Introduction
Gastric cancer (GC) is one of the most aggressive human malignancies, representing the fourth most common cancer and the second leading cause of cancer-related mortality worldwide [1]. Although the diagnosis and therapy of GC have improved, progressive GC remains highly lethal because of its aggressive metastatic behavior and the fact that it is often diagnosed at an advanced stage [2, 3]. Therefore, understanding the molecular mechanisms underlying GC and discovering specific biomarkers to monitor its progression are of great clinical significance.
Long noncoding RNAs (lncRNAs) are transcribed RNA molecules longer than 200 nucleotides in length [4] that act as scaffolds to affect chromatin assembly and remodeling, gene transcription, and protein translation or stability [5, 6]. Multiple lines of evidence suggest lncRNAs have important roles in tumor biology, typically leading to abnormal expression of gene products involved in the progression of human cancers [7]. For instance, dysregulation of HOTAIR, one of the best studied lncRNAs, can lead to altered target gene expression and contribute to the invasiveness and metastasis of different cancers [8, 9]. Moreover, some lncRNAs can be specific tumor biomarkers. One example is PCA3, which has been approved by the FDA as a diagnostic biomarker in prostate cancer [10].
The lncRNA MIR4435-2HG (also known as AK001796 and LINC00978) is encoded on human chromosome 2q13. Acting as an oncogene, MIR4435-2HG was initially characterized by its involvement in resveratrol-mediated cell growth inhibition in lung cancer [11]. Subsequently, MIR4435-2HG expression was found to positively correlate with poor outcome in breast cancer patients [12]. Recent studies, on the other hand, reported that MIR4435-2HG promoted GC growth and suggested its relevance as a diagnostic marker [13, 14]. Nevertheless, the molecular mechanisms linking MIR4435-2HG and GC remain largely undefined.
Using lncRNA expression microarray screening, RNA pull-down assays, and functional in vitro and in vivo studies, we characterized a novel interaction between MIR4435-2HG and desmoplakin (DSP), a desmosome-associated protein, in human GC. Knockdown and overexpression experiments confirmed that increased MIR4435-2HG expression in GC inhibits DSP activity, promoting Wnt/β-catenin signaling and epithelial-mesenchymal transition (EMT). Thus, our study provides new insight into the molecular mechanisms by which MIR4435-2HG contributes to the pathogenesis of GC.
Results
MIR4435-2HG knockdown inhibits growth and invasion of GC cells in vitro
To investigate the role of MIR4435-2HG in GC, its expression was stably knocked down in two GC cell lines (SNU5 and HGC-27) by transfection with MIR4435-2HG-specific shRNA (shMIR4435-2HG). Successful knockdown was confirmed by RT-qPCR (Supplementary Figure 1B). Compared to negative controls (NC, i.e. cells transfected with scrambled shRNA), both SNU5-shMIR4435-2HG cells and HGC-27-shMIR4435-2HG cells showed lower proliferation rates in CCK-8 assays (Figure 2A). Likewise, colony formation capacity was also decreased in both cell lines after transfection with MIR4435-2HG shRNA (Figure 2B). Meanwhile, flow cytometry analyses showed a higher percentage of cells in G1 phase and increased apoptosis rates after MIR4435-2HG-knockdown (Figure 2C and 2D). These findings indicate that down-regulation of MIR4435-2HG can arrest cells in G1 phase and promote apoptosis, effectively decreasing proliferation.
