Introduction
Telomeres are nucleoprotein complexes that cap chromosomes and, in most cells, shorten with each division. Leukocyte telomere length (LTL) is associated with age [1], and the rate of LTL attrition is more rapid during childhood (100-1000 bp/year) [2] than adulthood (20-100 bp/year) [3]. Biological variability in LTL is attributed to genetics [4], sex [5], ethnicity [6], and other determinants of health including diet [7], exercise [8], and substance use such as tobacco smoking [9, 10] and illicit drugs [11]. Shorter telomere length (TL) in leukocytes or peripheral blood mononuclear cells is also associated with human immunodeficiency virus (HIV) [12–15], diabetes [16], Alzheimer’s disease [17], and certain cancers [18, 19]. Among older individuals, TL is particularly predictive of cardiovascular disease [20–22] and mortality [23].
Mitochondria harbor multiple copies of mitochondrial DNA (mtDNA) per cell, which replicate independently from nuclear DNA. Changes in mtDNA quality or quantity, as well as decline in mitochondrial function, are implicated in biological aging [24] and are primarily linked to metabolic and mitochondrial disorders. Similarities also exist between whole blood (WB) mtDNA content per cell and LTL, whereby many diseases or conditions associated with shorter LTL are also associated with differences in mtDNA content. For example, decreased mtDNA content in various tissues is associated with HIV [25–27], diabetes [28], Alzheimer’s disease [29], various cancers [30, 31], and cardiovascular disease [32]. Tobacco smoking is also associated with altered mtDNA content in lung and buccal cells, though reports are inconsistent [33, 34]. Compared to LTL, which is consistently reported to decline with age, WB mtDNA content is not as robust a marker of aging, with studies having reported negative [24], positive [35, 36], non-linear [37, 38], or no relationships [39] between mtDNA content and age.
Several proposals have arisen connecting these two markers of cellular aging mechanistically. Short telomeres trigger p53 activation, which itself is associated with mitochondrial biogenesis [40]. It is also well recognized that telomerase translocates to and protects mitochondria in conditions of oxidative stress [41, 42]. Despite this, the relationship between LTL and WB mtDNA content is unclear based on cross-sectional studies [43–46] and has not yet been investigated longitudinally.
In the context of HIV, even people successfully treated with combination antiretroviral therapy (cART) have an accelerated aging phenotype compared to their HIV-negative peers [47]; they also have shorter LTL and abnormal mtDNA content. Despite a growing body of literature linking aging, HIV, and other diseases to either shorter LTL or altered WB mtDNA content individually, there is a paucity of research considering the two markers together, describing the relationship between the two, especially longitudinally. Given that HIV may provide a human model of biological aging, our goal was to characterize the relationship between LTL and WB mtDNA content in a cohort of girls and women living with and without HIV, both cross-sectionally and longitudinally.
Results
Study sample
Characteristics of the study participants are shown in Table 1. Cross-sectional analyses included 312 women living with HIV (WLWH) and 300 HIV-negative participants. WLWH were slightly younger, with a median (range) age of 41 (14-69) vs. 44 (15-78) years (P=0.042). WLWH were also more likely to be African/Caribbean/Black (ACB), current tobacco smokers, current prescribed opioid users, have past or present HBV or HCV infections, and less likely to be White, Asian, current drinkers, have a household income >15000 CAD/year, and have any college education compared to HIV-negative controls. Body mass index and cannabis use did not differ between groups. Among WLWH, 70% of participants had an undetectable HIV plasma viral load (pVL <50 HIV RNA copies/ml) and 82% were on cART at visit. Among all participants, the median[interquartile range] LTL and WB mtDNA content measurements were 7.2[6.5-7.9] and 105[81-139], respectively.
Table 1. Study sample characteristics.
| Cross-Sectional Sample (N=612) | Longitudinal Sample (N=296) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| HIV+ (N=312) | HIV- (N=300) | P-value | HIV+ (N=228) | HIV- (N=68) | P-value | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Age, years | 41 [31,50](14,69) | 44 [31,55](15,78) | 0.042 | 38 [29,46](12,67) | 36 [27,48](12,73) | 0.859 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Time between visits, years | 4 [2,6] (1,8) | 3 [2,5] (1,7) | < 0.001 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BMI, kg/m2 (N=603) | 24.9 [21.4,30.0] (15.0,48.6) | 24.6 [21.4,29.7] (14.0,52.9) | 0.608 | 24.0 [21.3,28.8] (16.0,46.6) | 23.5 [20.5,28.7] (15.3,42.3) | 0.526 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Ethnicity | < 0.001 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| White | 131 (42) | 153 (51) | 94 (41) | 37 (54) | 0.003 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| African/Black/Caribbean | 72 (23) | 19 (6) | 51 (22) | 3 (4) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Indigenous | 78 (25) | 79 (26) | 58 (25) | 16 (24) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Asian | 8 (3) | 28 (9) | 8 (4) | 7 (10) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Othera | 23 (7) | 21 (7) | 17 (7) | 5 (7) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Household income >$15000 CAD/year (N=553) | 127 (48) | 170 (59) | 0.009 | 95 (48) | 40 (67) | 0.011 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Highest level of education (N=552) Any College | 114 (43) | 200 (69) | < 0.001 | 93 (47) | 44 (73) | 0.005 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| High School Graduate | 54 (21) | 30 (10) | 38 (19) | 6 (10) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Some High School | 80 (30) | 53 (18) | 56 (28) | 9 (15) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Grade School | 15 (6) | 6 (2) | 11 (6) | 1 (2) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cigarette smokingb | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Current | 122 (39) | 85 (28) | 0.010 | 85 (37) | 13 (19) | 0.002 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Past | 69 (22) | 67 (22) | 54 (24) | 12 (18) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Never | 121 (39) | 148 (49) | 89 (39) | 43 (63) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cannabisb | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Current | 77 (25) | 72 (24) | 0.853 | 59 (26) | 15 (22) | 0.664 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Past | 67 (21) | 60 (20) | 49 (21) | 13 (19) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Never | 168 (54) | 168 (56) | 120 (53) | 40 (59) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alcoholb | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Current | 150 (48) | 199 (66) | < 0.001 | 113 (50) | 44 (65) | 0.039 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Past | 87 (28) | 61 (20) | 63 (28) | 17 (25) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Never | 75 (24) | 40 (13) | 52 (23) | 7 (10) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Current any opioid useb | 69 (22) 62 (20) | 32 (11) 25 (8) | < 0.001 | 51 (22) 47 (21) | 3 (4) 1 (1) | < 0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Prescribed opioid use | < 0.001 | < 0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Heroin use | 17 (5) | 13 (4) | 0.523 | 15 (7) | 0 (0) | 0.030 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| HIV Detectable pVL at visit, >50 copies/mlb,c | 95 (30) | 63 (28) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| HIV Peak pVL >100000 copies/ml (N=248) | 111 (45) | 87 (48) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CD4 count at visit, cells/μLb | 540 [348,733] (10,2380) | 500 [348,683] (14,1370) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CD4 nadir, cells/μL (N=260) | 221 [130,350] (1,1110) | 216 [118,330] (1,900) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ART-naïve (N=278) | 10 (4) | 11 (5) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| On cART at visitb | 255 (82) | 195 (86) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| HBV ever infected (N=395) | 29 (13) | 2 (1) | < 0.001 | 9 (26) | 0 (0) | 0.011 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| HCV ever infected | 109 (35) | 45 (15) | < 0.001 | 79 (35) | 9 (13) | < 0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Platelet Count, 109/L (N=429) | 227 [178,274] (40,663) | 235 [213,281] (82,464) | 0.005 | 233 [184,274] (45,489) | 272 [217,277] (172,316) | 0.240 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Relative LTL | 7.1 [6.4,7.8] (4.8,11.5) | 7.4 [6.7,8.1] (4.8,11.3) | 0.004 | 7.0 [6.3,7.8] (4.7,10.5) | 7.5 [6.9,8.4] (5.1,10.5) | < 0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ΔLTL/year | 0.004 | 0.01 [-0.10,0.11] (-0.94,1.80) | -0.05 [-0.26,0.09] (-1.18,0.74) | 0.020 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| WB mtDNA content | 101 [78,131] (4,265) | 112 [85,146] (4,379) | 143 [109,171] (4,277) | 129 [96,154] (4,231) | 0.025 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| WB ΔmtDNA content/year | -10 [-15,-1] (-76,77) | -7 [-16,8] (-32,37) | 0.013 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Data are number (%) of individuals or median [interquartile range] (range). Number of participants with available data indicated beside variables with incomplete data. P-values indicate Mann-Whitney U or Chi-Squared Tests depending on type of variable. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Abbreviations: ART, antiretroviral therapy; BMI, body mass index; cART, combination antiretroviral therapy; CAD, Canadian dollars; HBV, hepatitis B virus; HCV, hepatitis C virus; LTL, leukocyte telomere length; mtDNA, mitochondrial DNA; pVL, plasma viral load; WB, whole blood. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| a a combination of South Asian, Hispanic, or other | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| b at last visit for longitudinal sample | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| c 22 lost HIV viral control and 41 participants gained HIV viral control between longitudinal visits | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Cross-sectional predictors of LTL
Among all participants, the final multivariable model for LTL (N=612, R2=0.28) indicated that HIV infection (either with detectable or undetectable pVL), current tobacco smoking, and older age were independently associated with shorter LTL. In contrast, alcohol use and ACB or Asian ethnicities were associated with longer LTL (Figure 2A). Cannabis use showed no effect on LTL. In a subgroup of WLWH (Figure 2B), detectable HIV pVL was also independently associated with shorter LTL compared to undetectable HIV pVL (N=312, R2=0.31), in addition to the above factors. Among HIV-negative participants, current smoking remained associated (N=300, R2=0.28) with shorter LTL, but current alcohol did not (Figure 2C). Models showing non-standardized effect sizes are shown in supplement (Supplementary Figure 2). Although substance use was considered when building our model, to ensure the validity of our findings in the general population, we performed subgroup analyses among participants who never used tobacco, cannabis, or opioids (Supplementary Figure 3A–3C). Similarly, informed by past studies that showed effects on LTL by chronic co-infections [12], we analyzed participants with no history of either HCV or HBV infection (Supplementary Figure 3D). All models showed essentially the same effects as the main model, and in the smaller never smoker and never HBV/HCV-coinfected groups, the effect size of undetectable HIV infection was similar, but the confidence interval became broader and significance was lost. The effect of detectable HIV infection remained in all subgroup models. Taken together, these secondary models confirmed the robustness of the effects observed in our main model.
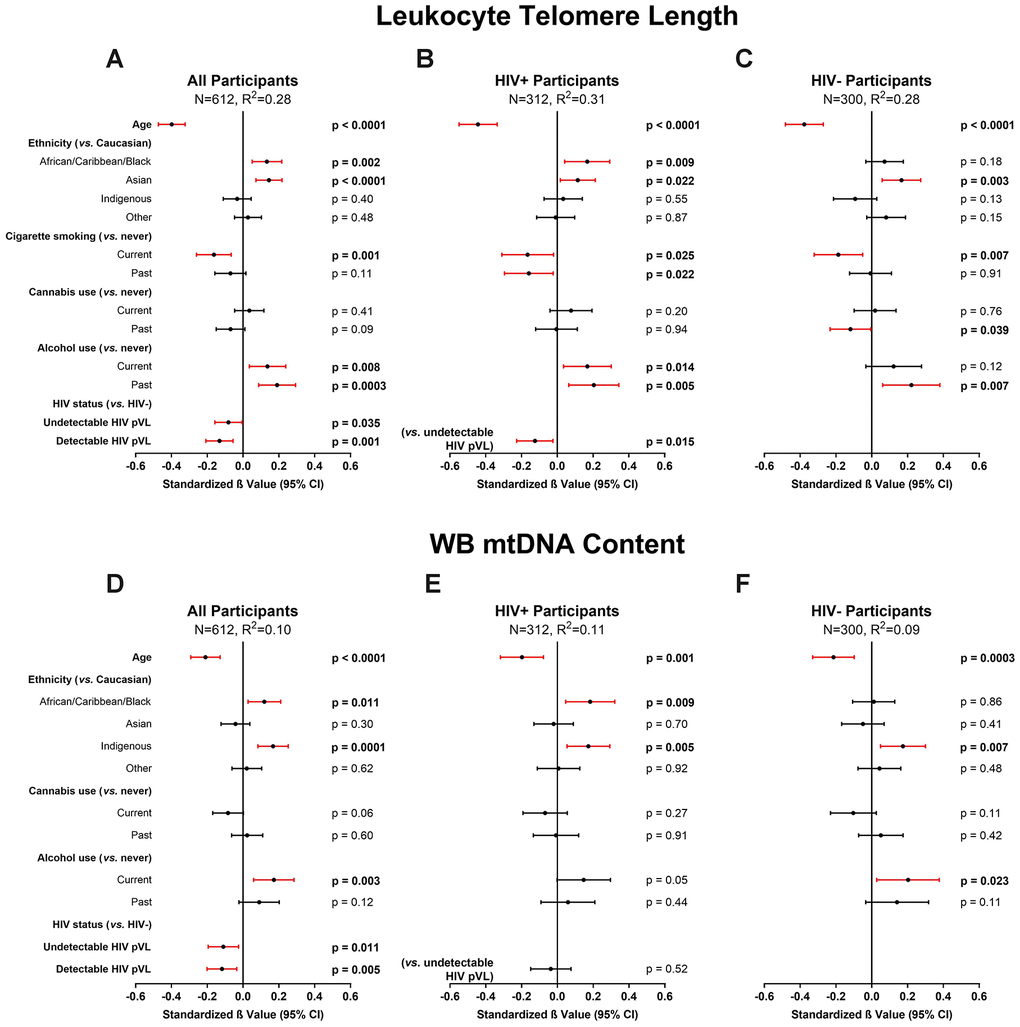
Figure 2. Multivariable modelling of cross-sectional leukocyte telomere length (LTL) and whole blood mitochondrial DNA (WB mtDNA) content. Final selected multivariable linear regression models of cross-sectional LTL (variance inflation factor (VIF) ≤2.1) in (A) all, (B) HIV+, and (C) HIV- participants, and WB mtDNA content (VIF≤1.3) in (D) all, (E) HIV+, and (F) HIV- participants. Final models among all participants were selected automatically by minimizing Akaike’s Information Criterion (AIC). Statistical significance depicted by red confidence intervals; negative standardized β values indicate associations with either shorter LTL or lower WB mtDNA content and vice versa. Coefficients of determination (R2) are shown for each model. The models show that after adjusting for age, ethnicity and substance use, HIV infection is independently associated with shorter LTL and lower WB mtDNA content. Detectable HIV viremia was associated with shorter LTL but not WB mtDNA content.
