Introduction
Although biological and chronological time can be dissociated to some extent by experimental manipulation, aging appears to be the most important risk factor for the deterioration of normal physiological functions and the manifestation of organ-specific or systemic pathologies, physical and mental decadency, and eventual death [1–3].
One species that – to a certain degree – escapes from the rule that natural life expectancy declines with body mass is the naked mole-rat (Heterocephalus glaber). The naked mole-rat (NMR) is a small poikilotherm rodent native to East Africa that lives strictly underground in social colonies (Kenya, Ethiopia and Somalia). Although this rodent has a similar size as the laboratory mouse (Mus musculus), it lives 10-20 times longer without showing any visible signs of aging [4]. Furthermore, the naked mole-rat can live for over 32 years in captivity [5], without facing any increased age-related risk of mortality, challenging Gompertz’s mortality law, and thus establishing the naked mole-rat as a non-aging mammal [6].
Not only naked mole-rats can live an extremely long life, but they also show a remarkably long healthspan associated with almost no decline in physiological or biochemical functions for more than 20 years [4,7]. For example, cardiac functions are well preserved in aged naked mole-rats [8], cognitive functions do not decline with age and the NMR brain seems to be naturally protected from neurodegenerative processes [9], and also very little pathologic alterations have been found in the kidneys of aged naked mole-rats [10].
In addition, typical signs of aging, such as loss of fertility, muscle atrophy, bone loss, changes in body composition or metabolism are mostly absent in the naked mole-rats [7,11–13]. Finally, the incidence of age-related diseases such as cancers or metabolic disorders is extremely low in the NMR [10,14].
Herein, we investigated age-dependent and species-specific differences in the metabolome of naked mole-rats and mice, with the objective to identify novel mechanisms that may explain the exceptional resistance of NMR against the advancement of time. We were able to identify several circulating metabolites previously associated to an increased healthspan and lifespan in other species that might explain the longevity phenotype of this model species.
Results and Discussion
Trans-species differences in the metabolome
Plasma samples from post-adolescent young (1-1.5 months), intermediate (6-10 months) and mature/old (20 months) mice were compared to plasma specimens from post-adolescent young (1 year), intermediate (4 years) and relatively mature/old (10 years) naked mole-rats. Samples were subjected to mass spectrometric metabolomics using several different extraction methods, matrices and chromatography methods (including gas and liquid chromatography) to extract a maximum of information on a wide spectrum of metabolites. Results were then filtered based on quality control criteria (see Materials and Methods) and subjected to unbiased hierarchical clustering to reveal age- and species-dependent differences (Figure 1, Supplementary Table 1). We also performed volcano plot-based comparisons between mice and naked mole-rats irrespective of their age (Figure 2a), as well as within the same species between young and old animals (Figure 2a, b). This approach clearly indicates that species differences are well more important than age, allowing to clearly separating the samples from two rodents (Figure 1). The number of metabolites that were significantly reduced in their plasma concentration in naked mole-rats as compared to mice was larger than the number of compounds that were enhanced (Figure 1 and 2, Supplementary Table 1). Bioinformatic analyses to understand the divergence in the metabolomes from mice and naked mole-rats failed to yield a simple pattern of differences (Supplementary Figure 1 and Supplementary Figure 2). Rather, the two species differ in multiple apparently unconnected pathways.
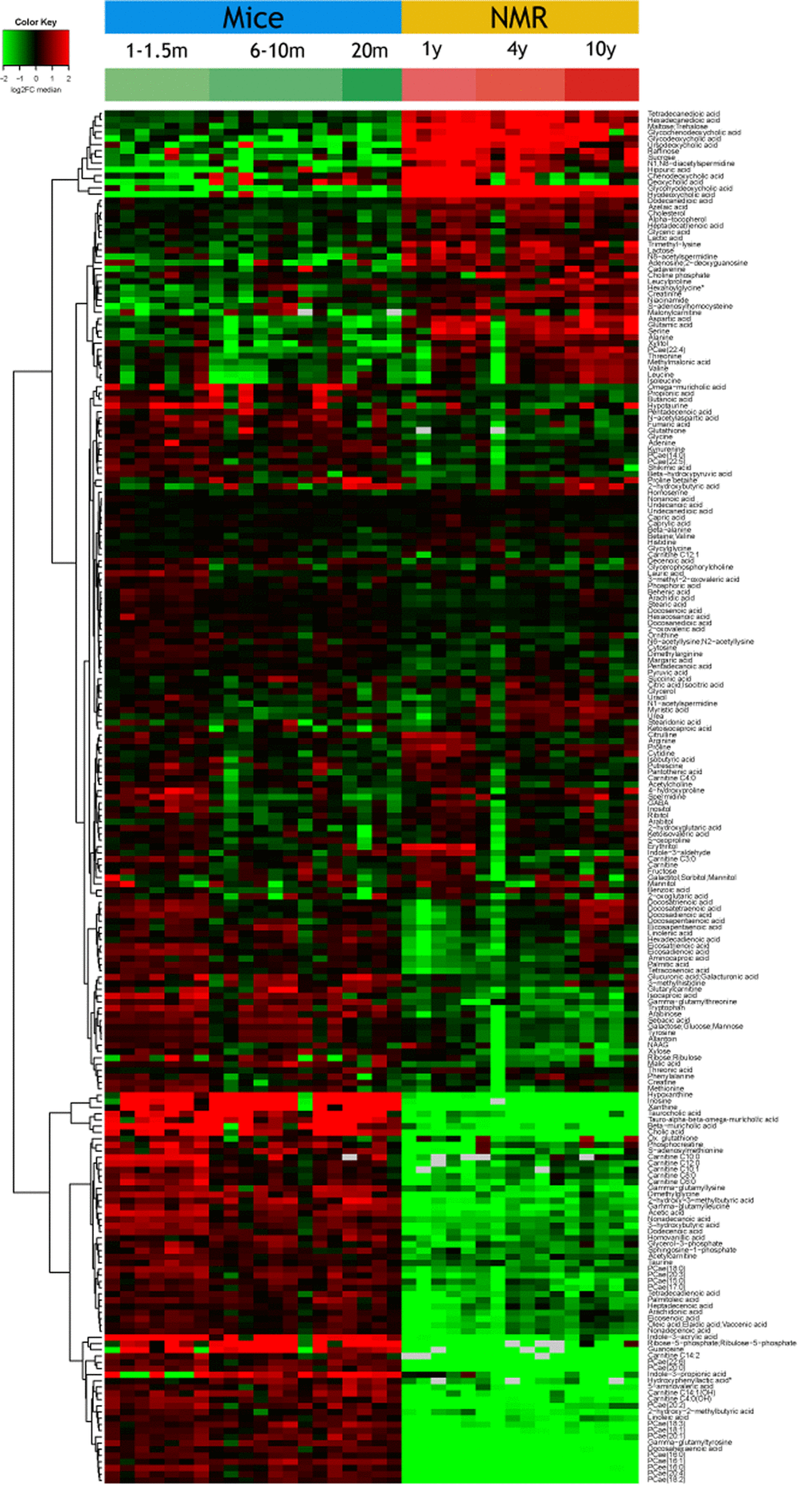

Figure 1. Overview of the plasma metabolome in the two rodents. The abundance of each metabolite is indicated for each mouse or naked mole-rat (NMR) as a heat map (red = high, green=low). Results were subjected to hierarchical clustering to indicate the increase (upper part) or decreased (lower part) of metabolites in NMR as compared to mice. Note that the raw data are listed in Supplementary Table 1.