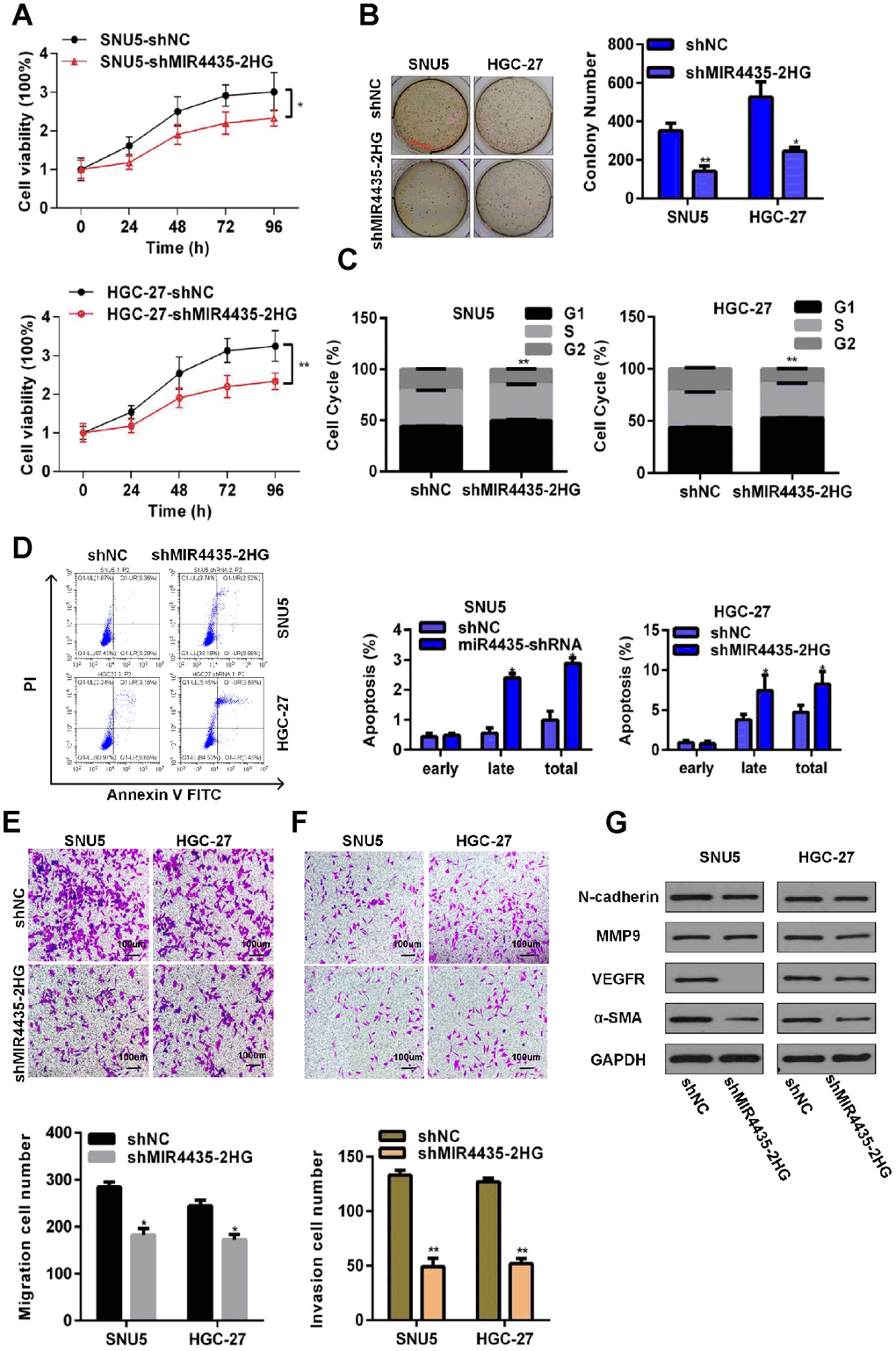
Figure 2. Depletion of MIR4435-2HG in GC cells inhibits cell growth and metastasis in vitro. (A) MIR4435-2HG knockdown inhibited cell proliferation (CCK8 assays) (*P < 0.05, **P < 0.01). (B) Colony formation assays were used to evaluate SNU5 and HGC-27 cell proliferation after inhibiting expression of MIR4435-2HG (*P < 0.05, **P < 0.01). (C) The cell cycle was analyzed using flow cytometry after knocking down MIR4435-2HG (**P < 0.01). (D) Flow cytometric apoptosis assays were used to analyze the incidence of apoptosis after silencing MIR4435-2HG (*P < 0.05). (E) MIR4435-2HG inhibition decreased cell migration in transwell assays (*P < 0.05). (F) Transwell assays used to assess the invasiveness of cells with downregulated MIR4435-2HG (**P < 0.01). (G) Expression of N-cadherin, MMP-9, VEGF and α-SMA in SNU5 and HGC-27 cells was detected by WB after decreasing MIR4435-2HG expression.
To evaluate whether down-regulation of MIR4435-2HG would affect the migration and invasion of SNU5 and HGC-27 cells, we conducted transwell assays. Results showed that MIR4435-2HG suppression attenuated migration and invasion in both cell lines (Figure 2E and 2F). To investigate the molecular bases of this inhibition, the expression of EMT-related proteins was examined by western blotting (WB). Markedly decreased expression of N-cadherin, VEGF, and α-SMA was observed in sh-MIR4435-2HG-transfected SNU5 (Figure 2G). Meanwhile, significantly decreased expression of MMP9, VEGF, and α-SMA was observed in sh-MIR4435-2HG-transfected HGC27 (Figure 2G). These findings suggest that MIR4435-2HG promotes migration and invasion of GC cells by inducing EMT.
MIR4435-2HG promotes GC cell tumorigenesis in vivo
To examine the effect of MIR4435-2HG on GC proliferation in vivo, HGC-27 cells stably transfected with sh-MIR4435 or scrambled control vectors (verified by RT-qPCR; Supplementary Figure 1C) were inoculated into nude mice. Tumor volumes were measured every week after injection. By week 6, tumor weights and volumes were significantly smaller in mice injected with HGC-27/sh-MIR4435-2HG cells (Figure 3A–3C).
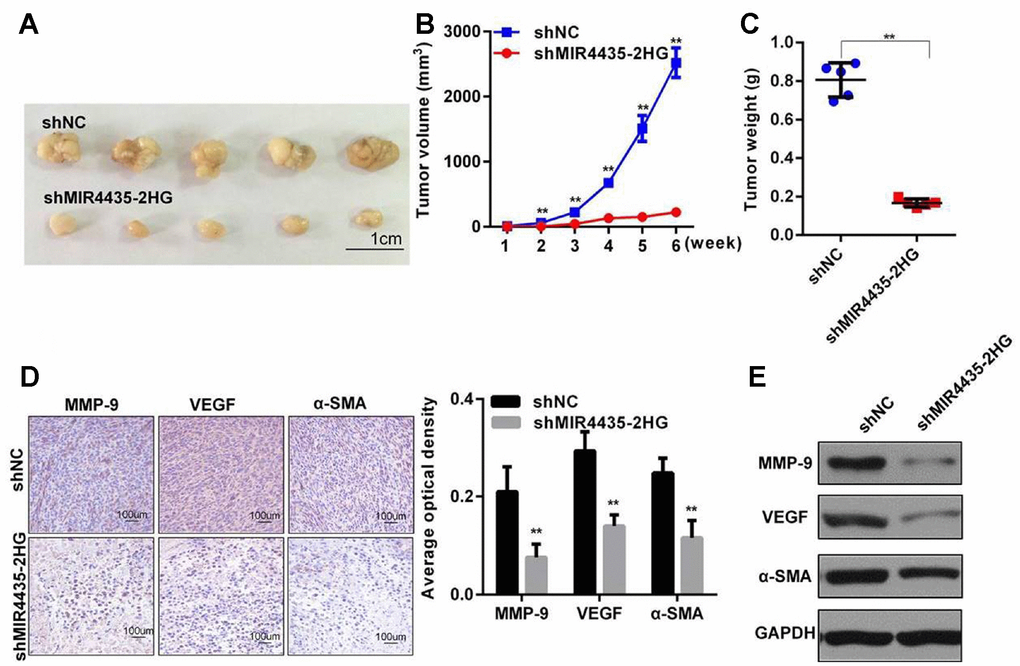
Figure 3. MIR4435-2HG promotes gastric cancer cell tumorigenesis in vivo. (A) MIR4435-2HG knockdown inhibited subcutaneous tumor formation in nude mouse models. Compared to the HGC-27/shNC group, the HGC-27/shMIR4435-2HG group showed smaller tumor sizes (n=5). (B) Quantitative comparison was made between the shNC and shMIR4435-2HG groups at weekly intervals using t-tests. Data are means ± SD for five samples (**P < 0.01). (C) Individual tumor weights from mice in the two groups (**P < 0.01). (D) Expression of N-Cadherin, MMP-9, VEGF and α-SMA in the two groups was detected using IHC. (E) Expression of MMP-9, VEGF and α-SMA in two groups were detected by WB and quantitated (**P < 0.01).