Cross-sectional predictors of WB mtDNA content
The final multivariable model for WB mtDNA content (N=612, R2=0.10) indicated that HIV infection and older age were independently associated with decreased WB mtDNA content, while ACB and Indigenous ethnicities, as well as current alcohol use, showed an association with increased WB mtDNA content (Figure 2D). Contrary to the LTL models, no difference was detected between participants with detectable or undetectable HIV pVL (Figure 2E). Once again, we carried out subgroup analyses as above and found that older age remained in all models while the effect of HIV was detected sporadically, likely due to lower power and wider confidence intervals (Supplementary Figure 4).
Sensitivity models were constructed including platelet count data that were available for a subset of study participants. Here, higher platelet count showed an association with higher WB mtDNA content (Supplementary Figure 5A–5C) and the relationship between older age and decreased WB mtDNA content remained, but the effect of HIV was lost. However, in these models, the sample size of the HIV-negative group was considerably reduced (N=121 vs. N=300).
Predictors of longitudinal change in LTL
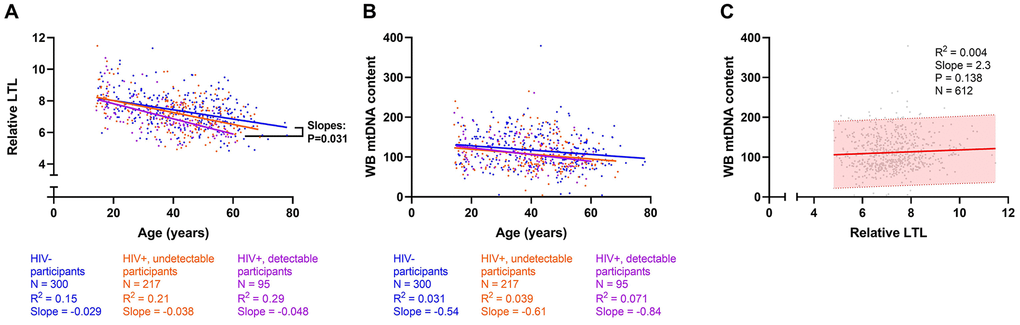
In the univariate analysis, overall relative LTL shortened by 0.036 units/year among all participants. This represents a loss of approximately 36 bp/year, in close agreement with the population attrition rate estimate (33 bp/year) seen in the cross-sectional analysis (Figure 1A). These univariate investigations were validated by multivariable modelling (N=296, R2=0.33), in which we observed that a slower decrease in WB mtDNA content/year was a strong predictor of faster LTL loss (Figure 4A), as was having longer LTL at baseline. Older age at baseline was also associated with faster LTL loss while past alcohol use, current opioid use, as well as ACB and Asian ethnicities were associated with slower loss of LTL (Figure 4A), reminiscent of the cross-sectional analysis. In contrast to the cross-sectional finding, tobacco smoking and undetectable HIV did not show an effect on LTL attrition rate. However, among WLWH, LTL was lost faster in those with a detectable HIV pVL at last visit (Figure 4B). Among HIV-negative participants (Figure 4C), only age, baseline LTL and WB ΔmtDNA content were included due to the reduced sample, and they all showed similar effects as those seen in the all-participants model. Models showing non-standardized effect sizes are shown in supplement (Supplementary Figure 6). In all subgroup analyses that excluded substance users or co-infections (Supplementary Figure 7), slower loss of WB mtDNA content/year and longer baseline LTL remained independently associated with faster LTL attrition. Similar to the cross-sectional subgroup models, the effect of HIV lost significance in some models, likely due to reduced power.
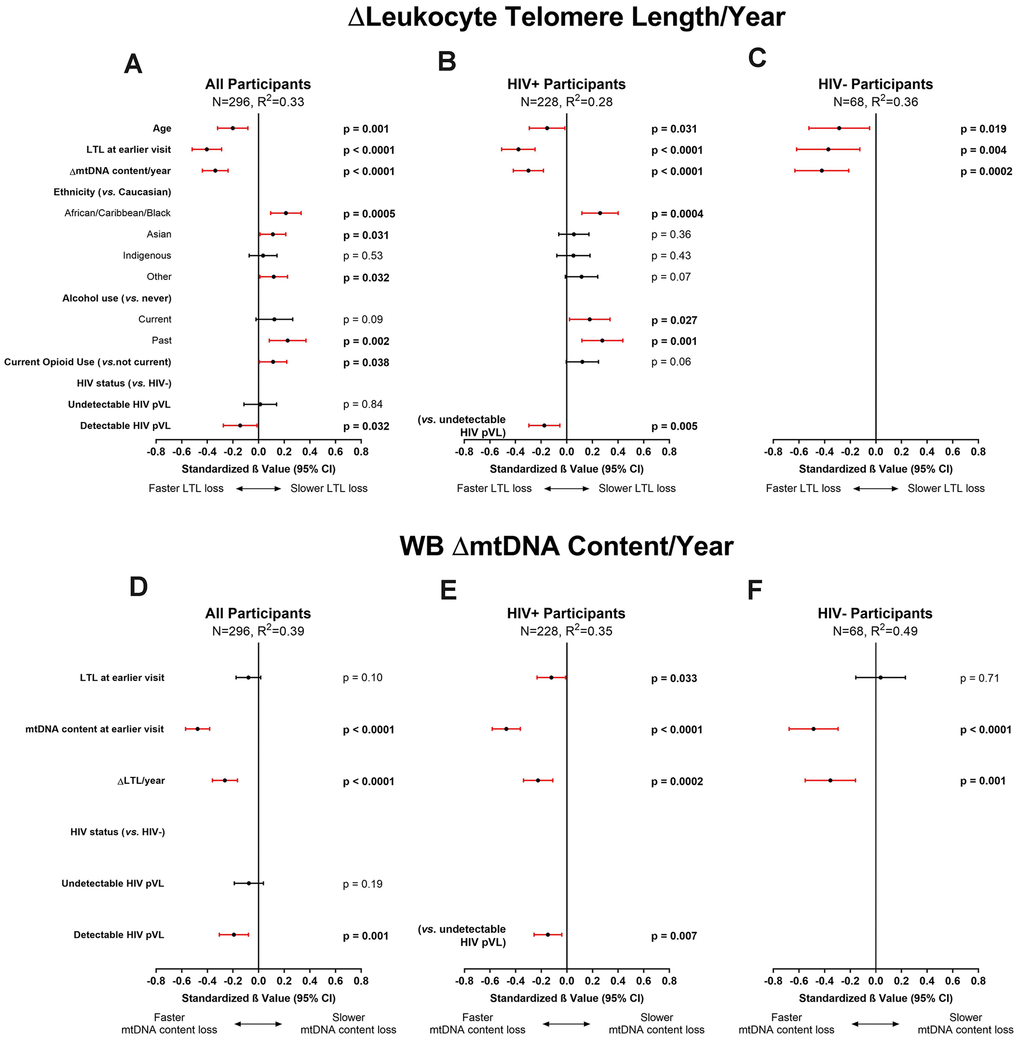
Figure 4. Multivariable modelling of longitudinal rates of change in leukocyte telomere length (LTL) and whole blood mitochondrial DNA (WB mtDNA) content. Final selected multivariable linear regression models of longitudinal ΔLTL/year (variance inflation factor (VIF) ≤1.5) in (A) all, (B) HIV+, and (C) HIV- participants, and WB ΔmtDNA content/year (VIF≤1.2) in (D) all, (E) HIV+, and (F) HIV- participants. Final models among all participants were selected automatically by minimizing Akaike’s Information Criterion (AIC). Statistical significance depicted by red confidence intervals; negative standardized β values indicate associations with either faster LTL loss or faster WB mtDNA content loss and vice versa. Coefficients of determination (R2) are shown for each model. The ΔLTL/year models show that after adjusting for age and LTL at baseline, ethnicity, and substance use, a slower loss of WB mtDNA/year is significantly independently associated with faster LTL attrition. Similarly, the models for WB ΔmtDNA content/year show that after adjusting for LTL and WB mtDNA at baseline, a slower rate of LTL attrition is independently associated with faster loss of WB mtDNA content. Detectable HIV viremia was associated with faster decline in both markers.