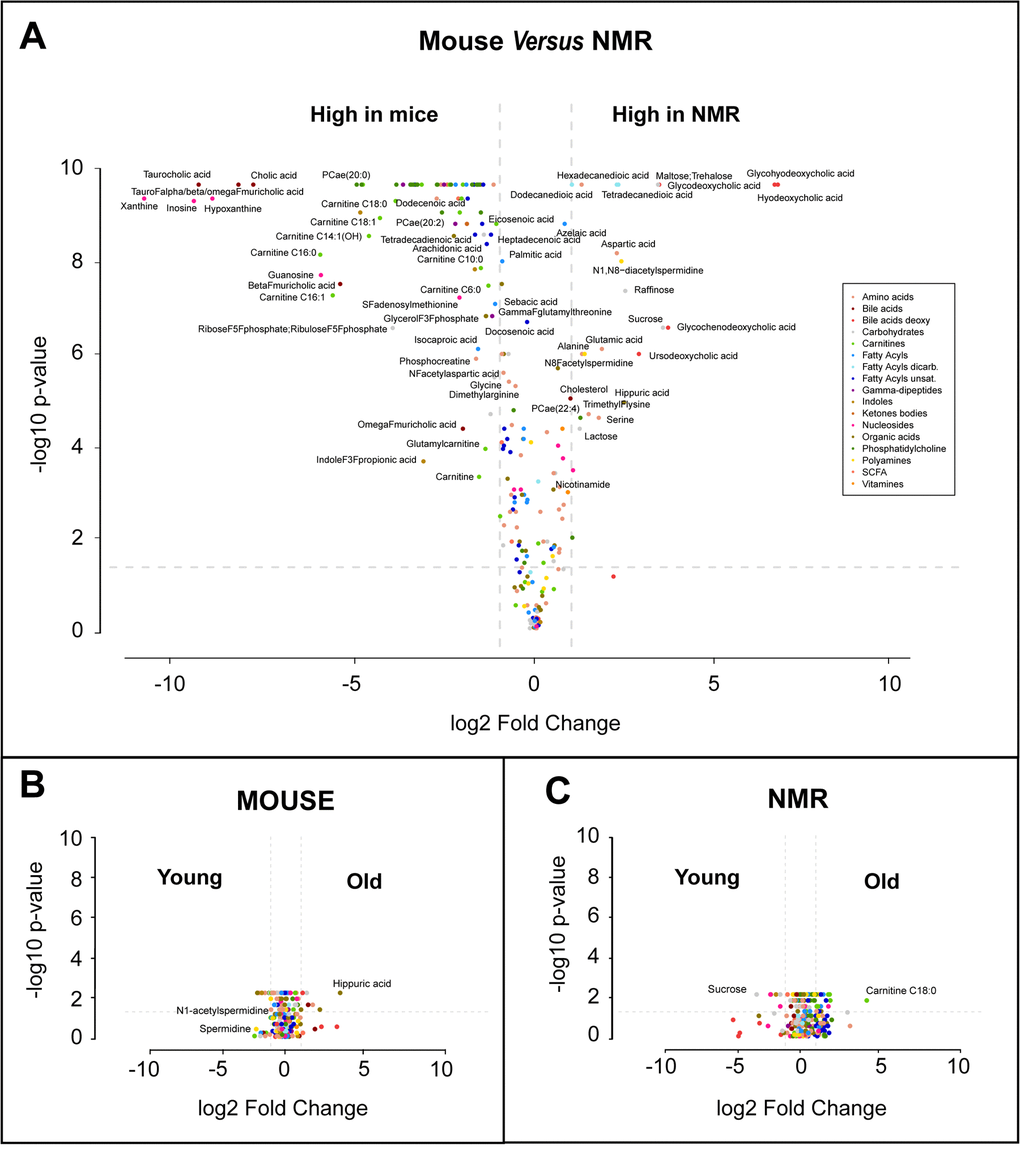
Figure 2. Volcano plots of metabolome differences. (A) Interspecies comparison. (B) Comparison between young (1-1.5 months) and old (20 months) mice. (C) Comparison between young (1 year) and old (10 months) naked mole-rats (NMR). The color code classifying different metabolic species used in A is also used in B and C. Selected metabolites are indicated.
Metabolites reduced in naked mole-rats
A large number of diverse circulating phosphatidyl cholines (abbreviated PCae) with distinct acyl chains (length 14 to 20) and unsaturation levels (from 0 to 4) were reduced in naked mole-rats. In addition, multiple unsaturated free fatty acids (e.g. arachidonic, dodecenoic, docosahexaenoic, heptadecanoic, linoleic, nonadecanoic, oleic, palimitoleic, tetradecadienoic acids) and carnitine-acyl esters (length 2-14) were decreased, pointing to a major alteration of lipid metabolism. The reduction of lipids and in particular, carnitine-acyl esters, that occurs in naked mole-rats might indicate efficient beta-oxidation. In line with a possible alteration of lipid metabolism, the major ketone body 3-hydroxybutyrate and two ketogenesis-associated metabolites, 2-hydroxy-2-methylbutyric acid and 2-hydroxy-3-methylbutyric acid, were depleted in the long-lived species. Furthermore, two primary bile acids (cholic and muricholic acids) and two tauro-conjugated bile acids (taurocholic, tauromuricholic acids) were strongly diminished. Several nucleic acid-relevant metabolites (guanosine, hypoxanthine, inosine, ribose-5-phosphate, and xanthine) were also reduced in the plasma from naked mole-rats compared to that of mice (Figure 1 and Figure 2a).
Several gamma-glutamyl amino acids (gamma-glutamyl leucine, gamma-glutamyl lysine, gamma-glutamyl threonine, and gamma-glutamyl tyrosine), which are proteolytic breakdown products were underrepresented in the plasma from naked mole-rats, in line with a prior report [15], suggesting a major reduction in protein turnover or an improved clearance of these metabolites (Figure 3a-d). The lysine degradation product 5-aminovaleric acid present also a reduced abundance in NMR compared to mice (Figure 3e). This metabolite is correlated positively with breast cancer risk in women [16]. Hydroxyphenyllactic acid, a tyrosine metabolite that increases with weight loss in obese women [17] and that correlates with ovarian cancer recurrence after surgery [18], was also decreased in NMR (Figure 3f). Furthermore, the tryptophan metabolite 3-Indole propionic acid (a bacterial metabolite), was diminished (Figure 3g). The same applies to taurine, a cysteine metabolite (Figure 3h), and dimethylglycine (Figure 3i), which increases with methionine restriction in mice [19]. However, methionine tended to be relatively lower in the long-lived species (Figure 3j), supporting the hypothesis [15] that long lived naked mole-rats bear characteristics of a methionine-restricted metabolism.
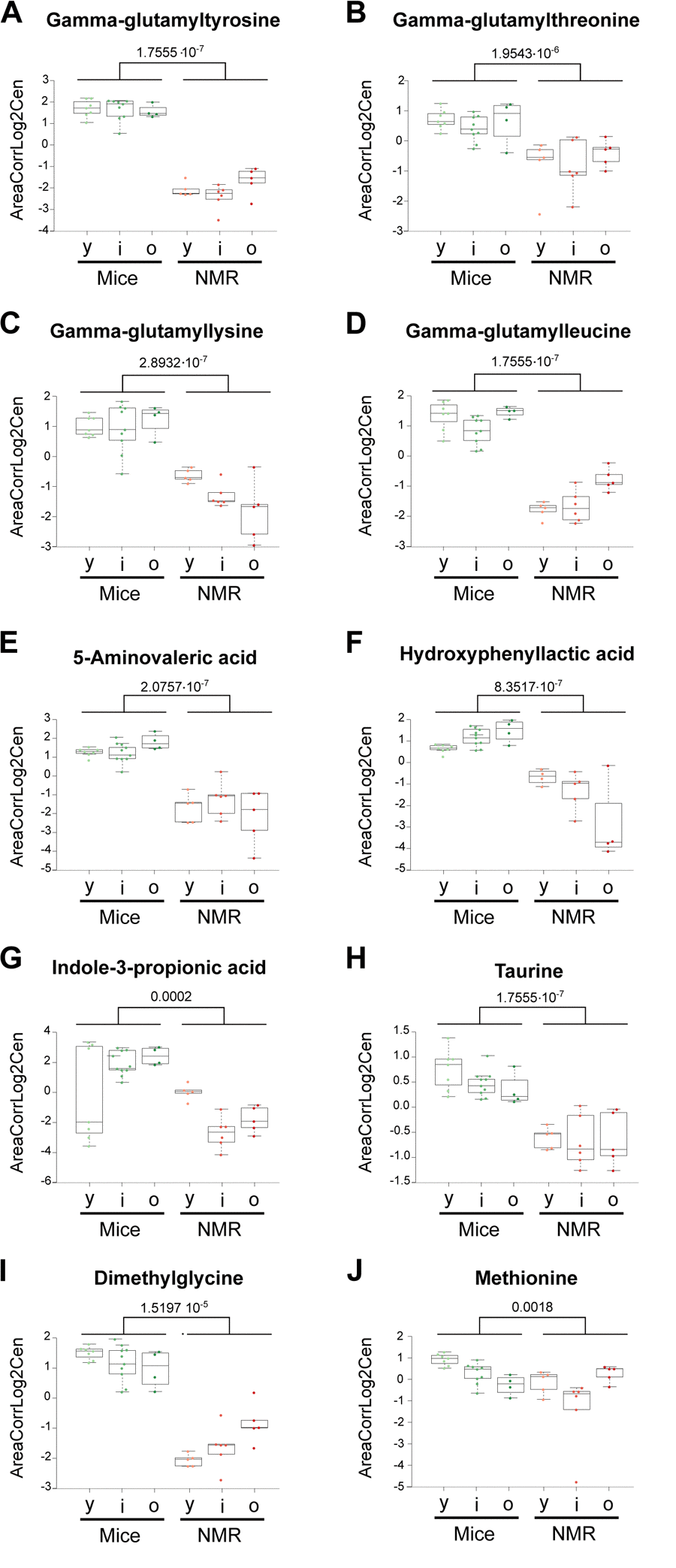
Figure 3. Amino acid derivatives that are reduced in naked mole-rats. Statistical comparisons were calculated by means of a two-sided Wilcoxon test. P-values are indicated.