Furthermore, several proteins important for cancerous progression were detected in xenografted tumors by IHC and WB assays. Results showed that the expression of E-Cadherin, MMP-9, VEGF, and α-SMA in HGC-27/sh-MIR4435-2HG tumors was significantly lower than in the HGC-27/shNC group (Figure 3D and 3E). These results, which are in accord with our in vitro findings, strongly suggest that MIR4435-2HG contributes to the progression of GC in vivo.
MIR4435-2HG binds to DSP and inhibits its expression
Research showed that lncRNAs may regulate intracellular signaling through their interaction with RNA-binding proteins [15, 16]. Therefore, RNA-pull-down experiments were performed to search for MIR4435-2HG-interacting proteins. A specific band associated with biotinylated sense MIR4435-2HG was identified by silver staining after the pull-down assay (Figure 4A, red box). This band was cut out, digested, and subjected to mass spectrometry, which identified DSP, DCD, DSC1, HEL-S-270, HRNR, and JUP as MIR4435-2HG-interacting proteins (Figure 4B, Supplementary Figure 1D). Next, we examined the effect of MIR4435-2HG silencing on the expression of the above proteins by RT-qPCR. Results showed that DSP and HRNR were up-regulated in HGC-27 cells transfected with shMIR4435-2HG (Figure 4C). Furthermore, we verified the interaction of DSP with MIR4435-2HG or HRNR with MIR4435-2HG by RNA pull-down assay; DSP was detected in sense, but not antisense, MIR4435-2HG pull-down protein complexes (Figure 4D). In contrast, HRNR could not interact with MIR4435-2HG. Finally, we studied the interaction between MIR4435-2HG and DSP in HGC-27/shMIR4435-2HG and HGC-27/shNC xenografts by WB and IHC assays. Results demonstrated that the expression of DSP was inversely related to that of MIR4435-2HG (Figure 4E). These data suggest that MIR4435-2HG interacts with DSP, reducing its expression.
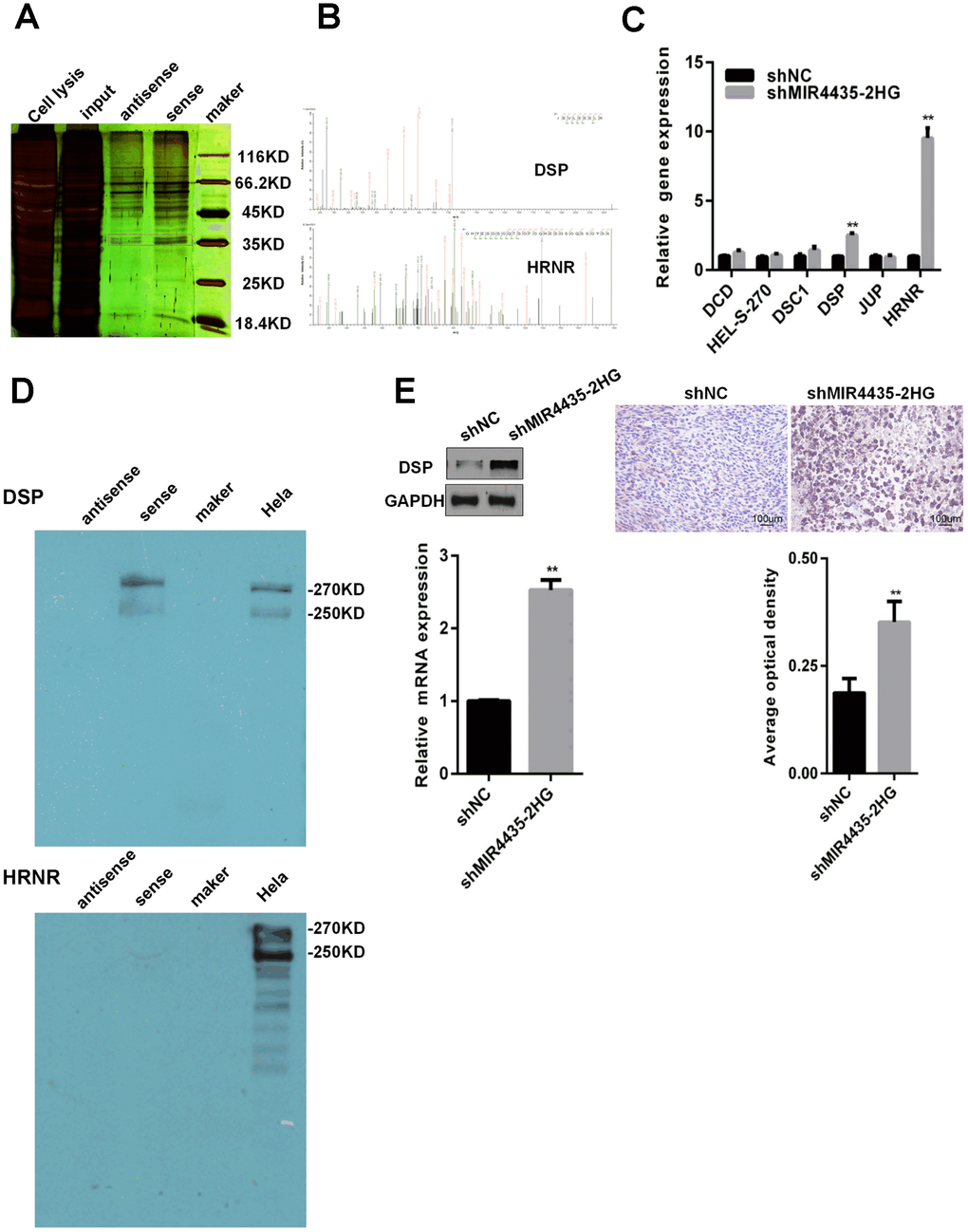
Figure 4. MIR4435-2HG binds to DSP protein and reduces DSP expression. (A) RNA pull-down experiments were performed to identify proteins interacting with MIR4435-2HG. Silver staining revealed a specific band associated with biotinylated sense MIR4435-2HG. (B) The biotinylated sense MIR4435-2HG band was excised, digested, and subjected to mass spectrometry, which identified DSP and HRNR as the MIR4435-2HG-interacting proteins. (C) Levels of DSP and HRNR were confirmed in the shMIR4435-2HG (HGC-27 cells) and control groups using RT-qPCR (**P < 0.01). (D) The interaction of DSP with MIR4435-2HG or HRNR with MIR4435-2HG by RNA pull-down assay. DSP was detected within sense MIR4435-2HG pull-down protein complexes but not with antisense MIR4435-2HG. (E) Expression levels of DSP within HGC-27/shMIR4435-2HG-xenografted and HGC-27/shNC-xenografted tumors were detected using WB and IHC (**P < 0.01).