Taken together, our data show that the main predictors of faster LTL attrition were having longer LTL at baseline and a slower WB mtDNA content decline over time. Our data also point toward faster LTL attrition among those who have a detectable HIV pVL while those who maintain viral control showed rates of LTL loss essentially identical to HIV-negative individuals.
Predictors of longitudinal change in WB mtDNA content
In WLWH and controls, WB mtDNA content decreased by medians of 10 and 7 copies of mtDNA per copy of nDNA/year, respectively (Table 1). Similar to what was observed for ΔLTL, in the final model of WB ΔmtDNA content (N=296, R2=0.39), detectable HIV infection, higher WB mtDNA content at baseline, and slower LTL attrition rate were each independently associated with a faster decline in WB mtDNA content (Figure 4D). Among WLWH (Figure 4E) and HIV-negative controls (Figure 4F), the same associations were seen. Furthermore, compared to WLWH with undetectable HIV pVL, having a detectable HIV pVL at last visit was associated with faster WB mtDNA content decline (Figure 4E). Subgroup analyses were highly similar to the all-participants model (Supplementary Figure 8) and the effect of viremia persisted in all models. Adding platelet counts to the WB ΔmtDNA content models had no effect, and platelet count itself showed no association with WB ΔmtDNA content (Supplementary Figure 9).
Overall, as seen for the change in LTL, the strongest predictors of faster decrease in WB mtDNA content were higher WB mtDNA content at baseline and slower LTL loss. In addition, having a detectable HIV pVL accelerated mtDNA loss.
Discussion
This is the first human cohort study to incorporate both LTL and WB mtDNA content measures and their rates of change over time, adjusting for relevant covariates. Our primary finding is that the rates of change in the two measures are inversely associated, meaning that while both decline with age, a faster decline in one marker is observed if the decline in the other is slower, and vice versa. We further show the negative impact of uncontrolled HIV viremia on both LTL and WB mtDNA content loss longitudinally. Given that both markers were measured using the same technique (monochrome multiplex quantitative polymerase chain reaction, MMqPCR) on the same cohort specimens, this study was uniquely suited to characterize the relative effect of predictors, as well as the influence of one marker on the other. These data highlight the importance of investigating both LTL and WB mtDNA content as complementary and non-independent markers of aging.
The inverse relationship between LTL and mtDNA content attrition rates, seen in both the ΔLTL/year and the ΔmtDNA content/year models, is a novel finding and sheds light on previously proposed mechanisms that implicate a relationship between telomere and mitochondrial biology. This effect is consistent with the dual functions of telomerase. While the classical function of telomerase is to maintain telomeres, in vitro research has demonstrated that under oxidative stress conditions, the enzymatic subunit of telomerase translocates to the mitochondria where it exerts a protective function against oxidative damage but appears to neglect telomere maintenance [41, 42]. Telomerase localization to mitochondria has also been demonstrated in vivo in the neurons of both mice and humans [48, 49]. A similar phenomenon may be occurring in leukocytes when mitochondria require protection, potentially resulting in mitigated mtDNA loss at the cost of higher LTL attrition rate, as seen here. If so, this would suggest that upon oxidative stress challenge, telomerase confers a necessary and immediate protection to the mitochondria at the expense of the long-term benefit of telomere maintenance. Furthermore, inter-related mechanisms of mitochondrial metabolism, telomerase-associated DNA damage sensing, and ribosome production, previously described as the mitochondrion/telomere nucleoprotein complex/ribosome (MTR) postulate [50], have been implicated in cellular aging, in which disruption of any one mechanism may lead to consequences throughout all three systems. The potential mechanistic relationships between LTL, WB mtDNA content, and other previously established markers of aging and/or senescence should also be explored. For example, HIV has been linked with changes in expression of cyclin-dependent kinase inhibitor 2A (CDKN2A) [51] and DNA methylation levels [52]. As yet, no direct evidence links the relationship between LTL and mtDNA content attrition rates to any of the mechanisms described above. Future longitudinal studies in an independent cohort with a longer interval between measures are needed to confirm this relationship and determine if this apparent “yin and yang” effect persists over longer periods of time.
The rate of LTL attrition observed agrees with previous research [3] and is consistent between our cross-sectional and longitudinal data. Although WB mtDNA content also declined with age, both univariable and multivariable analyses explained a smaller percentage of the total variance compared to that of LTL models. Given that the variability of each of the two assays is comparable [53, 54], this suggests that either WB mtDNA content is intrinsically more stochastic, or that the mtDNA analyses presented here missed key explanatory variables. Such yet unidentified variables may partially explain the inconsistency in previous attempts by researchers to establish WB mtDNA content as a robust marker of aging. A potential confounder that we addressed in WB mtDNA content models was platelets, which may lead to overestimation of WB mtDNA content. Two studies have shown that a very large increase in platelet mtDNA contribution would be necessary to meaningfully affect WB mtDNA content measurement [55, 56], and others found no relationship between platelet count and WB mtDNA content [26, 57]. Based on this and our own unpublished data in a non-HIV sample, we did not initially intend to include platelet count in our analyses. However, a recent study highlighted the importance of platelet count in studies investigating the effect of HIV on WB mtDNA content [27]. Therefore, to increase the rigor of our analysis, we analyzed a subset of our participants for whom platelet count data were available to investigate the role of platelets in WB mtDNA content and its change over time. We found that platelet count had no effect on our longitudinal models. Despite a modest reduction in power, the effects of HIV on faster WB mtDNA attrition, as well as the inverse relationship between the rates of change in LTL and WB mtDNA content over time, persisted. However, while platelet count affected WB mtDNA content cross-sectionally, its inclusion resulted in a substantial reduction in sample size and the effect of HIV lost statistical significance. It remains unclear whether the well-established relationship between HIV and decreased WB mtDNA content is partially driven by thrombocytopenia in people living with HIV.
We could not detect any significant cross-sectional relationship between LTL and WB mtDNA content, whether univariately or multivariately, in contrast to some studies in healthy adults that reported modest correlations (R2=0.15, and R2=0.03) [36, 44]. The reason for this is unclear and may be related to differences in the assays used and the range they yield. There could also be a bias toward not reporting negative findings (i.e. no correlation) by other studies. Furthermore, past studies did not examine this relationship while adjusting for covariates and confounders.
