Introduction
Biological aging is characterized by a progressive decline of physiological and metabolic functions across multiple organ systems. One of the key phenotypes of aging is the loss of skeletal muscle mass, a condition known as age-associated muscle atrophy or sarcopenia [1–3]. This is usually accompanied by reduced strength, muscle quality and mobility, increased risk of frailty and falls, lack of endurance and poor physical performance [4]. In the western population, >40% of adults over the age of 60 have difficulties with daily activities such as walking or standing up from a chair etc. and >30% suffer from some kind of physical disability [5]. After the age of 60, striking changes occur in muscle physiology, corresponding to a decline in muscle mass and deterioration of muscle strength by ~2% each year [6]. The pathophysiology of age-related decline in muscle mass and function is multifactorial, including biological factors such as hormonal imbalance, neurodegeneration and motor neuron loss, increased inflammation and circulating cytokines, as well as environmental factors such as physical inactivity, inadequate nutritional intake and psychosocial factors [4, 5, 7]. Furthermore, age-associated changes in muscle metabolism, such as mitochondrial dysfunction and insulin resistance, can have severe implications in muscle homeostasis and regeneration [8, 9]. We have shown that progressive resistance exercise training (PRT) induces meaningful increases in muscle strength, power, and functional mobility, however, the hypertrophic response is, on average, reduced in old compared to young, especially within men [10]. The hypertrophic response to PRT is also highly variable in older adults (reviewed in [11]).
The use of pharmacological interventions to augment the effect of PRT on muscle hypertrophy in older adults has been proposed [12–15]. The biguanide metformin has been studied in the context of exercise capacity, quality of life and mood states, and metabolic adaptations, such as insulin production and clearance, oxidative stress and cardiometabolic health in older adults with prediabetes and Type II diabetes [16–18]. Metformin has been extensively used since the 1950s, as the first-line treatment against Type II diabetes and is one of the most commonly prescribed drugs in the world, either as a monotherapy or in combination with insulin or other anti-hyperglycemic agents [19]. Metformin directly inhibits mitochondrial enzymes including complex I [20], activates AMP-activated protein kinase (AMPK) [21], inhibits NF-κB signaling and specifically blunts the secretion of proinflammatory cytokines in macrophages [22, 23]. Due to its role beyond anti-hyperglycemia in modulating several fundamental pathways disrupted during chronic diseases and aging, repurposing metformin to treat cardiovascular diseases, cognitive decline, cancers, neurodegenerative diseases and ultimately, aging as a whole, has been proposed [24–29].
Two studies have shown that short-term metformin treatment and exercise do not exhibit synergy, but work in an antagonistic manner, where metformin attenuates the insulin sensitizing effect of exercise [30, 31]. Metformin has been shown to induce physiologically subtle decreases in peak aerobic capacity evidenced by a reduction in peak oxygen uptake, peak heart rate, peak ventilation, peak respiratory exchange ratio and exercise duration [32]. However, another study in prediabetic adults concluded that metformin in combination with aerobic and resistance exercise training lowered proinsulin concentrations and increased insulin clearance [33]. Recently, Konopka et al showed in older adults that metformin blunted aerobic exercise training-induced improvements in cardiorespiratory fitness, insulin sensitivity, and prevented the gain in muscle mitochondrial respiration capacity [34]. Thus, there may be a complex interplay between molecular mechanisms of exercise adaptations and pathways affected by metformin.
We originally undertook the MASTERS Trial to test the hypothesis that metformin would act synergistically with PRT to reduce the number of exercise non-responders regularly observed among older adults by reducing muscle inflammation. However, in the MASTERS Trial, metformin plus PRT inhibited muscle growth after 14 weeks of training [35]. In vitro, acute metformin treatment of human primary myotubes undergoing electrical pulse stimulation (exercise mimetic capable of inducing myotube hypertrophy) had a repressive effect on mTORC1 signaling, and upregulated AMPK phosphorylation [35]. Thus, metformin’s impairment of cardiorespiratory fitness and muscle mass gains in physically active older adults must be reconciled with benefits associated with health span.