Of note, sphingosine-1-phosphates (S1P) was reduced in naked mole-rats compared to mice (Figure 4a), a finding that appears interesting because in humans, S1P levels inversely associate with arteriosclerosis [20], but increase in prostate cancer [21], and predict osteoporotic fractures in postmenopausal women [22]. Thus, S1P seems a biomarker of several health-related parameters. Other metabolites that were reduced in the long-living species were glycerol-3-phosphate, indole-3-acrylic acid (a plant auxin), phosphocreatine (Figure 1), S-adenosylmethione (the methyl group donor for methylation reactions) (Figure 4b), as well oxidized glutathione, pleading in favor of a strong antioxidant system [23]. Indeed, the ratio of reduced over oxidized glutathione was significantly higher in naked mole-rats than in mice (Figure 4c).
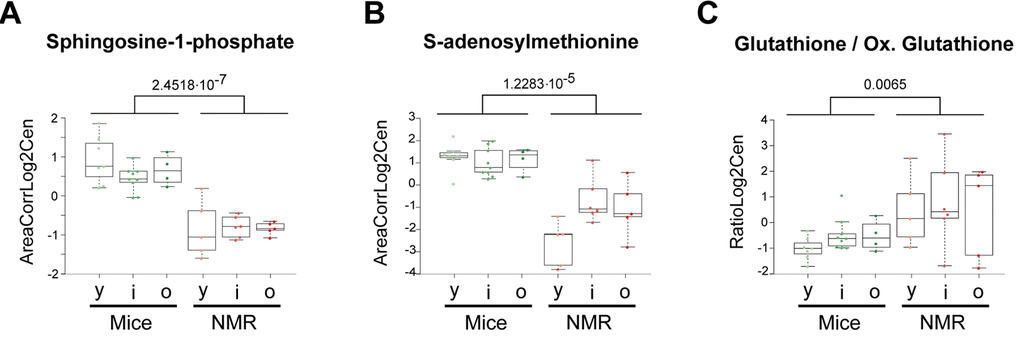
Figure 4. Selected metabolic alterations in naked mole-rats. Statistical comparisons were calculated by means of a two-sided Wilcoxon test. P-values are indicated.
Thus, a large panel of metabolites falling into distinct functional categories is comparatively low-abundant in naked mole-rats than in mice.
Metabolites increased in naked mole-rats
Opposite to the multiple abovementioned phosphatidyl cholines, PCae (22:4) was increased in naked mole-rats. Three aliphatic even-number medium-chain (12, 14 or 16 carbons) alpha-omega dicarboxylic acids (dodecanedioic, tetradecanedioic and heptadecatrienoic acids) were increased in NMR. These metabolites can be endogenously generated by omega oxidation of monocarboxylic acids or stem from vegetables [24]. Among these, dodecanedioic acid has been shown to improve glycemic control and to reduce muscle fatigue in type-2 diabetes patients, suggesting that it can improve metabolic flexibility [25,26]. In contrast, hexadecanedioic acid plasma levels are associated with high blood pressure in patients, and oral supplementation of hexadecanedioic acid causes hypertension in rats [27] (Figure 1).
Phosphocholine and cholesterol were enhanced in the long-lived rodent (Figure 1). Several bile acids were also increased, as this applies to one primary bile acid (chenodeoxycholic), three secondary bile acids (deoxycholic, hyodeoxycholic, ursodeoxycholic) and three glycoconjugated (glycodeoxycholic, glycochenodeoxycholic, glycohyodeoxycholic. This shift in circulating bile acids (Supplementary Figure 1) suggests species-differences in their metabolism, as this has been reported across mammalian species [28]. Bile acid supplementation has been shown to promote longevity in yeast [29,30], and supplementation with cholic acid increases longevity in short-lived progeroid mice [19]. This suggests a possible link between bile acid metabolism and the exceptional longevity of naked mole-rats.
Several free amino acids were particularly abundant in naked mole-rats compared to mice: alanine, aspartic acid, glutamic acid, isoleucine, leucine, serine and threonine (Figure 1, Figure 2a). Moreover, the dipeptide leucylproline and several amino acid derivatives were elevated: hexanoylglycine, S-adenosyl homocysteine and trimethyl lysine (Figure 5a-d). Interestingly, multiple sugars were more abundant in the plasma from the long-lived species, as this applies to lactose, maltose, raffinose and sucrose, as well as to the sugar alcohol xylitol. Two malonate derivatives, methylmalonic acid and malonylcarnitine were both overabundant in naked mole-rats. Lactate was increased, and so was adenosine, azelaic acid (generally considered as a fungal metabolite), creatinine, glyceric acid (a glycerol oxidation product) and hippuric acid (a biomarker of polyphenol uptake) (Figure 1, Figure 2a).
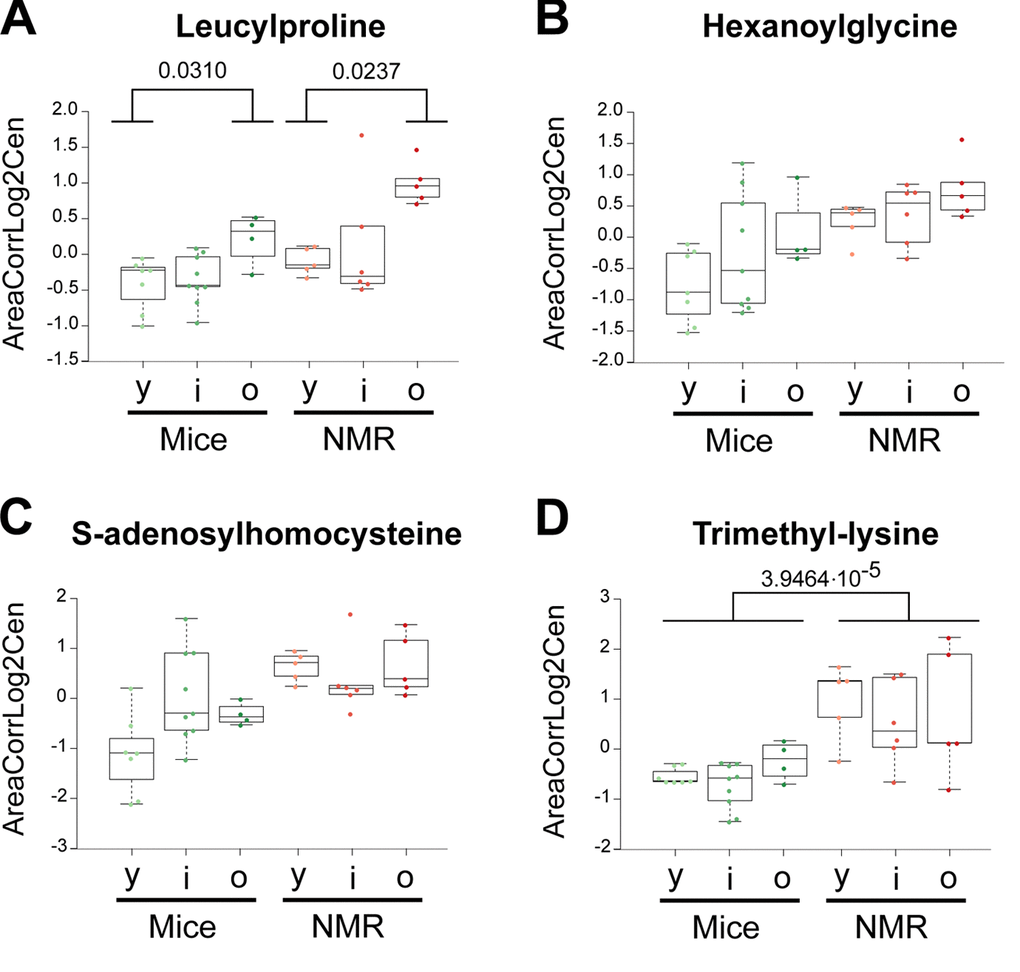
Figure 5. Amino acid derivatives that are elevated in naked mole-rats. Statistical comparisons were calculated by means of a two-sided Wilcoxon test. P-values are indicated.
The few metabolites that have been most convincingly linked to health-improving and antiaging effects in mice include two vitamins (B3, E) and polyamines. Nicotinamide (also called niacinamide, vitamin B3), which is known to extend health span and lifespan in mice [31,32], tend to increase in naked mole-rats and actually augmented with age in this species but not in mice (Figure 6a). The potent antioxidant α-tocopherol (vitamin E), which can enhance the lifespan of short-lived mouse strains [33,34], was also increased (Figure 6b). Several polyamines (cadaverine, N8-acetylspermidine, N1,N8-diacetylspermidine, but not ornithine nor putrescine) were elevated in the long-lived species (Figure 7). Importantly, spermidine and N1 acetylspermidine declined in aging mice, but remained at high levels in aging naked mole-rats (Figure 7c, d), echoing an abundant literature showing that spermidine supplementation promotes longevity in mice and other model organisms [35–38] and that increased nutrient uptake of spermidine reduces cardiovascular and cancer-related mortality in humans [39–41].