HIV infection and uncontrolled viremia have been previously associated with both shorter LTL [12] and lower mtDNA content [26, 27]. Based on non-standardized effect sizes, our study demonstrates that the independent effects on LTL by HIV infection, uncontrolled HIV viremia, and current tobacco smoking are each similar to approximately one decade of chronological aging. Furthermore, we show that the decline of both LTL and mtDNA content with age is accelerated in participants with uncontrolled HIV viremia, emphasizing the importance of maintaining HIV viral control. This is supported by model estimates showing that having uncontrolled viremia increases the LTL loss per year of aging from approximately 30 bp to more than 100 bp.
Previous research has shown early and rapid LTL decline following HIV acquisition [14, 15], which may result from an initial HIV-induced immune activation leading to rearrangement of leukocyte subsets. It is possible that a similar phenomenon is occurring here, in which immune cell proliferation triggered by actively replicating HIV results in faster apparent attrition of both LTL and WB mtDNA content. A recent study demonstrated faster mtDNA content attrition among WLWH who were approximately 50 years old compared to controls [27]. However, more research with longitudinal samples spanning longer periods of time is necessary to determine whether this effect is transient or carries long-term consequences. Indeed, these findings are in keeping with the model of HIV-mediated accelerated aging and may explain the link between HIV and aging comorbidities. While HIV represents a valuable model of accelerated human aging, other chronic and proinflammatory diseases such as diabetes, rheumatoid arthritis, and other chronic/latent viral infections associated with aging comorbidities may similarly modulate LTL and mtDNA content.
In support of previous research, we show that tobacco smoking is robustly independently associated with shorter LTL. In most subgroup analyses, this effect exists only among current smokers, suggesting that smoking cessation confers a benefit to LTL, as reported by others [10]. However, it is unclear why the benefit of smoking cessation was not detected among WLWH. It is possible that time since smoking cessation differed between groups. Alternatively, the mechanism by which LTL recovers after smoking cessation may be adversely affected by the presence of HIV. Smoking was also not associated with LTL attrition rate, likely because the time elapsed between longitudinal visits was insufficient for the effect of smoking to reach a detectable size.
In contrast, alcohol use was associated with longer LTL, slower LTL attrition, and increased WB mtDNA content. While this may seem to contradict some research reporting shorter LTL in people who abuse alcohol [58], other studies are less clear as to the relationship between LTL and moderate alcohol consumption [21, 59]. Our definition for this variable was any alcohol use regardless of quantity, which does not imply abuse and would also include low to moderate users. Moreover, unlike tobacco smoking or opioid use, alcohol use is lower among our WLWH. The apparent association between alcohol use and both longer LTL and increased WB mtDNA content may therefore be the result of nested effects with unknown factors more prevalent among the HIV-negative group, and not fully addressed by the covariates considered herein. It is also possible that we detected a spurious finding, as the association with alcohol disappeared in some subgroup analyses. Nevertheless, the associations between alcohol and the markers studied here are of interest and should be reproduced in an independent cohort, along with a more granular analysis of alcohol consumption, with respect to frequency and quantity.
While differences in LTL dynamics between men and women exist, this study includes only female participants. As such, we are not able to comment on whether the same predictors of LTL and WB mtDNA content would be seen in men. Furthermore, this analysis also does not consider the intensity and frequency of substance use. The role of sex and substance use should therefore be further examined.
In summary, we describe a novel relationship between longitudinal LTL and WB mtDNA dynamics, whereby better preservation of LTL appears to occur at the cost of mtDNA loss and vice versa. We also validate the cross-sectional predictors of LTL and WB mtDNA content that have been previously shown. In addition, we demonstrate that for women living with HIV, loss of viral control exacerbates both LTL and WB mtDNA content attrition and may contribute to biological aging. Taken together, our findings provide further evidence supporting the importance of consistently maintaining HIV viral control. Given that the effects of HIV viremia are believed to be largely related to chronic inflammation, it is possible that other proinflammatory conditions would similarly exacerbate cellular aging through related mechanisms. Our findings further imply that future longitudinal studies of either LTL or WB mtDNA content as a biomarker should also consider the other given their strong relationship.
Materials and Methods
Study sample
Study participants included all non-pregnant women and girls ≥12 years old living with or without HIV enrolled in the Children and Women: AntiRetrovirals and Markers of Aging (CARMA) cohort between December 2008 and August 2017. Enrolment and blood collection of WLWH took place during routine clinical visits at four sites across Canada. HIV-negative controls were purposely recruited to have similar sociodemographic characteristics as the HIV group. Written informed consent was provided by adult participants, and assent from pediatric participants was obtained with parent/legal guardian consent. This study was approved by the University of British Columbia Research Ethics Board at the Children’s and Women’s Hospital (H08-02018). Further details on CARMA enrolment have been previously described [12], and more information can be found in Supplementary methods.
Study participants total 328 WLWH and 318 HIV-negative women and girls. Of these, one participant was excluded because no blood was collected and 33 were excluded because of missing demographic or clinical data that were deemed essential a priori (Supplementary Figure 1). These variables included age, ethnicity, tobacco smoking, cannabis use, alcohol use, opioid use, HCV infection ever, and HIV status for all participants, as well as current HIV pVL, and CD4 count at visit for WLWH. Other data considered included BMI, household income, highest education level completed, HCV viremia, highest HIV pVL ever recorded (peak pVL), lowest CD4 count recorded, and platelet count (Supplementary Table 1). For participants with ≥2 study visits at least one year apart, the last visit was used for cross-sectional analyses, and the earliest (or baseline) and latest visits were used for longitudinal analyses.
In total, cross-sectional analyses included 312 WLWH and 300 HIV-negative participants. Longitudinal analyses included 228 WLWH and 68 HIV-negative participants.
Biospecimen collection and qPCR
WB was collected and stored at -80°C until genomic DNA was extracted from 100 μl of WB using the QIAamp DNA Mini Kit on the QIAcube (Qiagen, Hilden, Germany) according to the manufacturer’s blood and body fluid protocol. DNA was eluted in 100 μl of Buffer AE which contains 0.5 mM EDTA and 10 mM Tris-Cl, pH 9.0, and stored at -80oC until assayed.
Relative LTL and mtDNA content were measured in duplicate by MMqPCR as previously described [53, 54]. TL was defined as the ratio between telomeric DNA quantity and the copy number of a single-copy nuclear gene, albumin (ALB). MtDNA content was defined as the ratio of mitochondrial genome copy number, measured using a segment of the mitochondrial displacement loop, to ALB copy number. The TL assay was calibrated using fluorescent in situ hybridization such that each unit of relative TL approximates 1 kb of telomeric DNA, whereas the mtDNA content assay provides the mtDNA to nuclear DNA ratio. Both LTL and WB mtDNA content were assayed in whole blood, producing measurements for the average leukocyte. In a previous publication by our group, the intra- and inter-assay variabilities of the TL assay were measured among several operators to be 4.2-6.2% and 3.2-4.9%, respectively [53]. Similarly, the intra- and inter-assay variabilities of the mtDNA content assay were determined to be 4.3-7.9% and 2.9-9.2%, respectively [54].
Statistical analyses
To evaluate the relationship between potential explanatory variables and our measures of interest, namely LTL and mtDNA content and the change in these over time, possible associations were first explored with univariate analyses using Pearson’s or Spearman’s correlation, one-way analysis of variance (ANOVA), Kruskal-Wallis, Student’s t-, or Mann-Whitney U tests. Variables that were important in univariate analysis (P<0.15) were candidates for inclusion in multivariable models. The final models for each measure of interest included all essential variables as described above and were constructed in a stepwise manner, by minimizing the Akaike information criterion (AIC). While possible interactions were explored, none offered substantial model improvement. Chi-Square contingency tables, Kruskal-Wallis tests, and Spearman’s correlation were used to detect collinearity among variables, and variance inflation factors (VIFs) were calculated to characterize the influence of collinearity in the models. To further test the robustness of final models, subgroup analyses (such as among a specific ethnicity or HIV group) and sensitivity analyses were done to investigate the effect of HIV clinical variables and non-essential variables, but with reduced power.