The purpose of this study was to profile the muscle transcriptome response to PRT with or without adjunctive metformin in the MASTERS Trial, to identify potential mechanisms contributing to the blunted hypertrophic response in older adults [35]. Previously, global gene expression studies of human skeletal muscle aging have identified mitochondrial dysfunction, extracellular matrix organization, complement activation and ribosomal pathways comprising a differentially expressed aging signature [36, 37]. Effects of PRT on this signature, with and without metformin, may identify new intervention targets that counteract sarcopenia, and provide information on potential metformin-exercise interactions in muscle that are relevant to repurposing metformin to treat aging and age-related disorders.
Results
Global gene expression changes in skeletal muscle with placebo + progressive resistance training (plaPRT) or metformin + progressive resistance training (metPRT)
Research participants were randomized in a double-blind fashion to placebo or metformin for 2 weeks followed by 14 weeks of PRT with continued drug treatment. Vastus lateralis biopsies were obtained at baseline (prior to drug treatment) and at 16 weeks (following 14 weeks of training) [35]. Fourteen weeks of placebo plus PRT (plaPRT) induced differential expression of 2048 genes (FDR-adj p-value < 0.01), with 1161 genes upregulated and 887 downregulated (Figure 1A). The combination of metformin and PRT (metPRT) resulted in differential expression of 1435 genes (FDR-adj p-value < 0.01), with 817 genes upregulated and 618 downregulated compared to baseline (Figure 1B). Principal component analysis (PCA) demonstrated a clear effect of PRT with a similar shift in the global gene expression profiles in both treatment groups (Figure 1C), with the Venn diagram illustrating an overlap of 918 genes (Figure 1D). The correlation between fold changes of the 918 common genes between the two groups was 0.97 (p-value < 2.2e-16), with no genes showing any anticorrelation (Figure 1E). Eleven hundred thirty genes were exclusively altered in plaPRT, while 517 genes were changed only in metPRT. Gene lists are included in Supplementary Table 1. We found no significant change in inflammatory pathway gene expression with either plaPRT or metPRT, which argues against our original hypothesis that metformin and PRT reduce muscle inflammation.
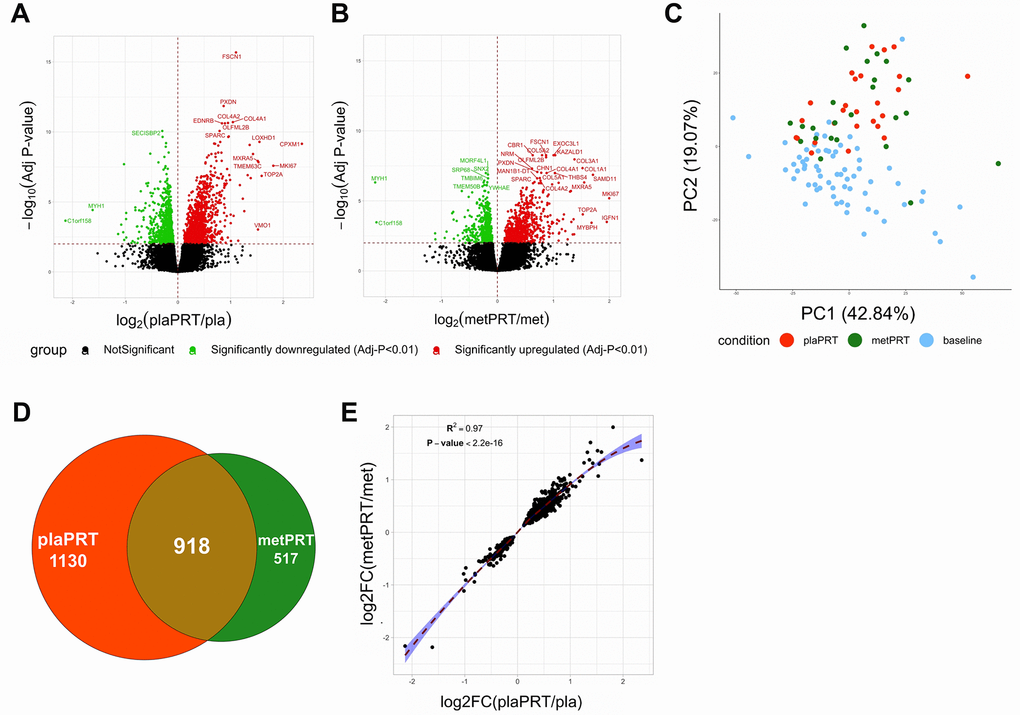
Figure 1. Metformin blunts the global transcriptomic changes induced by PRT in human skeletal muscle. (A) Volcano plot of 2048 DEG (q-value < 0.01) with plaPRT between 16 weeks and baseline; (B) Volcano plot of 1435 DEG (q-value < 0.01) with metPRT intervention between 16 weeks and baseline; (C) Principal component analysis on DEG shared between plaPRT (orange) and metPRT (green) compared to baseline (blue); (D) Venn diagram showing an overlap of DEG common between plaPRT and metPRT vs baseline; (E) Correlation plot between the fold changes of DEG common between plaPRT and metPRT.