DSP inhibits GC proliferation and metastasis through Wnt/β-catenin signaling inhibition
To clarify the effects of DSP on GC, we performed knockdown and overexpression assays in HGC-27 cells using siRNA (si-DSP/si-NC) or pCDNA3.1-DSP vector transfection (pCDNA-DSP/empty vector), respectively (Figure 5A). CCK8 assays showed that DSP knockdown and overexpression promoted and inhibited, respectively, the proliferation of HGC-27 cells (Figure 5B). Meanwhile, flow cytometry revealed that DSP down-regulation led to lower apoptotic rates, while DSP up-regulation promoted apoptosis of GC cells (Figure 5C). In transwell assays, DSP knockdown enhanced migration and invasion of HGC-27 cells, whereas DSP overexpression had the opposite effect (Figure 5D). These findings suggest that DSP expression restricts growth and metastasis of GC cells.
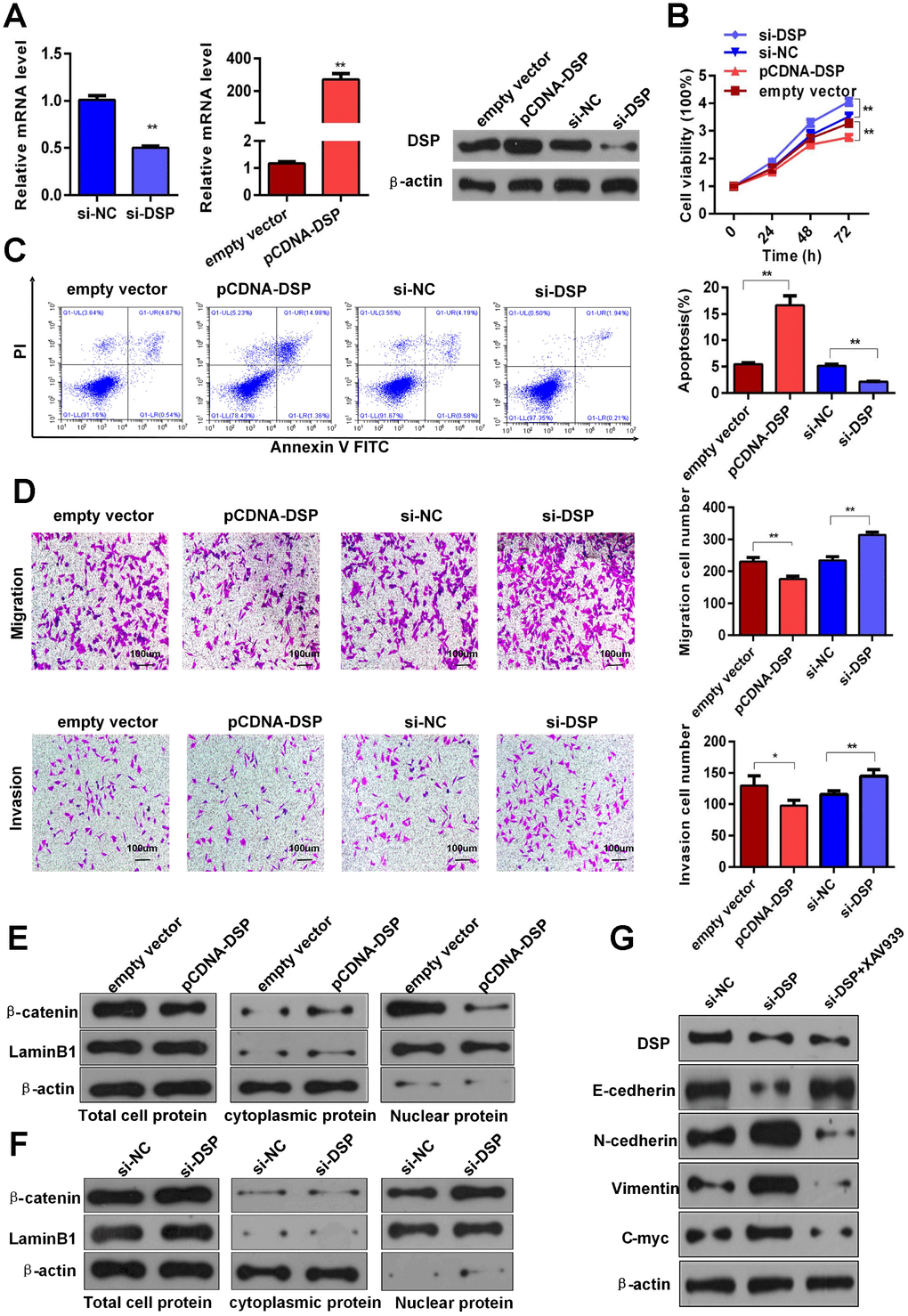
Figure 5. DSP inhibits GC growth and metastasis via Wnt/β-catenin signaling. (A) Depletion and overexpression of DSP were verified using RT-qPCR and WB. (B) CCK8 assays were performed after knockdown and overexpression of DSP (**P < 0.01). (C) Flow cytometric apoptosis assays were used to assess cell apoptosis after silencing and overexpression of DSP (**P < 0.01). (D) Transwell assays were used to determine the invasiveness of cells with down-regulated and up-regulated DSP (*P < 0.05, **P < 0.01). (E–F) Total, cytoplasmic and nuclear levels of β-catenin and lamin B1 were detected by WB after promoting or inhibiting DSP expression. (G) Expression levels of E-cadherin, N-cadherin, vimentin and c-myc were determined using WB after DSP knockdown or combined treatment with DSP siRNA and a Wnt/β-catenin inhibitor (XAV939a).