Statistical analyses were performed using XLSTAT version 2019.1.1.
Acknowledgments
The authors thank all study participants, collaborators and research staff at CARMA, as well as members of the Côté Lab for their guidance and support.
Conflicts of Interest
The authors have no conflicts of interest to disclose.
Funding
This work was supported by the Canadian Institutes of Health Research (CIHR) Team Grant on Cellular Aging and HIV Comorbidities in Women and Children 2013-2018 TCO-125269 (to HCFC). The authors acknowledge additional support from the University of British Columbia (UBC) Centre for Blood Research (CBR) Summer Studentship (to EK), and the CBR Graduate Student Award, UBC Four Year Doctoral Fellowship, and CIHR Doctoral Research Award (to AYYH).
References
- 1. Müezzinler A, Zaineddin AK, Brenner H. A systematic review of leukocyte telomere length and age in adults. Ageing Res Rev. 2013; 12:509–19. https://doi.org/10.1016/j.arr.2013.01.003 [PubMed]
- 2. Frenck RW
Jr , Blackburn EH, Shannon KM. The rate of telomere sequence loss in human leukocytes varies with age. Proc Natl Acad Sci USA. 1998; 95:5607–10. https://doi.org/10.1073/pnas.95.10.5607 [PubMed] - 3. Daniali L, Benetos A, Susser E, Kark JD, Labat C, Kimura M, Desai K, Granick M, Aviv A. Telomeres shorten at equivalent rates in somatic tissues of adults. Nat Commun. 2013; 4:1597. https://doi.org/10.1038/ncomms2602 [PubMed]
- 4. Hjelmborg JB, Dalgård C, Möller S, Steenstrup T, Kimura M, Christensen K, Kyvik KO, Aviv A. The heritability of leucocyte telomere length dynamics. J Med Genet. 2015; 52:297–302. https://doi.org/10.1136/jmedgenet-2014-102736 [PubMed]
- 5. Gardner M, Bann D, Wiley L, Cooper R, Hardy R, Nitsch D, Martin-Ruiz C, Shiels P, Sayer AA, Barbieri M, Bekaert S, Bischoff C, Brooks-Wilson A, et al, and Halcyon study team. Gender and telomere length: systematic review and meta-analysis. Exp Gerontol. 2014; 51:15–27. https://doi.org/10.1016/j.exger.2013.12.004 [PubMed]
- 6. Hunt SC, Chen W, Gardner JP, Kimura M, Srinivasan SR, Eckfeldt JH, Berenson GS, Aviv A. Leukocyte telomeres are longer in African Americans than in whites: the national heart, lung, and blood institute family heart study and the bogalusa heart study. Aging Cell. 2008; 7:451–58. https://doi.org/10.1111/j.1474-9726.2008.00397.x [PubMed]
- 7. Cassidy A, De Vivo I, Liu Y, Han J, Prescott J, Hunter DJ, Rimm EB. Associations between diet, lifestyle factors, and telomere length in women. Am J Clin Nutr. 2010; 91:1273–80. https://doi.org/10.3945/ajcn.2009.28947 [PubMed]
- 8. Ludlow AT, Zimmerman JB, Witkowski S, Hearn JW, Hatfield BD, Roth SM. Relationship between physical activity level, telomere length, and telomerase activity. Med Sci Sports Exerc. 2008; 40:1764–71. https://doi.org/10.1249/MSS.0b013e31817c92aa [PubMed]
- 9. Astuti Y, Wardhana A, Watkins J, Wulaningsih W, and PILAR Research Network. Cigarette smoking and telomere length: a systematic review of 84 studies and meta-analysis. Environ Res. 2017; 158:480–89. https://doi.org/10.1016/j.envres.2017.06.038 [PubMed]
- 10. Müezzinler A, Mons U, Dieffenbach AK, Butterbach K, Saum KU, Schick M, Stammer H, Boukamp P, Holleczek B, Stegmaier C, Brenner H. Smoking habits and leukocyte telomere length dynamics among older adults: results from the ESTHER cohort. Exp Gerontol. 2015; 70:18–25. https://doi.org/10.1016/j.exger.2015.07.002 [PubMed]
- 11. Yang Z, Ye J, Li C, Zhou D, Shen Q, Wu J, Cao L, Wang T, Cui D, He S, Qi G, He L, Liu Y. Drug addiction is associated with leukocyte telomere length. Sci Rep. 2013; 3:1542. https://doi.org/10.1038/srep01542 [PubMed]
- 12. Zanet DL, Thorne A, Singer J, Maan EJ, Sattha B, Le Campion A, Soudeyns H, Pick N, Murray M, Money DM, Côté HC, and CIHR Emerging Team Grant on HIV Therapy and Aging: CARMA. Association between short leukocyte telomere length and HIV infection in a cohort study: no evidence of a relationship with antiretroviral therapy. Clin Infect Dis. 2014; 58:1322–32. https://doi.org/10.1093/cid/ciu051 [PubMed]
- 13. Bestilny LJ, Gill MJ, Mody CH, Riabowol KT. Accelerated replicative senescence of the peripheral immune system induced by HIV infection. AIDS. 2000; 14:771–80. https://doi.org/10.1097/00002030-200005050-00002 [PubMed]
- 14. Gonzalez-Serna A, Ajaykumar A, Gadawski I, Muñoz-Fernández MA, Hayashi K, Harrigan PR, Côté HC. Rapid decrease in peripheral blood mononucleated cell telomere length after HIV seroconversion, but not HCV seroconversion. J Acquir Immune Defic Syndr. 2017; 76:e29–32. https://doi.org/10.1097/QAI.0000000000001446 [PubMed]
- 15. Leung JM, Fishbane N, Jones M, Morin A, Xu S, Liu JC, MacIsaac J, Milloy MJ, Hayashi K, Montaner J, Horvath S, Kobor M, Sin DD, et al. Longitudinal study of surrogate aging measures during human immunodeficiency virus seroconversion. Aging (Albany NY). 2017; 9:687–705. https://doi.org/10.18632/aging.101184 [PubMed]
- 16. Salpea KD, Talmud PJ, Cooper JA, Maubaret CG, Stephens JW, Abelak K, Humphries SE. Association of telomere length with type 2 diabetes, oxidative stress and UCP2 gene variation. Atherosclerosis. 2010; 209:42–50. https://doi.org/10.1016/j.atherosclerosis.2009.09.070 [PubMed]
- 17. Forero DA, González-Giraldo Y, López-Quintero C, Castro-Vega LJ, Barreto GE, Perry G. Meta-analysis of telomere length in Alzheimer’s disease. J Gerontol A Biol Sci Med Sci. 2016; 71:1069–73. https://doi.org/10.1093/gerona/glw053 [PubMed]
- 18. Willeit P, Willeit J, Mayr A, Weger S, Oberhollenzer F, Brandstätter A, Kronenberg F, Kiechl S. Telomere length and risk of incident cancer and cancer mortality. JAMA. 2010; 304:69–75. https://doi.org/10.1001/jama.2010.897 [PubMed]
- 19. Ma H, Zhou Z, Wei S, Liu Z, Pooley KA, Dunning AM, Svenson U, Roos G, Hosgood HD