Pathway overrepresentation within differentially expressed genes (DEG)
Both interventions modulated transcripts involved in anabolic cell signaling, extracellular matrix (ECM) organization, and RNA metabolism pathways, suggesting a conservation of these processes in response to PRT with metformin. The directionality of changes in gene expression demonstrate that ECM genes including collagen genes (COL1A1, COL3A1, COL4A1, COL6A2 etc.), ECM-receptor interaction genes, focal adhesion genes, and those encoding matrix metallopeptidases (MMP2, MMP11, MMP14) and laminin subunits were upregulated in both groups in response to training. On the other hand, genes belonging to mRNA splicing, RNA metabolism and post-transcriptional processing pathways were mostly downregulated. These included several nuclear ribonucleoproteins, serine and arginine rich splicing factors, RNA polymerase II subunit C, as well as pre-mRNA processing factors (Figures 2A, 2B, and Supplementary Table 2). However, the number of DEG was higher for all pathways in plaPRT than metPRT (Figure 2C).
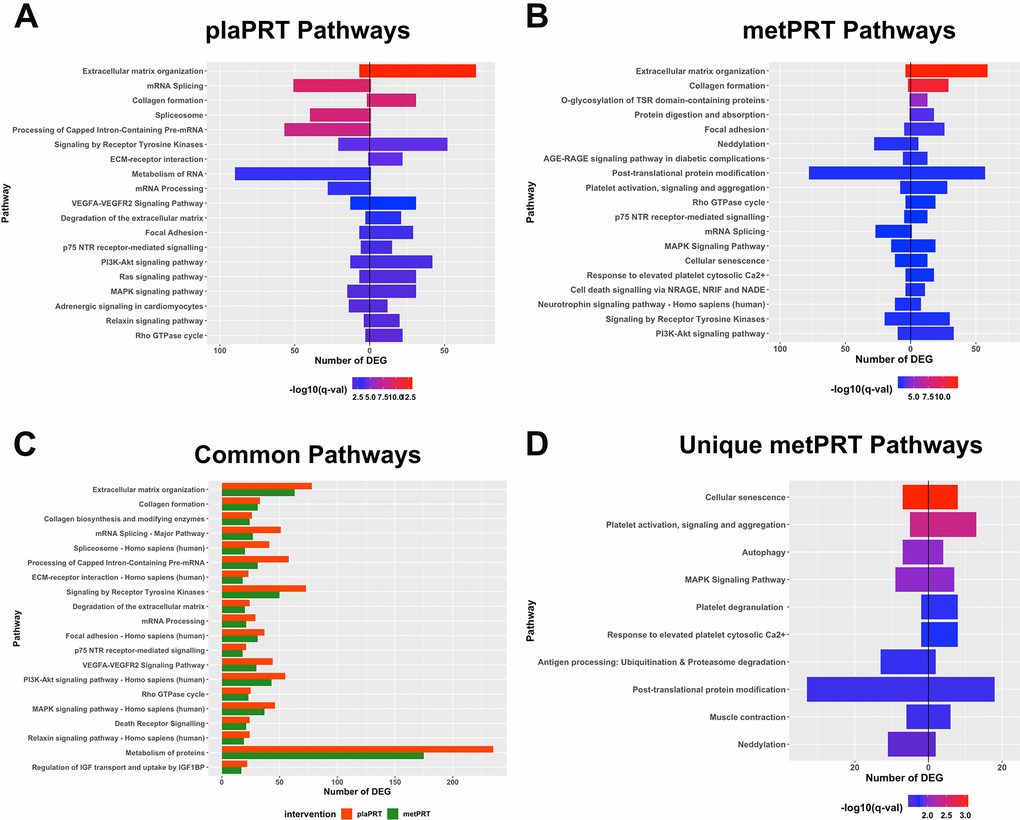
Figure 2. Pathway overrepresentation analyses on differentially expressed genes (DEG). (A) Pathways overrepresented in plaPRT-induced DEG with the length of the bar representing the number of DEG (upregulated genes to the right and downregulated genes to the left); (B) Pathways overrepresented in metPRT-induced DEG; (C) Common pathways overrepresented between the plaPRT (orange) and metPRT (green) groups; (D) Pathways overrepresented in the 517 DEG within the metPRT group that do not overlap with the DEG for the plaPRT group.
Of the 517 genes that changed exclusively in the metPRT group, pathway analyses show overrepresentation of aging hallmarks including cellular senescence, and autophagy [38], as well as post-translational modification pathways, specifically neddylation and ubiquitination (Figure 2D and Supplementary Table 3). Our previous work with short-term metformin treatment alone showed improvement in aging-induced pathways through inhibition of mTORC1 regulated genes in skeletal muscle of older adults [39]; however, it is unclear how changes in gene expression observed here in metPRT might alter the older adult muscle phenotype.