A previous study reported that DSP could function as a tumor suppressor through inhibition of the Wnt/β-catenin pathway in human lung cancer [17]. Considering the important role of Wnt/β-catenin pathway dysregulation in the progression of various tumors, including GC [18, 19], we next examined whether DSP could inhibit GC growth and metastasis by interfering with Wnt/β-catenin signaling. WB analyses revealed that total and nuclear expression of β-catenin was reduced in HGC-27 cells overexpressing DSP, while the expression of this protein in the cytoplasm was instead increased (Figure 5E). Contrast to these results, DSP knockdown increased both total and nuclear β-catenin levels (Figure 5F). Furthermore, we found evidence that DSP could inhibit EMT signaling. As shown by WB, DSP knockdown increased the expression of N-cadherin, vimentin, and c-Myc, and decreased E-cadherin expression. Accordingly, changes in N-cadherin, vimentin, and c-Myc were attenuated, and E-cadherin expression was rescued, after treatment with si-DSP and XAV939 (a Wnt/β-catenin attenuated, and E-cadherin expression was rescued, after treatment with si-DSP and XAV939 (a Wnt/β-catenin inhibitor) (Figure 5G). Meanwhile, the expressions of c-myc and β-catenin were obviously upregulated, whereas the level of DSP was obviously downregulated in GC specimens, compared with paired normal gastric tissues (Supplementary Figure 2A and 2B). Together, these data indicate that the capacity of DSP to inhibit GC growth and metastasis is in large part attributable to its ability to suppress Wnt/β-catenin signaling.
MIR4435-2HG regulates the Wnt/β-catenin signaling pathway through DSP to promote GC tumorigenesis
To test the hypothesis that MIR4435-2HG regulates the Wnt/β-catenin signaling through DSP to promote GC tumorigenesis and progression, we conducted rescue assays by co-transfecting shMIR4435-2HG and si-DSP into HGC-27 cells. As expected, CCK8 assays showed that DSP silencing partly rescued growth inhibition induced by shMIR4435-2HG (Figure 6A). In turn, apoptosis was reduced in cells co-transfected with shMIR44352HG and si-DSP, compared to cells transfected with shMIR4435-2HG or scrambled NC shRNA alone (Figure 6B and Supplementary Figure 1E). Similarly, partial restoration of migratory and invasive capacity was seen in MIR4435-2HG knockdown cells after si-DSP transfection. Under this condition, the number of migrating and invading cells was also lower in co-transfected with shMIR44352HG and si-DSP group than that transfected with shMIR4435-2HG (Figure 6C, 6D and Supplementary Figure 2C).
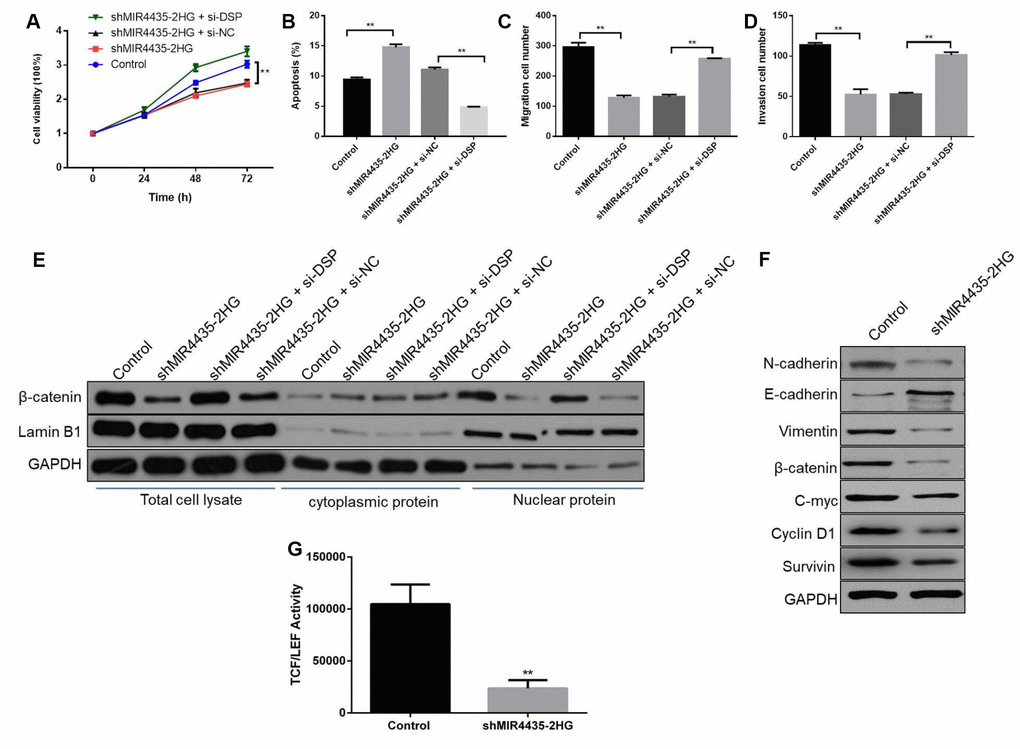
Figure 6. MIR4435-2HG regulates Wnt/β-catenin signaling through DSP to promote GC tumorigenesis and progression. (A) CCK8 assays were used with the HGC-27 cells co-transfected with shMIR4435-2HG and si-DSP (**P < 0.01). (B) The incidence of apoptosis among HGC-27 cells co-transfected with shMIR44352HG and si-DSP was determined using flow cytometry (**P < 0.01). (C, D) Transwell assays were used to assess cell migration and invasion in the group co-transfected with shMIR4435-2HG and si-DSP (**P < 0.01). (E) WB was performed to assess expression of β-catenin and lamin B1. (F) Levels of E-cadherin, N-cadherin, vimentin, c-Myc, cyclin D1, survivin and β-catenin within tumor tissue were determined using WB. (G) TCF/LEF transcriptional activity was determined by dual-luciferase assays in HGC-27 cells in response to MIR4435-2HG inhibition.
On the other hand, the expression of β-catenin was up-regulated in HGC-27 cells co-transfected with shMIR4435-2HG and si-DSP, compared with cells transfected with shMIR4435-2HG alone (Figure 6E). Moreover, knockdown of MIR4435-2HG increased E-cadherin and decreased N-cadherin, vimentin, c-Myc, β-catenin, cyclin D1 and survivin expression in tumor xenografts (Figure 6F and Supplementary Figure 2D). Meanwhile, the luciferase reporter assay revealed that inhibition of MIR4435-2HG markedly decreased the transactivating activity of β-catenin in HGC-27 cells (Figure 6G). In summary, these results indicate that MIR4435-2HG promotes GC growth and metastasis by activating Wnt/β-catenin signaling via DSP targeting.