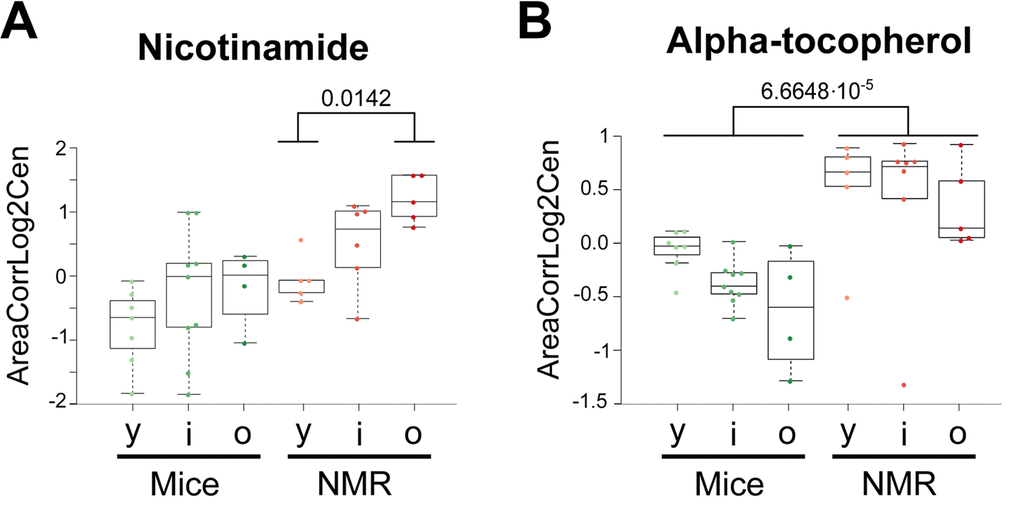
Figure 6. Vitamins that are elevated in naked mole-rats. Statistical comparisons were calculated by means of a two-sided Wilcoxon test. P-values are indicated.
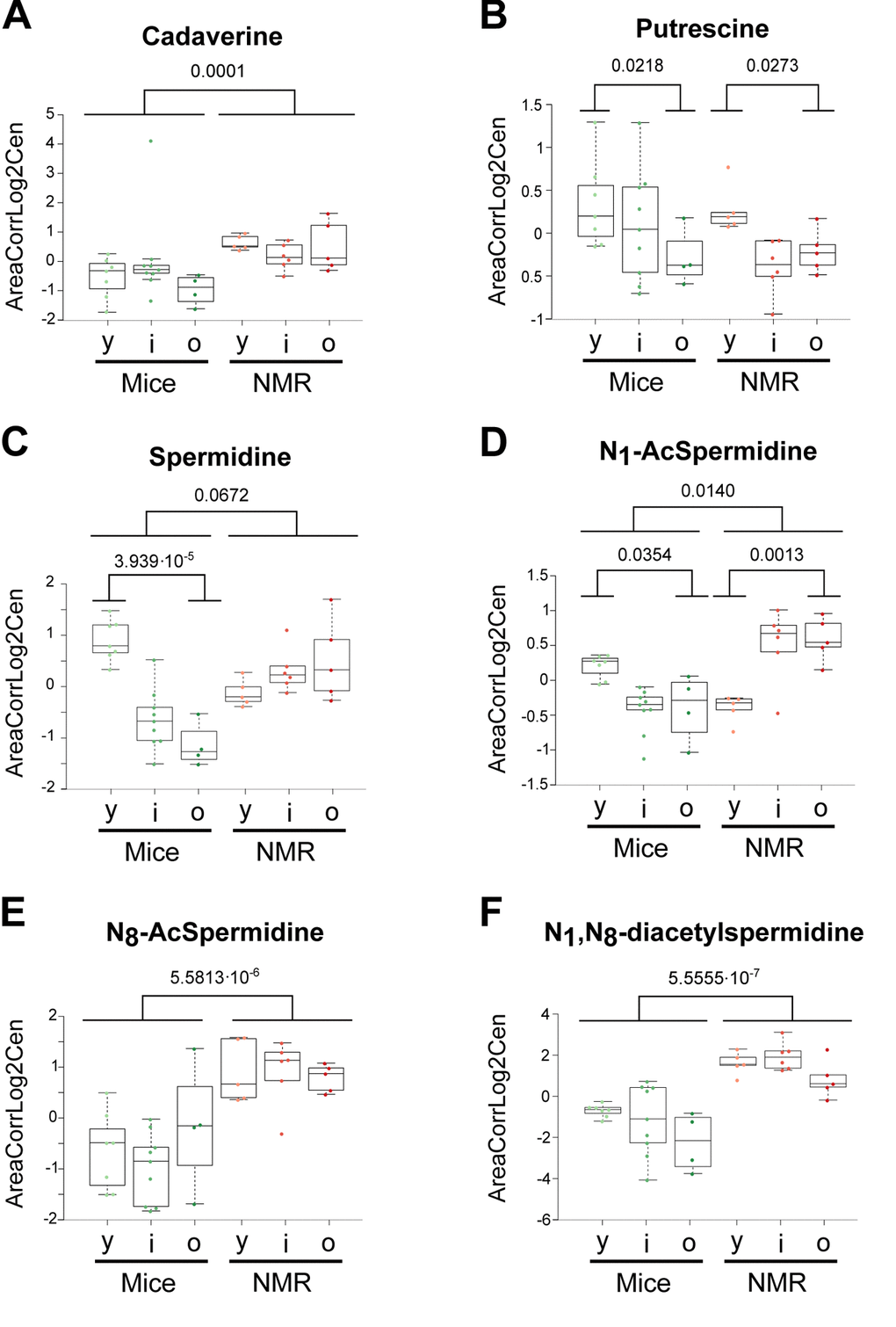
Figure 7. Influence of age and species differences on the abundance of polyamines and polyamine metabolites. Statistical comparisons were calculated by means of a two-sided Wilcoxon test. P-values are indicated.
Conclusions
Our present work reveals important differences in the metabolism between two species differing in their natural lifespan, namely, short-lived mice and long-lived naked mole-rats. These differences affect all major metabolic pathways, leading to alterations in the relative proportions of specific amino acid and their derivatives, free fatty acids and their carnitine esters, phosphatidyl cholines, bile acids, nucleic acids and their derivatives, protein degradation products and many other metabolites. A few differences may be hypothesis generating, as this applies to nicotinamide, α-tocopherol and polyamines which augmented in naked mole-rats, in the line with prior experiments showing that their continuous administration to mice can increase health span and longevity. That said, it will be necessary to inhibit the pathways involved in the intestinal absorption or synthesis of nicotinamide, α-tocopherol and polyamines in naked mole-rats and to consequently reduce their lifespan before a firm cause-effect-relationship between the accumulation of such ‘longevity molecules’ and the phenotype can be established.
Another difficulty inherent to the interpretation of the present results concerns the actual source of the longevity-associated molecules. For example, several molecules that are increased or reduced in naked mole-rats are most likely plant-derived: indole-3-acrylic acid and the saturated fatty acids with uneven-numbered carbon atoms (heptadecanoic and nonadecanoic acid), which all are diminished, and hippuric acid, which is augmented. Hippuric acid levels are known to rise with the consumption of phenolic compounds [42]. Thus, dietary differences in the two species (which do not receive the same chow) may dictate part of the discrepancies in the metabolome. Another molecule that is scarce in naked mole-rats is 3-Indole propionic acid, most likely a bacterial metabolite, contrasting with the fact that a fungal metabolite, azelaic acid, is overabundant. Hence, there may be major differences in the microbiota that explain some of the metabolic discrepancies found between the two species.
It is important to note that the longevity of the mammalian meta-organism composed by the host and is microbial communities is dictated by the systems property of this ecological unit. Thus, both mice and human show age-related shifts in the intestinal microbiota [43–45], and such shifts can actually contribute to the age-associated organismal decline [46], at least in the context of progeroid syndromes [47]. The body content of vitamins from the B series including nicotinamide as well as of polyamines and bile acids are strongly influenced by the intestinal microbiota (because bacteria are able to synthesize them), supporting the intriguing, yet-to-be-confirmed hypothesis that (some of) the longevity-associated metabolic alterations found in naked mole-rats actually reflect a particular gut flora [48]. Thus, one species that is highly abundant in the gut from naked mole-rats, Bacillus megaterium [49,50], is also particular efficient in polyamine biosynthesis [51,52]. These results call for an extensive functional characterization of the naked mole-rat microbiota with respect to its possible longevity-conferring properties. The polyamine spermidine is known to induce autophagy in multiple model organisms including mice [53–56], and autophagy induction accounts for its lifespan-extending effects [35–38]. Of note, autophagy reportedly is increased in naked mole-rats [57,58]. Hence, it will be tempting to investigate whether the depletion of polyamine (by inhibition of their biosynthesis by bacterial and host enzymes or by inhibition of their intestinal uptake) would reduce autophagic flux in naked mole-rats, thereby reducing the fitness of this species.