Reversal of the aging skeletal muscle transcriptome with PRT and metformin
In lieu of our findings that metPRT appeared to affect pathways associated with aging, we performed RNA-sequencing on skeletal muscle biopsies from 21 young individuals (mean age 24, 11 females, 10 males, Supplementary Table 4) under resting conditions and compared their transcriptome to that of week 0 baseline biopsies from the older adult trial participants. Our analysis revealed that 4654 DEG were present when comparing baseline biopsies (FDR-adj p-value < 0.01) with 2446 upregulated and 2208 downregulated (Figure 3A and Supplementary Table 5). Following 14 weeks of PRT, the week 16 time point from plaPRT compared to young demonstrated a dramatic decrease in DEG that were previously observed at baseline (2898 DEG), with 1089 new DEG apparent following 14 weeks of plaPRT (Figure 3B and Supplementary Table 5). MetPRT further reduced the number of DEG compared to young muscle to 2705, however, the number of unique DEG that were present after 14 weeks of PRT was also lower compared to placebo (837 DEG), consistent with our initial findings that metformin blunts the overall transcriptomic response to PRT (Figure 3B and Supplementary Table 5). Young vs old baseline DEG that were lost following PRT in both groups (1483) include RNA splicing and numerous genes involved in longevity-associated pathways (Figure 3C and Supplementary Table 6). After identifying DEG between young and old muscle at baseline that were no longer differentially expressed at week 16 in both plaPRT and metPRT, we identified 466 DEG that returned to young expression levels only in metPRT. Pathway overrepresentation analysis showed that these genes were largely involved in metabolism, particularly lipid metabolism (Supplementary Table 7).