Discussion
It is now widely established that mammalian genomes produce thousands of lncRNAs, some of which may contribute to carcinogenesis and metastasis [18, 20–22]. However, the normal functions of lncRNAs and the basis of their abnormal expression in tumors remain obscure. Through lncRNA expression microarray screening of clinical GC specimens, followed by expression and functional analyses in vitro and in vivo, our study identified the lncRNA MIR4435-2HG as a putative mediator of GC.
Up-regulation of MIR4435-2HG was observed in both human GC samples and GC cell lines, and was associated with more aggressive tumor invasion and late TNM stage. Accordingly, increased MIR4435-2HG expression was found to promote proliferation and restrict apoptosis of GC cells in vitro, and to stimulate tumorigenesis in mice. These results suggest the oncogenic potential of MIR4435-2HG on GC, consistent with a similar role reported for lung cancer, breast cancer, and esophageal squamous cell carcinoma [12, 23, 24]. Thus, MIR4435-2HG joins the list of several lncRNAs, including H19, HOTAIR, and GAPLINC, reported to influence GC development, progression, and metastasis [25–27].
One of the MIR4435-2HG-interacting proteins in our RNA pull-down assays, also verified by WB, was DSP, a founding member of the plakin family of proteins and an important component of desmosomal plaques [28]. Several studies suggested that loss of DSP is not uncommon in human tumors and may be an early step in carcinogenesis [29–32]. DSP has been reported to act as a tumor suppressor by inhibiting the Wnt/β-catenin signaling pathway in human lung cancer [17]. While dysregulated Wnt/β-catenin signaling is commonly observed in GC [18, 19, 33] a role for DSP in this malignancy has not been reported so far. Based on these data and our current findings, we speculated that MIR4435-2HG targets DSP to affect Wnt/β-catenin signaling and promote GC. Indeed, our results showed that MIR4435-2HG expression was closely associated with that of β-catenin and lamin B1, two key players in the activation of the WNT canonical pathway [34]. After ligation of WNT proteins to their respective cell surface receptors, β-catenin is released from the degradation complex and translocated into the cell nucleus to effect gene expression changes [35]. Our study indicated that up-regulation of DSP in GC cells decreased the expression of nuclear β-catenin, thus suppressing the activation of the Wnt/β-catenin pathway. Moreover, DSP knockdown led to decreased expression of N-cadherin, vimentin, and c-Myc, and increased expression of E-cadherin, suggesting that DSP inhibits the metastatic potential of GC cells by preventing EMT. Accordingly, we found that the EMT promoted by DSP knockdown was attenuated by a Wnt/β-catenin inhibitor. Altogether, these results revealed that up-regulation of the lncRNA MIR4435-2HG in GC cells reduces DSP activity, and this event stimulates the growth and metastasis of GC by inducing Wnt/β-catenin signaling and EMT. To our knowledge, this is the first study that reports the interaction between MIR4435-2HG and DSP, exposing a novel mechanism at play in the progression and metastasis of GC and perhaps other tumors.
Further studies are needed to validate MIR4435-2HG as a potential diagnostic or prognostic maker for GC, as well as its relevance as a therapeutic target.
Materials and Methods
Tissue samples and cell lines
GC specimens were obtained from patients of the First Affiliated Hospital of Zhejiang University, with confirmatory diagnoses based on histopathology. Informed consents prior to surgery were obtained for every patient. Human gastric cell lines (SNU5, HGC27 and SGC7901) and a normal gastric epithelial cell line (GES-1) were purchased from Shanghai Cell Bank of Chinese Academy of Sciences (Shanghai, China). HGC27, SGC7901, and GES-1 cells were cultured in RPMI 1640; SNU5 cells were cultured in IMDM medium with 10% fetal bovine serum (Gibco).
LncRNA expression microarray
The Affymetrix Human Gene 2.0 ST Array (Affymetrix, Santa Clara, CA, USA) was used to analyze lncRNA in four paired GC and normal gastric samples. Sample preparation and procedures were preformed based on the manufacturer’s standard protocols. Raw data was extracted by Feature Extraction software 10.7 (Agilent Technologies, Inc.) and normalized by Quantile algorithm using the GeneSpring GX v11.5 software package (Agilent Technologies). Genes with a fold change >2.0 or <0.5 (P < 0.05) were noted as statistically significant.
RNA extraction and quantitative reverse transcription PCR (RT-qPCR)
Procedures for RT-qPCR are described in Supplementary Methods. All primer sequences used in this study are shown in Supplementary Table 1.
Cell transfections
Plasmid vectors (pCDNA-DSP, sh-MIR4435-2HG, and empty vectors) and siRNA (si-DSP and scrambled negative control siRNA) were transfected into SNU5 or HGC-27 cells using Lipofectamine 2000 (Invitrogen, Carlsbad, California, USA) in Opti-MEM medium (Gibco, Carlsbad, California, USA) according to manufacturer’s instructions. The siRNA targeting DSP was synthesized by Invitrogen. All siRNA and shRNA sequences are shown in Supplementary Table 2.
Cell proliferation, flow cytometry, and transwell assay
Procedures for these assays are described in Supplementary Methods.
In vivo tumorigenesis
Ten male athymic BALB/c mice aged 3-4 weeks were purchased from Hubei Provincial Center for Disease Control and Prevention. All animal experiments were performed according to the protocols approved by the Institutional Animal Care and Use Committee at the Zhejiang University. For xenograft models, 4 × 106 HGC-27 cells transfected with sh-MIR4435 vector or sh-NC were injected subcutaneously in the right flank of mice (five mice per group). Tumor size was measured every 7 days using the equation V(mm3) = 0.5 × a × b2 (V, volume; a, length; b, width). After 6 weeks, the mice were sacrificed and tumors were weighed and processed for qPCR, immunohistochemistry, or western blot analyses.
Immunohistochemical (IHC) assay
Description of the IHC assay is provided in Supplementary Methods.
Western blot (WB) analyses
WB techniques are described in Supplementary Methods.
RNA pull-down assay and mass spectrometry analysis
The DNA templates of sense and antisense MIR4435 fragments used for transcription were cloned using PCR; primer sequences are listed in Supplementary Table 1. Sense and antisense MIR4435 strands were in vitro transcribed with the TranscriptAid T7 High Yield transcription Kit (Thermo Fisher Scientific, USA) and then labeled with desthiobiotin using Pierce RNA3′ End Desthiobiotinylation Kit (Thermo Fisher Scientific, USA). RNA pull-down was performed using the Pierce™ Magnetic RNA-Protein Pull-Down Kit (Thermo Fisher Scientific, USA) and protein bands of interest were analyzed with an UltiMate 3000 Rapid Separation LC (RSLC) system.