In synthesis, we identified several metabolites that may explain the exceptional longevity of naked mole-rats. Future studies are required to understand the mechanistic bases of their accumulation as well as their actual contribution to the suppression of the aging process.
Materials and Methods
Animal maintenance and experimental procedure
Animal experimental protocol was approved by the French National Ethical Committee ComEth Anses/ENVA/UPEC (authorization N°12114-2017110916247504-v3). Nineteen male naked mole-rats (1-year-old (n=6), 4 years old (n=6) and 10 years old (n=7)) and twenty-two C57BL6/J male mice (1-1.5 months old (n=7), 6-10 months old (n=10) and 20 months old(n=5)) were used in this study. Naked mole-rats were housed in Plexiglas cages interconnected with tubes to simulate burrows with tunnel systems. Naked mole-rats were kept in the dark, at a temperature of 28-30°C, and 70% humidity.
All animals were fasted overnight and all blood samples were collected in the morning (8-10 am). The blood was collected in heparinized tubes and centrifuged at 3000 rpm, for 10 min at 4°C. Plasma samples (30-50µl) were collected and stored (− 80 °C) until metabolomic analysis.
Standard and reagents
Acetonitrile (Sigma Aldrich); Isopropanol (Sigma Aldrich); Methanol (Sigma Aldrich); Chloroform (Sigma Aldrich); Acetic acid (Sigma Aldrich); Formic acid (Sigma Aldrich); Methoxyamine hydrochloride (Sigma Aldrich); MSTFA - N-Methyl-N-(trimethylsilyl) trifluoroacetamide (Sigma Aldrich); Pyridine (Sigma Aldrich); 3 nitrophenylhydrazine (Sigma Aldrich); N-(3-Dimethylaminopropyl)-N′-ethylcarbodiimide; hydrochloride (EDC) (Sigma Aldrich); Sulfosalicylic acid (Sigma Aldrich).
Sample preparation plasma (lithium heparin)
A volume of 25 µL of plasma were mixed with 250 µL a cold solvent mixture with ISTD (MeOH/Water/Chloroform, 9/1/1, -20°C), into 1.5 mL microtube, vortexed and centrifuged (10 min at 15000 g, 4°C) to obtain protein precipitation. Then upper phase of supernatant was split in three parts: 50 µL were used for GC-MS experiment in injection vial, 30 µL were used for the SCFA (Short Chain Fatty Acids) UHPLC-MS method, and 50 µL were used for others UHPLC-MS experimentations.
GC-MS aliquot was evaporated and 50 µL of methoxyamine (20 mg/mL in pyridine) were added on dried extracts, then stored at room temperature in dark, during 16 hours. The day after, 80 µL of MSTFA was added and final derivatization occurred at 40°C during 30 minutes. Samples were directly injected into GC-MS.
Concerning the UHPLC-MS aliquots (for SCFA method), 15 µl of 200mM 3-NPH and 15 µl of EDC (150mM) were added. The whole was heated at 40°C during 1h. 60 µl of H20 was added and the whole was injected into UHPLC-MS.
Concerning the LC-MS aliquots, the 50 µl collected supernatant was evaporated at 40°C in a pneumatically-assisted concentrator (Techne DB3, Staffordshire, UK). The LC-MS dried extracts were solubilized with 150 µL of MilliQ water. Samples were aliquoted for LC methods and backup. Biological samples and QC aliquots are kept at -80°C until injection or transferred in vials for direct analysis by UHPLC/MS.
Concerning the rest of the supernatant and the pellet, 90 µl of methanol with 2% of sulfosalicylic acid (SSA) was added before vortex and centrifugation (10 min at 15000g, 4°C). 130 µl of the supernatant were transferred in a microtube and evaporated. The dried sample were spiked with 100 µl of MilliQ water before injection in UHPLC/MS of the polyamines method.
Aliquots for analysis were transferred in polypropylene vials and injected into UHPLC-MS or kept at -80°C until injection.
Widely-targeted analysis of intracellular metabolites gas chromatography (GC) coupled to a triple quadrupole (QQQ) mass spectrometer
GC-MS/MS method was performed on a 7890B gas chromatography coupled to a triple quadrupole 7000C (Agilent Technologies, Waldbronn, Germany) equipped with a High sensitivity electronic impact source (EI) operating in positive mode.
Front inlet temperature was 250°C, injection was performed in splitless mode. Transfer line and ion-source temperature were 250°C and 230°C, respectively. Septum purge flow was fixed at 3 mL/min, purge flow to split vent operated at 80 mL/min during 1 min and gas saver mode was set to 15 mL/min after 5 min.
Helium gas flowed through column (J&WScientificHP-5MS, 30m x 0.25 mm, i.d. 0.25 mm, d.f., Agilent Technologies Inc.) at 1 mL/min. Column temperature was held at 60°C for 1 min, then raised to 210°C (10°C/min), followed by a step to 230°C (5°C/min) and reached 325°C (15°C/min), and be hold at this temperature for 5 min.
Collision gas was nitrogen. Scan mode used was MRM for biological samples. Peak detection and integration of analytes were performed using Agilent Mass Hunter quantitative software (B.07.01).
Targeted analysis of bile acids by ion pairing ultra-high-performance liquid chromatography (UHPLC) coupled to a Triple Quadrupole (QQQ) mass spectrometer
Targeted analysis was performed on a RRLC 1260 system coupled to a Triple Quadrupole 6410 (Agilent Technologies) equipped with an electrospray source operating in positive mode. Gas temperature was set to 325°C with a gas flow of 12 L/min. Capillary voltage was set to 4.5 kV.
10 μl of sample were injected on a Column Poroshell 120 EC-C8 (100 mm x 2.1 mm particle size 2.7 µm) from Agilent technologies, protected by a guard column XDB-C18 (5 mm × 2.1 mm particle size 1.8 μm) and heated at 40°C by a Pelletier oven.
Gradient mobile phase consisted of water with 0.2% of formic acid (A) and acetonitrile/isopropanol (1/1; v/v) (B) freshly made. Flow rate was set to 0.3 mL/min, and gradient as follow: initial condition was 70% phase A and 30% phase B, maintained during 1.5 min. Molecules were then eluted using a gradient from 30% to 60% phase B over 9 min. Column was washed using 98% mobile phase B for 2 minutes and equilibrated using 30% mobile phase B for 2 min. After each injection, needle was washed twice with isopropanol and thrice with water. Autosampler was kept at 4°C.
At the end of batch analysis, column was rinsed with 0.3 mL/min of MilliQ water (phase A) and acetonitrile (phase B) as follow: 10% phase B during 20 minutes, to 90% phase B in 20 minutes, and maintained during 20 minutes before shutdown.
Collision gas was nitrogen. Scan mode used was the MRM for biological samples. Peak detection and integration of the analytes were performed using the Agilent Mass Hunter quantitative software (B.07.01).
Targeted analysis of polyamines by ion pairing ultra-high performance liquid chromatography (UHPLC) coupled to a Triple Quadrupole (QQQ) mass spectrometer
Targeted analysis was performed on a RRLC 1260 system coupled to a Triple Quadrupole 6410 (Agilent Technologies) equipped with an electrospray source operating in positive mode. The gas temperature was set to 350°C with a gas flow of 12 l/min. The capillary voltage was set to 3.5 kV.
10 μl of sample were injected on a Column Kinetex C18 (150 mm x 2.1 mm particle size 2.6 µm) from Phenomenex, protected by a guard column C18 (5 mm × 2.1 mm) and heated at 40°C by a Pelletier oven. Heat the column more than the room temperature allowed rigorous control of the column temperature.
The gradient mobile phase consisted of water with 0.1% of Heptafluorobutyric acid (HFBA, Sigma-Aldrich) (A) and acetonitrile with 0.1% of HFBA (B) freshly made. The flow rate was set to 0.2 ml/min, and gradient as follows: initial condition was 95% phase A and 5% phase B. Molecules were then eluted using a gradient from 5% to 40% phase B over 10 min. The column was washed using 90% mobile phase B for 2.5 minutes and equilibrated using 5% mobile phase B for 4 min. The autosampler was kept at 4°C.
The collision gas was nitrogen. The scan mode used was the MRM for biological samples. Peak detection and integration of analytes were performed using the Agilent Mass Hunter quantitative software (B.07.01).
Targeted analysis of Short Chain Fatty Acid by ion pairing ultra-high performance liquid chromatography (UHPLC) coupled to a 6500+ QTRAP mass spectrometer
Targeted analysis was performed on a RRLC 1260 system (Agilent Technologies, Waldbronn, Germany) coupled to a 6500+ QTRAP (Sciex, Darmstadt, Germany) equipped with an electrospray ion source.