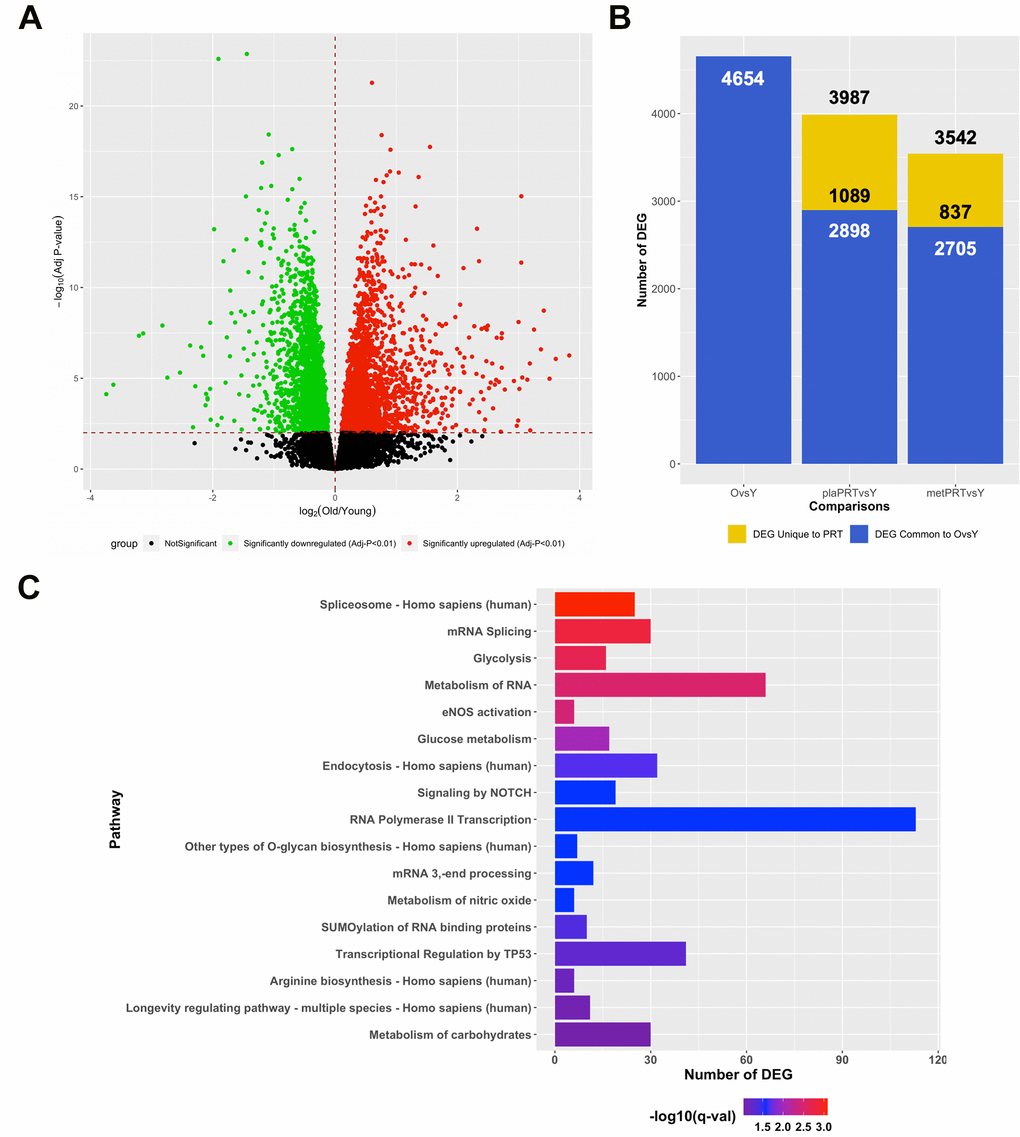
Figure 3. PRT reverts aging transcriptome towards that of young resting muscle and the effects of metformin are additive. (A) Volcano plot of 4654 DEG (q-value < 0.01) between young and old muscle at baseline; (B) Number of DEG observed when comparing young baseline skeletal muscle to old baseline skeletal muscle (4654 DEG) and young baseline to old 16 weeks with either plaPRT (3987 DEG) or metPRT (3542 DEG). Dark blue represents DEG common to throughout all time points and yellow represents those unique to week 16 time point in either plaPRT or metPRT; (C) Pathways overrepresented from those genes common to both groups following 14-weeks of PRT that were no different from young.
