y. Hypoxia/ischemia up-regulates Id2 expression in neuronal cells in vivo and in vitro. Neurosci Lett. 2013; 554:88–93. https://doi.org/10.1016/j.neulet.2013.08.044 [PubMed]
- 7. Mercer TR, Dinger ME, Mattick JS. Long non-coding RNAs: insights into functions. Nat Rev Genet. 2009; 10:155–59. https://doi.org/10.1038/nrg2521 [PubMed]
- 8. Gabory A, Ripoche MA, Yoshimizu T, Dandolo L. The H19 gene: regulation and function of a non-coding RNA. Cytogenet Genome Res. 2006; 113:188–93. https://doi.org/10.1159/000090831 [PubMed]
- 9. Byun HM, Wong HL, Birnstein EA, Wolff EM, Liang G, Yang AS. Examination of IGF2 and H19 loss of imprinting in bladder cancer. Cancer Res. 2007; 67:10753–58. https://doi.org/10.1158/0008-5472.CAN-07-0329 [PubMed]
- 10. Tian F, Tang Z, Song G, Pan Y, He B, Bao Q, Wang S. Loss of imprinting of IGF2 correlates with hypomethylation of the H19 differentially methylated region in the tumor tissue of colorectal cancer patients. Mol Med Rep. 2012; 5:1536–40. https://doi.org/10.3892/mmr.2012.833 [PubMed]
- 11. Lottin S, Adriaenssens E, Dupressoir T, Berteaux N, Montpellier C, Coll J, Dugimont T, Curgy JJ. Overexpression of an ectopic H19 gene enhances the tumorigenic properties of breast cancer cells. Carcinogenesis. 2002; 23:1885–95. https://doi.org/10.1093/carcin/23.11.1885 [PubMed]
- 12. Hashad D, Elbanna A, Ibrahim A, Khedr G. Evaluation of the Role of Circulating Long Non-Coding RNA H19 as a Promising Novel Biomarker in Plasma of Patients with Gastric Cancer. J Clin Lab Anal. 2016; 30:1100–05. https://doi.org/10.1002/jcla.21987 [PubMed]
- 13. Wu W, Hu Q, Nie E, Yu T, Wu Y, Zhi T, Jiang K, Shen F, Wang Y, Zhang J, You Y. Hypoxia induces H19 expression through direct and indirect Hif-1α activity, promoting oncogenic effects in glioblastoma. Sci Rep. 2017; 7:45029. https://doi.org/10.1038/srep45029 [PubMed]
- 14. Matouk IJ, Mezan S, Mizrahi A, Ohana P, Abu-Lail R, Fellig Y, Degroot N, Galun E, Hochberg A. The oncofetal H19 RNA connection: hypoxia, p53 and cancer. Biochim Biophys Acta. 2010; 1803:443–51. https://doi.org/10.1016/j.bbamcr.2010.01.010 [PubMed]
- 15. Wang J, Cao B, Han D, Sun M, Feng J. Long Non-coding RNA H19 Induces Cerebral Ischemia Reperfusion Injury via Activation of Autophagy. Aging Dis. 2017; 8:71–84. https://doi.org/10.14336/AD.2016.0530 [PubMed]
- 16. Wang J, Zhao H, Fan Z, Li G, Ma Q, Tao Z, Wang R, Feng J, Luo Y. Long Noncoding RNA H19 Promotes Neuroinflammation in Ischemic Stroke by Driving Histone Deacetylase 1-Dependent M1 Microglial Polarization. Stroke. 2017; 48:2211–21. https://doi.org/10.1161/STROKEAHA.117.017387 [PubMed]
- 17. Luo M, Li Z, Wang W, Zeng Y, Liu Z, Qiu J. Upregulated H19 contributes to bladder cancer cell proliferation by regulating ID2 expression. FEBS J. 2013; 280:1709–16. https://doi.org/10.1111/febs.12185 [PubMed]
- 18. Zhao TF, Jia HZ, Zhang ZZ, Zhao XS, Zou YF, Zhang W, Wan J, Chen XF. LncRNA H19 regulates ID2 expression through competitive binding to hsa-miR-19a/b in acute myelocytic leukemia. Mol Med Rep. 2017; 16:3687–93. https://doi.org/10.3892/mmr.2017.7029 [PubMed]
- 19. Wang WT, Ye H, Wei PP, Han BW, He B, Chen ZH, Chen YQ. LncRNAs H19 and HULC, activated by oxidative stress, promote cell migration and invasion in cholangiocarcinoma through a ceRNA manner. J Hematol Oncol. 2016; 9:117. https://doi.org/10.1186/s13045-016-0348-0 [PubMed]