The instrument was operated using multiple reaction monitoring (MRM) in negative ion mode with unit resolution for both Q1 and Q3.
The optimized MS/MS conditions were: ion spray source temperature at 450°C, curtain (CUR) gas pressure at 40 psi, gas 1 (GS1) pressure at 30 psi and gas 2 (GS2) pressure at 70 psi. Ion-spray voltage (IS) was set at -4500V, collision-activated dissociation (CAD) at High, entrance potential (EP) at -10V and declustering potential (DP) at -80V.
10 μl of sample were injected on a Column Zorbax Eclipse XBD C18 (100 mm x 2.1 mm particle size 1.8 µm) from Agilent technologies, protected by a guard column XDB-C18 (5 mm × 2.1 mm particle size 1.8 μm) and heated at 50°C by a Pelletier oven.
Gradient mobile phase consisted of water with 0.01% of formic acid (A) and acetonitrile with 0.01% of formic acid (B). Flow rate was set to 0.4 mL/min, and gradient as follow: initial condition was 80% phase A and 20% phase B, maintained during 6 min. Molecules were then eluted using a gradient from 20% to 45% phase B over 7 min. Column was washed using 95% mobile phase B for 5 minutes and equilibrated using 20% mobile phase B for 4 min. The autosampler was kept at 4°C.
At the end of batch analysis, column was rinsed with 0.3 mL/min of MilliQ water (phase A) and acetonitrile (phase B) as follow: 10% phase B during 20 minutes, to 90% phase B in 20 minutes, and maintained during 20 minutes before shutdown.
The software used to operate the mass spectrometer was Analyst (Version 1.7). Peak detection, integration and quantification of the analytes were performed using MultiQuant quantitative software (Version 3.0.3).
Untargeted analysis of intracellular metabolites by ultra-high performance liquid chromatography (UHPLC) coupled to a Q-Exactive mass spectrometer. Reversed phase acetonitrile method
The profiling experiment was performed with a Dionex Ultimate 3000 UHPLC system (Thermo Scientific) coupled to a Q-Exactive (Thermo Scientific) equipped with an electrospray source operating in both positive and negative mode and full scan mode from 100 to 1200 m/z. The Q-Exactive parameters were: sheath gas flow rate 55 au, auxiliary gas flow rate 15 au, spray voltage 3.3 kV, capillary temperature 300°C, S-Lens RF level 55 V. The mass spectrometer was calibrated with sodium acetate solution dedicated to low mass calibration.
10 μL of sample were injected on a SB-Aq column (100 mm × 2.1 mm particle size 1.8 μm) from Agilent Technologies, protected by a guard column XDB-C18 (5 mm × 2.1 mm particle size 1.8 μm) and heated at 40°C by a Pelletier oven. The gradient mobile phase consists of water with 0.2% of acetic acid (A) and acetonitrile (B). The flow rate was set to 0.3 mL/min. Initial condition is 98% phase A and 2% phase B. Molecules were then eluted using a gradient from 2% to 95% phase B in 22 min. The column was washed using 95% mobile phase B for 2 minutes and equilibrated using 2% mobile phase B for 4 min.
The autosampler was kept at 4°C.
Peak detection and integration were performed using the Thermo Xcalibur quantitative software (3.1.).
Statistical methods
Raw data were processed and cleaned with R using the GRMeta package (located on Github/kroemerlab). Quality controls consisted in eliminating too low ion signal responses (signal to noise ratio less than 5) and biased metabolites (standard deviation of quality control pooled samples more than 26%). Kruskal-Wallis test with no adjustment were conducted on processed data with R. The heatmap was constructed with log2 normalized data centered around the average abundance from all samples. Metabolites were subjected to hierarchical clustering following the ward. D2 method and using the Euclidean distance. Volcano plots were constructed with a two-sided Wilcoxon to calculate p-values and subtractions of means (on a log2 scale) of the groups (Mice/NMR, and Old/Young).
Conflicts of Interest
GK has been holding research contracts with Bayer Healthcare, Glaxo Smyth Kline, Institut Mérieux, Lytix Pharma, Nucana, Oncolinx, PharmaMar, Sotio and Vasculox. GK is on the Board of Directors of the Bristol Myers Squibb Foundation France. GK is a scientific co-founder of everImmune and Samsara Therapeutics. FM is a scientific co-founder The Longevity Labs and Samsara Therapeutics.
Funding
GK is supported by the Ligue contre le Cancer (équipe labellisée); Agence National de la Recherche (ANR) – Projets blancs; ANR under the frame of E-Rare-2, the ERA-Net for Research on Rare Diseases; Association pour la Recherche sur le Cancer (ARC); Cancéropôle Ile-de-France; Chancelerie des universités de Paris (Legs Poix), Fondation pour la Recherche Médicale (FRM); a donation by Elior; European Research Area Network on Cardiovascular Diseases (ERA-CVD, MINOTAUR); Gustave Roussy Odyssea, the European Union Horizon 2020 Project Oncobiome; Fondation Carrefour; High-end Foreign Expert Program in China (GDW20171100085 and GDW20181100051), Institut National du Cancer (INCa); Inserm (HTE); Institut Universitaire de France; LeDucq Foundation; the LabEx Immuno-Oncology (ANR-18-IDEX-0001); the RHU Torino Lumière; the Seerave Foundation; the SIRIC Stratified Oncology Cell DNA Repair and Tumor Immune Elimination (SOCRATE); and the SIRIC Cancer Research and Personalized Medicine (CARPEM).
F.M. is grateful to the Austrian Science Fund FWF (SFB LIPOTOX F3007 & F3012, W1226, P29203, P29262, P27893, P 31727 and the Austrian Federal Ministry of Education, Science and Research and the University of Graz for grants “Unkonventionelle Forschung-InterFast” and “flysleep” (BMWFW-80.109/0001-WF/V/3b/2015) as well as the field of excellence program BioHealth. We acknowledge support from NAWI Graz and the BioTechMed-Graz flagship project “EPIAge”.
M.V. is grateful to the Ecole Nationale Vétérinaire d’Alfort (EnvA) and the Association De Prévoyance Santé (ADPS) partners of Allianz, for their invaluable help in establishing and funding the animal facility.
M.P.L. is supported by Fondation pour la Recherche Médicale (FRM FDT201904008383).
References
- 1. López-Otín C, Blasco MA, Partridge L, Serrano M, Kroemer G. The hallmarks of aging. Cell. 2013; 153:1194–217. https://doi.org/10.1016/j.cell.2013.05.039 [PubMed]
- 2. López-Otín C, Galluzzi L, Freije JM, Madeo F, Kroemer G. Metabolic Control of Longevity. Cell. 2016; 166:802–21. https://doi.org/10.1016/j.cell.2016.07.031 [PubMed]
- 3. Lopez-Otín C, Kroemer G. Decelerating ageing and biological clocks by autophagy. Nat Rev Mol Cell Biol. 2019; 20:385–86. https://doi.org/10.1038/s41580-019-0149-8 [PubMed]
- 4. Buffenstein R. Negligible senescence in the longest living rodent, the naked mole-rat: insights from a successfully aging species. J Comp Physiol B. 2008; 178:439–45. https://doi.org/10.1007/s00360-007-0237-5 [PubMed]
- 5. Lewis KN, Soifer I, Melamud E, Roy M, McIsaac RS, Hibbs M, Buffenstein R. Unraveling the message: insights into comparative genomics of the naked mole-rat. Mamm Genome. 2016; 27:259–78. https://doi.org/10.1007/s00335-016-9648-5 [PubMed]
- 6. Ruby JG, Smith M, Buffenstein R. Naked Mole-Rat mortality rates defy gompertzian laws by not increasing with age. eLife. 2018; 7:e31157.
- 7. Lagunas-Rangel FA, Chávez-Valencia V. Learning of nature: The curious case of the naked mole rat. Mech Ageing Dev. 2017; 164:76–81. https://doi.org/10.1016/j.mad.2017.04.010 [PubMed]
- 8. Grimes KM, Voorhees A, Chiao YA, Han HC, Lindsey ML, Buffenstein R. Cardiac function of the naked mole-rat: ecophysiological responses to working underground. Am J Physiol Heart Circ Physiol. 2014; 306:H730–7. https://doi.org/10.1152/ajpheart.00831.2013 [PubMed]
- 9. Edrey YH, Medina DX, Gaczynska M, Osmulski PA, Oddo S, Caccamo A, Buffenstein R. Amyloid beta and the longest-lived rodent: the naked mole-rat as a model for natural protection from Alzheimer’s disease. Neurobiol Aging. 2013; 34:2352–60. https://doi.org/10.1016/j.neurobiolaging.2013.03.032 [PubMed]
- 10. Edrey YH, Hanes M, Pinto M, Mele J, Buffenstein R. Successful aging and sustained good health in the naked mole rat: a long-lived mammalian model for biogerontology and biomedical research. ILAR J. 2011; 52:41–53. https://doi.org/10.1093/ilar.52.1.41 [PubMed]
- 11. O’Connor TP, Lee A, Jarvis JU, Buffenstein R. Prolonged longevity in naked mole-rats: age-related changes in metabolism, body composition and gastrointestinal function. Comp Biochem Physiol A Mol Integr Physiol. 2002; 133:835–42. https://doi.org/10.1016/S1095-6433(02)00198-8 [PubMed]
- 12. Lewis KN, Buffenstein R. The Naked Mole-Rat: A Resilient Rodent Model of Aging, Longevity, and Healthspan. A Resilient Rodent Model of Aging, Longevity, and Healthspan. Handbook of the Biology of Aging: Eighth Edition. 2015.