Discussion
In this study, we characterized the effect of metformin on the skeletal muscle transcriptomic response to PRT in older individuals from the MASTERS Trial [35]. Our results indicate that PRT induces substantial transcriptional changes in skeletal muscle with upregulation of genes involved in extracellular matrix (ECM) remodeling and downregulation of mRNA processing genes being the most affected. We also provide evidence that metformin alters skeletal muscle adaptations to PRT at the transcriptomic level, consistent with a decrease in physiological gains in response to PRT in lean body mass, and thigh muscle mass and area, as we have previously reported in this cohort [35]. Recently, the complex interaction between metformin and aerobic exercise training was described, showing that metformin also blunts improvements in physiological functions and mitochondrial adaptations otherwise promoted by aerobic exercise training in older adults [34]. Similarly, the present findings suggest that the key molecular cues underlying adaptations to PRT are attenuated by metformin. Nevertheless, we discovered gene sets differentially regulated by the combination of PRT and metformin that may promote health benefits separate from PRT-induced muscle hypertrophy.
Skeletal muscle ECM is crucial to force transmission, structural integrity and muscle stem cell dynamics [40, 41], and ECM remodeling is critical for muscle fiber growth [42]. A large number of genes associated with pathways involved in ECM composition and reorganization were upregulated in both plaPRT and metPRT groups, but to a lesser magnitude in metPRT, potentially contributing to the reduced growth response. Resistance exercise training in middle aged adults has been shown to upregulate collagens and metallopeptidases [43]. In particular, MMP2 plays an integral role in skeletal muscle hypertrophy by remodeling the ECM [44]. We found significant up-regulation of MMP2 mRNA in response to PRT in both groups, suggesting that metformin may not interfere with this specific effect of PRT that may contribute to muscle remodeling.
Genes belonging to mRNA splicing and post-transcriptional processing pathways were mostly downregulated in both groups. Aging is accompanied by a dysregulation in the splicing machinery including a rise in splicing factors, especially the heterogeneous nuclear ribonucleoproteins (hnRNPs) [45, 46]. We observed a systematic suppression of multiple hnRNPs and mRNAs encoding RNA processing and splicing factors, with down regulation of RNA processing pathways more prominent in plaPRT. Several studies have demonstrated that alternative splicing is highly enriched in energetically demanding tissues such as muscle and brain [47]. Upregulation of alternative splicing in skeletal muscle with aging may activate damage-response mechanisms at a time when energy becomes scarce [46]. Physical activity in older adults was shown to offset the changes in splicing machinery [48] and although the functional relevance has yet to be established, this effect may reduce the number of genes that are differentially spliced and the number of splicing errors that produce non-functional proteins that tend to increase with age [49].
To our knowledge only one other study has performed transcriptome analysis using microarrays to compare the effects of PRT in healthy older subjects to that of young resting skeletal muscle [50]. This seminal study illustrated that 596 genes were differentially expressed at baseline between young and old. Following 6 months of resistance exercise training a total of 179 of the 596 were no longer significantly different to that of young. Although our DEG lists are much larger, which is likely due to the greater number of older subjects at baseline between studies (65 vs 25), as well as our end point (26 plaPRT and 24 metPRT vs 14), we did find commonalities between gene lists including cell cycle inhibitors p21 and p15, as well as many metabolism related genes. Melov S et al. gene ontology analysis demonstrated that metabolic and mitochondrial function were largely impacted by aging and that resistance exercise training had a significant effect on reversing these age-related deficits [50]. Presently, we observed a similar response with PRT in both cohorts; however, adjunctive metformin further reduced the number of DEG between young and old, which appeared largely due to metformin’s effects on genes related to metabolism; pathway overrepresentation indicated a return to young gene expression patterns related to lipid metabolism preferentially in metPRT. In our primary findings of the trial, we reported an increase in AMPK/ACC phosphorylation in metPRT, that may affect lipogenesis [35]. The known inhibition of complex I by metformin [20] coupled with the increased energy demand of PRT may have led to compensatory activation of pathways involved in lipid metabolism that were previously dysregulated with age at baseline. However, it should be noted that outcomes of the MASTERS Trial showed that the PRT-induced decrease in low density muscle (which contains more intramyocellular lipid than normal density muscle) was similar between groups, whereas plaPRT gained significantly more normal density muscle area than metPRT [35]. Thus, the combined effects of metformin and exercise on muscle density and lipid content require further study.