3rd , Shen M, Wei Q. Shortened telomere length is associated with increased risk of cancer: a meta-analysis. PLoS One. 2011; 6:e20466. https://doi.org/10.1371/journal.pone.0020466 [PubMed] - 20. Fitzpatrick AL, Kronmal RA, Gardner JP, Psaty BM, Jenny NS, Tracy RP, Walston J, Kimura M, Aviv A. Leukocyte telomere length and cardiovascular disease in the cardiovascular health study. Am J Epidemiol. 2007; 165:14–21. https://doi.org/10.1093/aje/kwj346 [PubMed]
- 21. Bekaert S, De Meyer T, Rietzschel ER, De Buyzere ML, De Bacquer D, Langlois M, Segers P, Cooman L, Van Damme P, Cassiman P, Van Criekinge W, Verdonck P, De Backer GG, et al, and Asklepios investigators. Telomere length and cardiovascular risk factors in a middle-aged population free of overt cardiovascular disease. Aging Cell. 2007; 6:639–47. https://doi.org/10.1111/j.1474-9726.2007.00321.x [PubMed]
- 22. van der Harst P, van der Steege G, de Boer RA, Voors AA, Hall AS, Mulder MJ, van Gilst WH, van Veldhuisen DJ, and MERIT-HF Study Group. Telomere length of circulating leukocytes is decreased in patients with chronic heart failure. J Am Coll Cardiol. 2007; 49:1459–64. https://doi.org/10.1016/j.jacc.2007.01.027 [PubMed]
- 23. Cawthon RM, Smith KR, O’Brien E, Sivatchenko A, Kerber RA. Association between telomere length in blood and mortality in people aged 60 years or older. Lancet. 2003; 361:393–95. https://doi.org/10.1016/S0140-6736(03)12384-7 [PubMed]
- 24. Mengel-From J, Thinggaard M, Dalgård C, Kyvik KO, Christensen K, Christiansen L. Mitochondrial DNA copy number in peripheral blood cells declines with age and is associated with general health among elderly. Hum Genet. 2014; 133:1149–59. https://doi.org/10.1007/s00439-014-1458-9 [PubMed]
- 25. Miró O, López S, Martínez E, Pedrol E, Milinkovic A, Deig E, Garrabou G, Casademont J, Gatell JM, Cardellach F. Mitochondrial effects of HIV infection on the peripheral blood mononuclear cells of HIV-infected patients who were never treated with antiretrovirals. Clin Infect Dis. 2004; 39:710–16. https://doi.org/10.1086/423176 [PubMed]
- 26. Côté HC, Brumme ZL, Craib KJ, Alexander CS, Wynhoven B, Ting L, Wong H, Harris M, Harrigan PR, O’Shaughnessy MV, Montaner JS. Changes in mitochondrial DNA as a marker of nucleoside toxicity in HIV-infected patients. N Engl J Med. 2002; 346:811–20. https://doi.org/10.1056/NEJMoa012035 [PubMed]
- 27. Sun J, Longchamps RJ, Piggott DA, Castellani CA, Sumpter JA, Brown TT, Mehta SH, Arking DE, Kirk GD. Association between HIV infection and mitochondrial DNA copy number in peripheral blood: a population-based, prospective cohort study. J Infect Dis. 2019; 219:1285–93. https://doi.org/10.1093/infdis/jiy658 [PubMed]
- 28. Lee HK, Song JH, Shin CS, Park DJ, Park KS, Lee KU, Koh CS. Decreased mitochondrial DNA content in peripheral blood precedes the development of non-insulin-dependent diabetes mellitus. Diabetes Res Clin Pract. 1998; 42:161–67. https://doi.org/10.1016/s0168-8227(98)00110-7 [PubMed]
- 29. Delbarba A, Abate G, Prandelli C, Marziano M, Buizza L, Arce Varas N, Novelli A, Cuetos F, Martinez C, Lanni C, Memo M, Uberti D. Mitochondrial alterations in peripheral mononuclear blood cells from Alzheimer’s disease and mild cognitive impairment patients. Oxid Med Cell Longev. 2016; 2016:5923938. https://doi.org/10.1155/2016/5923938 [PubMed]
- 30. Wang Y, Liu VW, Xue WC, Cheung AN, Ngan HY. Association of decreased mitochondrial DNA content with ovarian cancer progression. Br J Cancer. 2006; 95:1087–91. https://doi.org/10.1038/sj.bjc.6603377 [PubMed]
- 31. Meierhofer D, Mayr JA, Foetschl U, Berger A, Fink K, Schmeller N, Hacker GW, Hauser-Kronberger C, Kofler B, Sperl W. Decrease of mitochondrial DNA content and energy metabolism in renal cell carcinoma. Carcinogenesis. 2004; 25:1005–10. https://doi.org/10.1093/carcin/bgh104 [PubMed]
- 32. Chen S, Xie X, Wang Y, Gao Y, Xie X, Yang J, Ye J. Association between leukocyte mitochondrial DNA content and risk of coronary heart disease: a case-control study. Atherosclerosis. 2014; 237:220–26. https://doi.org/10.1016/j.atherosclerosis.2014.08.051 [PubMed]
- 33. Lee HC, Lu CY, Fahn HJ, Wei YH. Aging- and smoking-associated alteration in the relative content of mitochondrial DNA in human lung. FEBS Lett. 1998; 441:292–96. https://doi.org/10.1016/s0014-5793(98)01564-6 [PubMed]
- 34. Masayesva BG, Mambo E, Taylor RJ, Goloubeva OG, Zhou S, Cohen Y, Minhas K, Koch W, Sciubba J, Alberg AJ, Sidransky D, Califano J. Mitochondrial DNA content increase in response to cigarette smoking. Cancer Epidemiol Biomarkers Prev. 2006; 15:19–24. https://doi.org/10.1158/1055-9965.EPI-05-0210 [PubMed]
- 35. Hosgood HD
3rd , Liu CS, Rothman N, Weinstein SJ, Bonner MR, Shen M, Lim U, Virtamo J, Cheng WL, Albanes D, Lan Q. Mitochondrial DNA copy number and lung cancer risk in a prospective cohort study. Carcinogenesis. 2010; 31:847–49. https://doi.org/10.1093/carcin/bgq045 [PubMed] - 36. Campa D, Barrdahl M, Santoro A, Severi G, Baglietto L, Omichessan H, Tumino R, Bueno-de-Mesquita HB, Peeters PH, Weiderpass E, Chirlaque MD, Rodríguez-Barranco M, Agudo A, et al. Mitochondrial DNA copy number variation, leukocyte telomere length, and breast cancer risk in the European Prospective Investigation into Cancer and Nutrition (EPIC) study. Breast Cancer Res. 2018; 20:29. https://doi.org/10.1186/s13058-018-0955-5 [PubMed]
- 37. Knez J, Winckelmans E, Plusquin M, Thijs L, Cauwenberghs N, Gu Y, Staessen JA, Nawrot TS, Kuznetsova T. Correlates of peripheral blood mitochondrial DNA content in a general population. Am J Epidemiol. 2016; 183:138–46. https://doi.org/10.1093/aje/kwv175 [PubMed]
- 38. Liu CS, Tsai CS, Kuo CL, Chen HW, Lii CK, Ma YS, Wei YH. Oxidative stress-related alteration of the copy number of mitochondrial DNA in human leukocytes. Free Radic Res. 2003; 37:1307–17. https://doi.org/10.1080/10715760310001621342 [PubMed]