- 13. Pinto M, Jepsen KJ, Terranova CJ, Buffenstein R. Lack of sexual dimorphism in femora of the eusocial and hypogonadic naked mole-rat: a novel animal model for the study of delayed puberty on the skeletal system. Bone. 2010; 46:112–20. https://doi.org/10.1016/j.bone.2009.08.060 [PubMed]
- 14. Delaney MA, Ward JM, Walsh TF, Chinnadurai SK, Kerns K, Kinsel MJ, Treuting PM. Initial Case Reports of Cancer in Naked Mole-rats (Heterocephalus glaber). Vet Pathol. 2016; 53:691–96. https://doi.org/10.1177/0300985816630796 [PubMed]
- 15. Lewis KN, Rubinstein ND, Buffenstein R. A window into extreme longevity; the circulating metabolomic signature of the naked mole-rat, a mammal that shows negligible senescence. Geroscience. 2018.40:105–21. https://doi.org/10.1007/s11357-018-0014-2 [PubMed]
- 16. Lécuyer L, Dalle C, Lyan B, Demidem A, Rossary A, Vasson MP, Petera M, Lagree M, Ferreira T, Centeno D, Galan P, Hercberg S, Deschasaux M, et al. Plasma metabolomic signatures associated with long-term breast cancer risk in the SU.VI.MAX prospective cohort. Cancer Epidemiol Biomarkers Prev. 2019. https://doi.org/10.1158/1055-9965.EPI-19-0154 [PubMed]
- 17. Palau-Rodriguez M, Garcia-Aloy M, Miñarro A, Bernal-Lopez MR, Brunius C, Gómez-Huelgas R, Landberg R, Tinahones FJ, Andres-Lacueva C. Effects of a long-term lifestyle intervention on metabolically healthy women with obesity: metabolite profiles according to weight loss response. Clin Nutr. 2019; S0261-5614(19)30036-6/.
- 18. Zhang F, Zhang Y, Ke C, Li A, Wang W, Yang K, Liu H, Xie H, Deng K, Zhao W, Yang C, Lou G, Hou Y, Li K. Predicting ovarian cancer recurrence by plasma metabolic profiles before and after surgery. Metabolomics. 2018; 14:65. https://doi.org/10.1007/s11306-018-1354-8 [PubMed]
- 19. Bárcena C, Quirós PM, Durand S, Mayoral P, Rodríguez F, Caravia XM, Mariño G, Garabaya C, Fernández-García MT, Kroemer G, Freije JM, López-Otín C. Methionine Restriction Extends Lifespan in Progeroid Mice and Alters Lipid and Bile Acid Metabolism. Cell Rep. 2018; 24:2392–403. https://doi.org/10.1016/j.celrep.2018.07.089 [PubMed]
- 20. Soltau I, Mudersbach E, Geissen M, Schwedhelm E, Winkler MS, Geffken M, Peine S, Schoen G, Debus ES, Larena-Avellaneda A, Daum G. Serum-sphingosine-1-phosphate concentrations are inversely associated with atherosclerotic diseases in humans. PLoS One. 2016; 11:e0168302. https://doi.org/10.1371/journal.pone.0168302 [PubMed]
- 21. Nunes J, Naymark M, Sauer L, Muhammad A, Keun H, Sturge J, Stebbing J, Waxman J, Pchejetski D. Circulating sphingosine-1-phosphate and erythrocyte sphingosine kinase-1 activity as novel biomarkers for early prostate cancer detection. Br J Cancer. 2012; 106:909–15. https://doi.org/10.1038/bjc.2012.14 [PubMed]
- 22. Ardawi MM, Rouzi AA, Al-Senani NS, Qari MH, Elsamanoudy AZ, Mousa SA. High Plasma Sphingosine 1-phosphate Levels Predict Osteoporotic Fractures in Postmenopausal Women: The Center of Excellence for Osteoporosis Research Study. J Bone Metab. 2018; 25:87–98. https://doi.org/10.11005/jbm.2018.25.2.87 [PubMed]
- 23. Munro D, Baldy C, Pamenter ME, Treberg JR. The exceptional longevity of the naked mole-rat may be explained by mitochondrial antioxidant defenses. Aging Cell. 2019; 18:e12916. https://doi.org/10.1111/acel.12916 [PubMed]
- 24. Mingrone G, Castagneto-Gissey L, Macé K. Use of dicarboxylic acids in type 2 diabetes. Br J Clin Pharmacol. 2013; 75:671–76. https://doi.org/10.1111/j.1365-2125.2012.04177.x [PubMed]
- 25. Schild HO. Some aspects of receptor pharmacology of ergotamine. Postgrad Med J. 1976 (Suppl 1); 9–11. [PubMed]
- 26. Salinari S, Bertuzzi A, Gandolfi A, Greco AV, Scarfone A, Manco M, Mingrone G. Dodecanedioic acid overcomes metabolic inflexibility in type 2 diabetic subjects. Am J Physiol Endocrinol Metab. 2006; 291:E1051–58. https://doi.org/10.1152/ajpendo.00631.2005 [PubMed]
- 27. Menni C, Graham D, Kastenmüller G, Alharbi NH, Alsanosi SM, McBride M, Mangino M, Titcombe P, Shin SY, Psatha M, Geisendorfer T, Huber A, Peters A, et al. Metabolomic identification of a novel pathway of blood pressure regulation involving hexadecanedioate. Hypertension. 2015; 66:422–29. https://doi.org/10.1161/HYPERTENSIONAHA.115.05544 [PubMed]
- 28. García-Cañaveras JC, Donato MT, Castell JV, Lahoz A. Targeted profiling of circulating and hepatic bile acids in human, mouse, and rat using a UPLC-MRM-MS-validated method. J Lipid Res. 2012; 53:2231–41. https://doi.org/10.1194/jlr.D028803 [PubMed]
- 29. Goldberg AA, Richard VR, Kyryakov P, Bourque SD, Beach A, Burstein MT, Glebov A, Koupaki O, Boukh-Viner T, Gregg C, Juneau M, English AM, Thomas DY, Titorenko VI. Chemical genetic screen identifies lithocholic acid as an anti-aging compound that extends yeast chronological life span in a TOR-independent manner, by modulating housekeeping longevity assurance processes. Aging (Albany NY). 2010; 2:393–414. https://doi.org/10.18632/aging.100168 [PubMed]
- 30. Burstein MT, Kyryakov P, Beach A, Richard VR, Koupaki O, Gomez-Perez A, Leonov A, Levy S, Noohi F, Titorenko VI. Lithocholic acid extends longevity of chronologically aging yeast only if added at certain critical periods of their lifespan. Cell Cycle. 2012; 11:3443–62. https://doi.org/10.4161/cc.21754 [PubMed]
- 31. Mitchell SJ, Bernier M, Aon MA, Cortassa S, Kim EY, Fang EF, Palacios HH, Ali A, Navas-Enamorado I, Di Francesco A, Kaiser TA, Waltz TB, Zhang N, et al. Nicotinamide Improves Aspects of Healthspan, but Not Lifespan, in Mice. Cell Metab. 2018; 27:667–676.e4. https://doi.org/10.1016/j.cmet.2018.02.001 [PubMed]
- 32. Williams PA, Harder JM, Foxworth NE, Cochran KE, Philip VM, Porciatti V, Smithies O, John SW. Vitamin B3 modulates mitochondrial vulnerability and prevents glaucoma in aged mice. Science. 2017; 355:756–60. https://doi.org/10.1126/science.aal0092 [PubMed]
- 33. Nagai T, Yamada K, Kim HC, Kim YS, Noda Y, Imura A, Nabeshima YI, Nabeshima T. Nabeshima Y ichi, Nabeshima T. Cognition impairment in the genetic model of aging klotho gene mutant mice: a role of oxidative stress. FASEB J. 2003; 17:50–52. https://doi.org/10.1096/fj.02-0448fje
- 34. McDonald SR, Forster MJ. Lifelong vitamin E intake retards age-associated decline of spatial learning ability in apoE-deficient mice. Age (Dordr). 2005; 27:5–16. https://doi.org/10.1007/s11357-005-4003-x [PubMed]
- 35. Eisenberg T, Knauer H, Schauer A, Büttner S, Ruckenstuhl C, Carmona-Gutierrez D, Ring J, Schroeder S, Magnes C, Antonacci L, Fussi H, Deszcz L, Hartl R, et al. Induction of autophagy by spermidine promotes longevity. Nat Cell Biol. 2009; 11:1305–14. https://doi.org/10.1038/ncb1975 [PubMed]