- 20. Li M, Chen H, Zhao Y, Gao S, Cheng C. H19 Functions as a ceRNA in Promoting Metastasis Through Decreasing miR-200s Activity in Osteosarcoma. DNA Cell Biol. 2016; 35:235–40. https://doi.org/10.1089/dna.2015.3171 [PubMed]
- 21. Liang WC, Fu WM, Wang YB, Sun YX, Xu LL, Wong CW, Chan KM, Li G, Waye MM, Zhang JF. H19 activates Wnt signaling and promotes osteoblast differentiation by functioning as a competing endogenous RNA. Sci Rep. 2016; 6:20121. https://doi.org/10.1038/srep20121 [PubMed]
- 22. Jiao ZY, Tian Q, Li N, Wang HB, Li KZ. Plasma long non-coding RNAs (lncRNAs) serve as potential biomarkers for predicting breast cancer. Eur Rev Med Pharmacol Sci. 2018; 22:1994–99. https://doi.org/10.26355/eurrev_201804_14727 [PubMed]
- 23. Zhou X, Yin C, Dang Y, Ye F, Zhang G. Identification of the long non-coding RNA H19 in plasma as a novel biomarker for diagnosis of gastric cancer. Sci Rep. 2015; 5:11516. https://doi.org/10.1038/srep11516 [PubMed]
- 24. Wang J, Yang K, Yuan W, Gao Z. Determination of Serum Exosomal H19 as a Noninvasive Biomarker for Bladder Cancer Diagnosis and Prognosis. Med Sci Monit. 2018; 24:9307–16. https://doi.org/10.12659/MSM.912018 [PubMed]
- 25. Luo J, Li Q, Pan J, Li L, Fang L, Zhang Y. Expression level of long noncoding RNA H19 in plasma of patients with nonsmall cell lung cancer and its clinical significance. J Cancer Res Ther. 2018; 14:860–63. https://doi.org/10.4103/jcrt.JCRT_733_17 [PubMed]
- 26. Liu Y, He A, Liu B, Huang Z, Mei H. Potential Role of lncRNA H19 as a Cancer Biomarker in Human Cancers Detection and Diagnosis: A Pooled Analysis Based on 1585 Subjects. Biomed Res Int. 2019; 2019:9056458. https://doi.org/10.1155/2019/9056458 [PubMed]
- 27. Zhao YH, Ji TF, Luo Q, Yu JL. Long non-coding RNA H19 induces hippocampal neuronal apoptosis via Wnt signaling in a streptozotocin-induced rat model of diabetes mellitus. Oncotarget. 2017; 8:64827–39. https://doi.org/10.18632/oncotarget.17472 [PubMed]
- 28. Hong Y, He H, Sui W, Zhang J, Zhang S, Yang D. [Corrigendum] Long non-coding RNA H19 promotes cell proliferation and invasion by acting as a ceRNA of miR-138 and releasing EZH2 in oral squamous cell carcinoma. Int J Oncol. 2018; 53:915. https://doi.org/10.3892/ijo.2018.4428 [PubMed]
- 29. Zhou X, Ye F, Yin C, Zhuang Y, Yue G, Zhang G. The Interaction Between MiR-141 and lncRNA-H19 in Regulating Cell Proliferation and Migration in Gastric Cancer. Cell Physiol Biochem. 2015; 36:1440–52. https://doi.org/10.1159/000430309 [PubMed]
- 30. Liang WC, Fu WM, Wong CW, Wang Y, Wang WM, Hu GX, Zhang L, Xiao LJ, Wan DC, Zhang JF, Waye MM. The lncRNA H19 promotes epithelial to mesenchymal transition by functioning as miRNA sponges in colorectal cancer. Oncotarget. 2015; 6:22513–25. https://doi.org/10.18632/oncotarget.4154 [PubMed]
- 31. Zhang J, Wang Y, Zhu P, Wang X, Lv M, Feng H. siRNA-mediated silence of protease-activated receptor-1 minimizes ischemic injury of cerebral cortex through HSP70 and MAP2. J Neurol Sci. 2012; 320:6–11. https://doi.org/10.1016/j.jns.2012.05.040 [PubMed]
- 32. Longa EZ, Weinstein PR, Carlson S, Cummins R. Reversible middle cerebral artery occlusion without craniectomy in rats. Stroke. 1989; 20:84–91. https://doi.org/10.1161/01.STR.20.1.84 [PubMed]
- 33. Subramanian M, Li XL, Hara T, Lal A. A biochemical approach to identify direct microRNA targets. Methods Mol Biol. 2015; 1206:29–37. https://doi.org/10.1007/978-1-4939-1369-5_3 [PubMed]