Luciferase reporter assay
HGC-27 cells were plated into 24-well plates and incubated for 24 h. The cells were co-transfected with wild-type TCF reporter plasmid TOP Flash (Upstate Biotechnology, Lake Placid, NY, USA) or mutated TCF reporter plasmid FOP flash (Upstate Biotechnology), and shMIR4435-2HG plasmid or NC using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) respectively. After 48 h of incubation, the relative luciferase activities in cell lysates were detected using a Dual-Luciferase Reporter Assay System (Promega, Madison, WI, USA).
Statistical analyses
All experiments were separately repeated 3 times. All data are expressed as the mean ± SD. Each bar represents the mean ± SD of 3 independent experiments. Statistical significance between two or multiple groups was analyzed by t-test or one-way ANOVA using GraphPad Prism 6.0. Statistical significance was assumed if P < 0.05.
Conflicts of Interest
The authors have no conflicts of interest to disclose.
Funding
This work was supported by the Zhejiang Province Science and Technology Bureau major research grant to Haiyong Wang (grant number 2018C03022, 2018); and grants to Lisong Teng from the National Natural Science foundation (grant number 81272676) and Zhejiang Provincial Science and Technology Project (grants numbers 2014C03040-2, 2011C23087).
References
- 1. Siegel RL, Miller KD, Jemal A. Cancer Statistics, 2017. CA Cancer J Clin. 2017; 67:7–30. https://doi.org/10.3322/caac.21387 [PubMed]
- 2. Cristescu R, Lee J, Nebozhyn M, Kim KM, Ting JC, Wong SS, Liu J, Yue YG, Wang J, Yu K, Ye XS, Do IG, Liu S, et al. Molecular analysis of gastric cancer identifies subtypes associated with distinct clinical outcomes. Nat Med. 2015; 21:449–56. https://doi.org/10.1038/nm.3850 [PubMed]
- 3. Rahman R, Asombang AW, Ibdah JA. Characteristics of gastric cancer in Asia. World J Gastroenterol. 2014; 20:4483–90. https://doi.org/10.3748/wjg.v20.i16.4483 [PubMed]
- 4. Gutschner T, Diederichs S. The hallmarks of cancer: a long non-coding RNA point of view. RNA Biol. 2012; 9:703–19. https://doi.org/10.4161/rna.20481 [PubMed]
- 5. Tsai MC, Manor O, Wan Y, Mosammaparast N, Wang JK, Lan F, Shi Y, Segal E, Chang HY. Long noncoding RNA as modular scaffold of histone modification complexes. Science. 2010; 329:689–93. https://doi.org/10.1126/science.1192002 [PubMed]
- 6. Nagano T, Fraser P. No-nonsense functions for long noncoding RNAs. Cell. 2011; 145:178–81. https://doi.org/10.1016/j.cell.2011.03.014 [PubMed]
- 7. Mercer TR, Dinger ME, Mattick JS. Long non-coding RNAs: insights into functions. Nat Rev Genet. 2009; 10:155–59. https://doi.org/10.1038/nrg2521 [PubMed]
- 8. Gupta RA, Shah N, Wang KC, Kim J, Horlings HM, Wong DJ, Tsai MC, Hung T, Argani P, Rinn JL, Wang Y, Brzoska P, Kong B, et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature. 2010; 464:1071–76. https://doi.org/10.1038/nature08975 [PubMed]
- 9. Liu XH, Sun M, Nie FQ, Ge YB, Zhang EB, Yin DD, Kong R, Xia R, Lu KH, Li JH, De W, Wang KM, Wang ZX. Lnc RNA HOTAIR functions as a competing endogenous RNA to regulate HER2 expression by sponging miR-331-3p in gastric cancer. Mol Cancer. 2014; 13:92. https://doi.org/10.1186/1476-4598-13-92 [PubMed]
- 10. Prensner JR, Rubin MA, Wei JT, Chinnaiyan AM. Beyond PSA: the next generation of prostate cancer biomarkers. Sci Transl Med. 2012; 4:127rv3. https://doi.org/10.1126/scitranslmed.3003180 [PubMed]
- 11. Liu B, Pan CF, Ma T, Wang J, Yao GL, Wei K, Chen YJ. Long non-coding RNA AK001796 contributes to cisplatin resistance of non-small cell lung cancer. Mol Med Rep. 2017; 16:4107–12. https://doi.org/10.3892/mmr.2017.7081 [PubMed]
- 12. Deng LL, Chi YY, Liu L, Huang NS, Wang L, Wu J. LINC00978 predicts poor prognosis in breast cancer patients. Sci Rep. 2016; 6:37936. https://doi.org/10.1038/srep37936 [PubMed]
- 13. Ke D, Li H, Zhang Y, An Y, Fu H, Fang X, Zheng X. The combination of circulating long noncoding RNAs AK001058, INHBA-AS1, MIR4435-2HG, and CEBPA-AS1 fragments in plasma serve as diagnostic markers for gastric cancer. Oncotarget. 2017; 8:21516–25. https://doi.org/10.18632/oncotarget.15628 [PubMed]
- 14. Fu M, Huang Z, Zang X, Pan L, Liang W, Chen J, Qian H, Xu W, Jiang P, Zhang X. Long noncoding RNA LINC00978 promotes cancer growth and acts as a diagnostic biomarker in gastric cancer. Cell Prolif. 2018; 51:51. https://doi.org/10.1111/cpr.12425 [PubMed]
- 15. Huarte M, Guttman M, Feldser D, Garber M, Koziol MJ, Kenzelmann-Broz D, Khalil AM, Zuk O, Amit I, Rabani M, Attardi LD, Regev A, Lander ES, et al. A large intergenic noncoding RNA induced by p53 mediates global gene repression in the p53 response. Cell. 2010; 142:409–19. https://doi.org/10.1016/j.cell.2010.06.040 [PubMed]