- 36. Eisenberg T, Abdellatif M, Schroeder S, Primessnig U, Stekovic S, Pendl T, Harger A, Schipke J, Zimmermann A, Schmidt A, Tong M, Ruckenstuhl C, Dammbrueck C, et al. Cardioprotection and lifespan extension by the natural polyamine spermidine. Nat Med. 2016; 22:1428–38. https://doi.org/10.1038/nm.4222 [PubMed]
- 37. Madeo F, Eisenberg T, Pietrocola F, Kroemer G. Spermidine in health and disease. Science. 2018; 359:eaan2788. https://doi.org/10.1126/science.aan2788 [PubMed]
- 38. Madeo F, Carmona-Gutierrez D, Hofer SJ, Kroemer G. Caloric Restriction Mimetics against Age-Associated Disease: Targets, Mechanisms, and Therapeutic Potential. Cell Metab. 2019; 29:592–610. https://doi.org/10.1016/j.cmet.2019.01.018 [PubMed]
- 39. Kiechl S, Pechlaner R, Willeit P, Notdurfter M, Paulweber B, Willeit K, Werner P, Ruckenstuhl C, Iglseder B, Weger S, Mairhofer B, Gartner M, Kedenko L, et al. Higher spermidine intake is linked to lower mortality: a prospective population-based study. Am J Clin Nutr. 2018; 108:371–80. https://doi.org/10.1093/ajcn/nqy102 [PubMed]
- 40. Madeo F, Carmona-Gutierrez D, Kepp O, Kroemer G. Spermidine delays aging in humans. Aging (Albany NY). 2018; 10:2209–11. https://doi.org/10.18632/aging.101517 [PubMed]
- 41. Pietrocola F, Castoldi F, Kepp O, Carmona-Gutierrez D, Madeo F, Kroemer G. Spermidine reduces cancer-related mortality in humans. Autophagy. 2018; 15:362–65. https://doi.org/10.1080/15548627.2018.1539592 [PubMed]
- 42. Henning SM, Wang P, Abgaryan N, Vicinanza R, de Oliveira DM, Zhang Y, Lee RP, Carpenter CL, Aronson WJ, Heber D. Phenolic acid concentrations in plasma and urine from men consuming green or black tea and potential chemopreventive properties for colon cancer. Mol Nutr Food Res. 2013; 57:483–93. https://doi.org/10.1002/mnfr.201200646 [PubMed]
- 43. Peng W, Yi P, Yang J, Xu P, Wang Y, Zhang Z, Huang S, Wang Z, Zhang C. Association of gut microbiota composition and function with a senescence-accelerated mouse model of Alzheimer’s Disease using 16S rRNA gene and metagenomic sequencing analysis. Aging (Albany NY). 2018; 10:4054–65. https://doi.org/10.18632/aging.101693 [PubMed]
- 44. Pasolli E, Asnicar F, Manara S, Zolfo M, Karcher N, Armanini F, Beghini F, Manghi P, Tett A, Ghensi P, Collado MC, Rice BL, DuLong C, et al. Extensive Unexplored Human Microbiome Diversity Revealed by Over 150,000 Genomes from Metagenomes Spanning Age, Geography, and Lifestyle. Cell. 2019; 176:649–662.e20. https://doi.org/10.1016/j.cell.2019.01.001 [PubMed]
- 45. Stebegg M, Silva-Cayetano A, Innocentin S, Jenkins TP, Cantacessi C, Gilbert C, Linterman MA. Heterochronic faecal transplantation boosts gut germinal centres in aged mice. Nat Commun. 2019; 10:2443. https://doi.org/10.1038/s41467-019-10430-7 [PubMed]
- 46. Finlay BB, Pettersson S, Melby MK, Bosch TC. The Microbiome Mediates Environmental Effects on Aging. BioEssays. 2019; e1800257. https://doi.org/10.1002/bies.201800257 [PubMed]
- 47. Barcena C, Valdes-Mas R, Quiros P, Mayoral P, Durand S, Garatachea N, Rodriguez F, Garabaya C, Lucia A, Kroemer G, Freije J, Lopez-Otin C. Healthspan and lifespan extension by fecal microbiota transplantation in progeroid mice. Nat Med. 2019. Epub ahead of print. https://doi.org/10.1038/s41591-019-0504-5
- 48. Debebe T, Biagi E, Soverini M, Holtze S, Hildebrandt TB, Birkemeyer C, Wyohannis D, Lemma A, Brigidi P, Savkovic V, König B, Candela M, Birkenmeier G. Unraveling the gut microbiome of the long-lived naked mole-rat. Sci Rep. 2017; 7:9590. https://doi.org/10.1038/s41598-017-10287-0 [PubMed]
- 49. Debebe T, Holtze S, Morhart M, Hildebrandt TB, Rodewald S, Huse K, Platzer M, Wyohannes D, Yirga S, Lemma A, Thieme R, König B, Birkenmeier G. Analysis of cultivable microbiota and diet intake pattern of the long-lived naked mole-rat. Gut Pathog. 2016; 8:25. https://doi.org/10.1186/s13099-016-0107-3 [PubMed]
- 50. Cong W, Xing J, Feng Y, Wang J, Fu R, Yue B, He Z, Lin L, Yang W, Cheng J, Sun W, Cui S. The microbiota in the intestinal and respiratory tracts of naked mole-rats revealed by high-throughput sequencing. BMC Microbiol. 2018; 18:89. https://doi.org/10.1186/s12866-018-1226-4 [PubMed]
- 51. Setlow P. Spermidine biosynthesis during germination and subsequent vegetative growth of Bacillus megaterium spores. J Bacteriol. 1974; 120:311–15. [PubMed]
- 52. Zhou C, Ma Z, Zhu L, Xiao X, Xie Y, Zhu J, Wang J. Rhizobacterial strain Bacillus megaterium BOFC15 induces cellular polyamine changes that improve plant growth and drought resistance. Int J Mol Sci. 2016; 17:E976. https://doi.org/10.3390/ijms17060976 [PubMed]
- 53. Morselli E, Mariño G, Bennetzen MV, Eisenberg T, Megalou E, Schroeder S, Cabrera S, Bénit P, Rustin P, Criollo A, Kepp O, Galluzzi L, Shen S, et al. Spermidine and resveratrol induce autophagy by distinct pathways converging on the acetylproteome. J Cell Biol. 2011; 192:615–29. https://doi.org/10.1083/jcb.201008167 [PubMed]
- 54. Minois N, Carmona-Gutierrez D, Bauer MA, Rockenfeller P, Eisenberg T, Brandhorst S, Sigrist SJ, Kroemer G, Madeo F. Spermidine promotes stress resistance in Drosophila melanogaster through autophagy-dependent and -independent pathways. Cell Death Dis. 2012; 3:e401. https://doi.org/10.1038/cddis.2012.139 [PubMed]
- 55. Bauer MA, Carmona-Gutiérrez D, Ruckenstuhl C, Reisenbichler A, Megalou EV, Eisenberg T, Magnes C, Jungwirth H, Sinner FM, Pieber TR, Fröhlich KU, Kroemer G, Tavernarakis N, Madeo F. Spermidine promotes mating and fertilization efficiency in model organisms. Cell Cycle. 2013; 12:346–52. https://doi.org/10.4161/cc.23199 [PubMed]
- 56. Pietrocola F, Lachkar S, Enot DP, Niso-Santano M, Bravo-San Pedro JM, Sica V, Izzo V, Maiuri MC, Madeo F, Mariño G, Kroemer G. Spermidine induces autophagy by inhibiting the acetyltransferase EP300. Cell Death Differ. 2015; 22:509–16. https://doi.org/10.1038/cdd.2014.215 [PubMed]
- 57. Zhao S, Lin L, Kan G, Xu C, Tang Q, Yu C, Sun W, Cai L, Xu C, Cui S. High autophagy in the naked mole rat may play a significant role in maintaining good health. Cell Physiol Biochem. 2014; 33:321–32. https://doi.org/10.1159/000356672 [PubMed]
- 58. Triplett JC, Tramutola A, Swomley A, Kirk J, Grimes K, Lewis K, Orr M, Rodriguez K, Cai J, Klein JB, Perluigi M, Buffenstein R, Butterfield DA. Age-related changes in the proteostasis network in the brain of the naked mole-rat: implications promoting healthy longevity. Biochim Biophys Acta. 2015; 1852:2213–24. https://doi.org/10.1016/j.bbadis.2015.08.002 [PubMed]