- 39. Xing J, Chen M, Wood CG, Lin J, Spitz MR, Ma J, Amos CI, Shields PG, Benowitz NL, Gu J, de Andrade M, Swan GE, Wu X. Mitochondrial DNA content: its genetic heritability and association with renal cell carcinoma. J Natl Cancer Inst. 2008; 100:1104–12. https://doi.org/10.1093/jnci/djn213 [PubMed]
- 40. Sahin E, Colla S, Liesa M, Moslehi J, Müller FL, Guo M, Cooper M, Kotton D, Fabian AJ, Walkey C, Maser RS, Tonon G, Foerster F, et al. Telomere dysfunction induces metabolic and mitochondrial compromise. Nature. 2011; 470:359–65. https://doi.org/10.1038/nature09787 [PubMed]
- 41. Ahmed S, Passos JF, Birket MJ, Beckmann T, Brings S, Peters H, Birch-Machin MA, von Zglinicki T, Saretzki G. Telomerase does not counteract telomere shortening but protects mitochondrial function under oxidative stress. J Cell Sci. 2008; 121:1046–53. https://doi.org/10.1242/jcs.019372 [PubMed]
- 42. Haendeler J, Dröse S, Büchner N, Jakob S, Altschmied J, Goy C, Spyridopoulos I, Zeiher AM, Brandt U, Dimmeler S. Mitochondrial telomerase reverse transcriptase binds to and protects mitochondrial DNA and function from damage. Arterioscler Thromb Vasc Biol. 2009; 29:929–35. https://doi.org/10.1161/ATVBAHA.109.185546 [PubMed]
- 43. Kim JH, Kim HK, Ko JH, Bang H, Lee DC. The relationship between leukocyte mitochondrial DNA copy number and telomere length in community-dwelling elderly women. PLoS One. 2013; 8:e67227. https://doi.org/10.1371/journal.pone.0067227 [PubMed]
- 44. Tyrka AR, Carpenter LL, Kao HT, Porton B, Philip NS, Ridout SJ, Ridout KK, Price LH. Association of telomere length and mitochondrial DNA copy number in a community sample of healthy adults. Exp Gerontol. 2015; 66:17–20. https://doi.org/10.1016/j.exger.2015.04.002 [PubMed]
- 45. Zole E, Ranka R. Mitochondrial DNA copy number and telomere length in peripheral blood mononuclear cells in comparison with whole blood in three different age groups. Arch Gerontol Geriatr. 2019; 83:131–37. https://doi.org/10.1016/j.archger.2019.04.007 [PubMed]
- 46. Alegría-Torres JA, Velázquez-Villafaña M, López-Gutiérrez JM, Chagoyán-Martínez MM, Rocha-Amador DO, Costilla-Salazar R, García-Torres L. Association of leukocyte telomere length and mitochondrial DNA copy number in children from salamanca, Mexico. Genet Test Mol Biomarkers. 2016; 20:654–59. https://doi.org/10.1089/gtmb.2016.0176 [PubMed]
- 47. Deeks SG. HIV infection, inflammation, immunosenescence, and aging. Annu Rev Med. 2011; 62:141–55. https://doi.org/10.1146/annurev-med-042909-093756 [PubMed]
- 48. Eitan E, Braverman C, Tichon A, Gitler D, Hutchison ER, Mattson MP, Priel E. Excitotoxic and radiation stress increase TERT levels in the mitochondria and cytosol of cerebellar purkinje neurons. Cerebellum. 2016; 15:509–17. https://doi.org/10.1007/s12311-015-0720-6 [PubMed]
- 49. Spilsbury A, Miwa S, Attems J, Saretzki G. The role of telomerase protein TERT in Alzheimer’s disease and in tau-related pathology in vitro. J Neurosci. 2015; 35:1659–74. https://doi.org/10.1523/JNEUROSCI.2925-14.2015 [PubMed]
- 50. Davies RW, Morris B. Molecular Biology of the Neuron. Molecular and Cellular Neurobiology Series. Oxford: Oxford University Press. 2004. https://doi.org/10.1093/acprof:oso/9780198509981.001.0001
- 51. Pathai S, Lawn SD, Gilbert CE, McGuinness D, McGlynn L, Weiss HA, Port J, Christ T, Barclay K, Wood R, Bekker LG, Shiels PG. Accelerated biological ageing in HIV-infected individuals in South Africa: a case-control study. AIDS. 2013; 27:2375–84. https://doi.org/10.1097/QAD.0b013e328363bf7f [PubMed]
- 52. Horvath S, Levine AJ. HIV-1 infection accelerates age according to the epigenetic clock. J Infect Dis. 2015; 212:1563–73. https://doi.org/10.1093/infdis/jiv277 [PubMed]
- 53. Hsieh AY, Saberi S, Ajaykumar A, Hukezalie K, Gadawski I, Sattha B, Côté HC. Optimization of a relative telomere length assay by monochromatic multiplex real-time quantitative PCR on the LightCycler 480: sources of variability and quality control considerations. J Mol Diagn. 2016; 18:425–37. https://doi.org/10.1016/j.jmoldx.2016.01.004 [PubMed]
- 54. Hsieh AY, Budd M, Deng D, Gadawska I, Côté HC. A monochrome multiplex real-time quantitative PCR assay for the measurement of mitochondrial DNA content. J Mol Diagn. 2018; 20:612–20. https://doi.org/10.1016/j.jmoldx.2018.05.001 [PubMed]
- 55. Urata M, Koga-Wada Y, Kayamori Y, Kang D. Platelet contamination causes large variation as well as overestimation of mitochondrial DNA content of peripheral blood mononuclear cells. Ann Clin Biochem. 2008; 45:513–14. https://doi.org/10.1258/acb.2008.008008 [PubMed]
- 56. Hurtado-Roca Y, Ledesma M, Gonzalez-Lazaro M, Moreno-Loshuertos R, Fernandez-Silva P, Enriquez JA, Laclaustra M. Adjusting MtDNA quantification in whole blood for peripheral blood platelet and leukocyte counts. PLoS One. 2016; 11:e0163770. https://doi.org/10.1371/journal.pone.0163770 [PubMed]
- 57. Budd MA, Calli K, Samson L, Bowes J, Hsieh AY, Forbes JC, Bitnun A, Singer J, Kakkar F, Alimenti A, Maan EJ, Lewis ME, Gentile C, et al. Blood mitochondrial DNA content in HIV-exposed uninfected children with autism spectrum disorder. Viruses. 2018; 10:77. https://doi.org/10.3390/v10020077 [PubMed]
- 58. Pavanello S, Hoxha M, Dioni L, Bertazzi PA, Snenghi R, Nalesso A, Ferrara SD, Montisci M, Baccarelli A. Shortened telomeres in individuals with abuse in alcohol consumption. Int J Cancer. 2011; 129:983–92. https://doi.org/10.1002/ijc.25999 [PubMed]
- 59. Dixit S, Whooley MA, Vittinghoff E, Roberts JD, Heckbert SR, Fitzpatrick AL, Lin J, Leung C, Mukamal KJ, Marcus GM. Alcohol consumption and leukocyte telomere length. Sci Rep. 2019; 9:1404. https://doi.org/10.1038/s41598-019-38904-0 [PubMed]