- 16. Rinn JL, Kertesz M, Wang JK, Squazzo SL, Xu X, Brugmann SA, Goodnough LH, Helms JA, Farnham PJ, Segal E, Chang HY. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell. 2007; 129:1311–23. https://doi.org/10.1016/j.cell.2007.05.022 [PubMed]
- 17. Yang L, Chen Y, Cui T, Knösel T, Zhang Q, Albring KF, Huber O, Petersen I. Desmoplakin acts as a tumor suppressor by inhibition of the Wnt/β-catenin signaling pathway in human lung cancer. Carcinogenesis. 2012; 33:1863–70. https://doi.org/10.1093/carcin/bgs226 [PubMed]
- 18. Li K, Dan Z. [Research progress of Wnt/β-catenin signaling pathway in prevention and treatment of gastric cancer]. Nan Fang Yi Ke Da Xue Xue Bao. 2014; 34:1852–56. [PubMed]
- 19. Clevers H, Nusse R. Wnt/β-catenin signaling and disease. Cell. 2012; 149:1192–205. https://doi.org/10.1016/j.cell.2012.05.012 [PubMed]
- 20. Guttman M, Amit I, Garber M, French C, Lin MF, Feldser D, Huarte M, Zuk O, Carey BW, Cassady JP, Cabili MN, Jaenisch R, Mikkelsen TS, et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature. 2009; 458:223–27. https://doi.org/10.1038/nature07672 [PubMed]
- 21. Yang X, Song JH, Cheng Y, Wu W, Bhagat T, Yu Y, Abraham JM, Ibrahim S, Ravich W, Roland BC, Khashab M, Singh VK, Shin EJ, et al. Long non-coding RNA HNF1A-AS1 regulates proliferation and migration in oesophageal adenocarcinoma cells. Gut. 2014; 63:881–90. https://doi.org/10.1136/gutjnl-2013-305266 [PubMed]
- 22. Ma Y, Yang Y, Wang F, Moyer MP, Wei Q, Zhang P, Yang Z, Liu W, Zhang H, Chen N, Wang H, Wang H, Qin H. Long non-coding RNA CCAL regulates colorectal cancer progression by activating Wnt/β-catenin signalling pathway via suppression of activator protein 2α. Gut. 2016; 65:1494–504. https://doi.org/10.1136/gutjnl-2014-308392 [PubMed]
- 23. Yang Q, Xu E, Dai J, Liu B, Han Z, Wu J, Zhang S, Peng B, Zhang Y, Jiang Y. A novel long noncoding RNA AK001796 acts as an oncogene and is involved in cell growth inhibition by resveratrol in lung cancer. Toxicol Appl Pharmacol. 2015; 285:79–88. https://doi.org/10.1016/j.taap.2015.04.003 [PubMed]
- 24. Liu B, Pan CF, Yao GL, Wei K, Xia Y, Chen YJ. The long non-coding RNA AK001796 contributes to tumor growth via regulating expression of p53 in esophageal squamous cell carcinoma. Cancer Cell Int. 2018; 18:38. https://doi.org/10.1186/s12935-018-0537-8 [PubMed]
- 25. Li H, Yu B, Li J, Su L, Yan M, Zhu Z, Liu B. Overexpression of lncRNA H19 enhances carcinogenesis and metastasis of gastric cancer. Oncotarget. 2014; 5:2318–29. https://doi.org/10.18632/oncotarget.1913 [PubMed]
- 26. Okugawa Y, Toiyama Y, Hur K, Toden S, Saigusa S, Tanaka K, Inoue Y, Mohri Y, Kusunoki M, Boland CR, Goel A. Metastasis-associated long non-coding RNA drives gastric cancer development and promotes peritoneal metastasis. Carcinogenesis. 2014; 35:2731–39. https://doi.org/10.1093/carcin/bgu200 [PubMed]
- 27. Hu Y, Wang J, Qian J, Kong X, Tang J, Wang Y, Chen H, Hong J, Zou W, Chen Y, Xu J, Fang JY. Long noncoding RNA GAPLINC regulates CD44-dependent cell invasiveness and associates with poor prognosis of gastric cancer. Cancer Res. 2014; 74:6890–902. https://doi.org/10.1158/0008-5472.CAN-14-0686 [PubMed]
- 28. Leung CL, Green KJ, Liem RK. Plakins: a family of versatile cytolinker proteins. Trends Cell Biol. 2002; 12:37–45. https://doi.org/10.1016/S0962-8924(01)02180-8 [PubMed]
- 29. Chun MG, Hanahan D. Genetic deletion of the desmosomal component desmoplakin promotes tumor microinvasion in a mouse model of pancreatic neuroendocrine carcinogenesis. PLoS Genet. 2010; 6:e1001120. https://doi.org/10.1371/journal.pgen.1001120 [PubMed]
- 30. Papagerakis S, Shabana AH, Pollock BH, Papagerakis P, Depondt J, Berdal A. Altered desmoplakin expression at transcriptional and protein levels provides prognostic information in human oropharyngeal cancer. Hum Pathol. 2009; 40:1320–29. https://doi.org/10.1016/j.humpath.2009.02.002 [PubMed]
- 31. Davies EL, Gee JM, Cochrane RA, Jiang WG, Sharma AK, Nicholson RI, Mansel RE. The immunohistochemical expression of desmoplakin and its role in vivo in the progression and metastasis of breast cancer. Eur J Cancer. 1999; 35:902–07. https://doi.org/10.1016/S0959-8049(99)00031-3 [PubMed]
- 32. Beaudry VG, Jiang D, Dusek RL, Park EJ, Knezevich S, Ridd K, Vogel H, Bastian BC, Attardi LD. Loss of the p53/p63 regulated desmosomal protein Perp promotes tumorigenesis. PLoS Genet. 2010; 6:e1001168. https://doi.org/10.1371/journal.pgen.1001168 [PubMed]
- 33. Hamada F. [Wnt signaling and cancer]. Kaibogaku Zasshi. 2009; 84:111–12. [PubMed]
- 34. Barker N, Clevers H. Catenins, Wnt signaling and cancer. BioEssays. 2000; 22:961–65. https://doi.org/10.1002/1521-1878(200011)22:11<961::AID-BIES1>3.0.CO;2-T [PubMed]
- 35. Tell S, Yi H, Jockovich ME, Murray TG, Hackam AS. The Wnt signaling pathway has tumor suppressor properties in retinoblastoma. Biochem Biophys Res Commun. 2006; 349:261–69. https://doi.org/10.1016/j.bbrc.2006.08.044 [PubMed]