A few limitations reduce the scope of our interpretation of the data, including the time between the last bout of exercise and tissue collection (3 days). It is possible that metformin inhibited mTOR signaling acutely after each exercise bout, impairing growth, but did not affect the new homeostasis following training. Although we reported down-regulation of mTOR-associated pathways in muscle following 6 weeks of metformin alone in a crossover study design in older adults [51], chronic effects of metformin on mTOR regulated pathways were less apparent within the context of exercise training. Another limitation acknowledged in our primary outcomes paper is the lack of a sedentary control group given metformin for 16 weeks. We attempted to extrapolate metformin-specific effects, but without a sedentary group, it is not possible to assess effects due to metformin or the combined effects of PRT + metformin on gene expression.
In conclusion, the blunted transcriptomic response to PRT in the presence of metformin is consistent with the blunted muscle hypertrophic growth response reported in the primary outcome of the MASTERS Trial [35]. However, specific effects of adjunctive metformin on the muscle transcriptomic response to PRT, separate from those related to muscle remodeling, may alter age-associated deficits in muscle metabolism to improve function. Metformin and PRT have beneficial effects on health that extend well-beyond skeletal muscle that should be considered, but a metformin-exercise interaction in muscle must be defined in more detail to inform recommending metformin for healthy, physically active older adults.
Materials and Methods
Study design, participants and interventions
The Metformin to Augment Strength Training Effective Response in Seniors (MASTERS) Trial is a randomized, controlled, double blind trial comparing the effects of metformin versus placebo during a 14 week progressive resistance exercise training (PRT) intervention in healthy men and women ≥ 65 years of age. Participants were recruited at University of Kentucky and University of Alabama at Birmingham, UAB. The detailed study design [52] and participant characteristics [35] have been published previously.
Participants were randomized to receive either placebo or metformin for the duration of the trial. Subjects underwent a two week drug or placebo wash-in period prior to beginning PRT. Those who were randomized to metformin were titrated up to the target dose by taking 1 tablet per day (850 mg) for 7 days, followed by 2 tablets per day (1700 mg) for the remainder of the trial.
All study subjects underwent 14 weeks of PRT, supervised by trained personnel. We employed a variable intensity prescription across the three training days each week (high/low/high) based on the results of our previous dose-response trial which showed this prescription optimized strength and muscle mass gains in older adults [53]. Vastus lateralis muscle biopsies were obtained prior to drug initiation, after the 2 week wash-in period and 3 days after the final bout of training. Primary outcomes of the trial, muscle size and strength, have been reported [35]. Vastus lateralis muscle biopsies from young individuals matched for body mass index (Supplementary Table 4) were obtained through the Center for Muscle Biology at the University of Kentucky.
Library preparation
Total RNA was isolated from baseline muscle biopsies in 37 plaPRT and 28 metPRT participants and from 16-week post-training muscle biopsies from 26 plaPRT and 24 metPRT participants (average age 71 years old). Of these, 24 plaPRT and 23 metPRT participants had biopsies at both timepoints. Additionally, total RNA was isolated from muscle biopsies in 21 young healthy donors (average age 24 years old). Approximately 35 mg of muscle was subjected to bead homogenization in Qiazol (Qiagen, Valencia, CA) and RNA purified using miRNeasy Mini Kits (Qiagen) and stored at -80° C. RNA content, integrity and purity were determined with a Nanodrop 2000 spectrophotometer (Thermo Fisher, Waltham, MA) and the 2100 Bioanalyzer (Agilent, Santa Clara, CA). A minimum RNA Integrity Number (RIN) of 6.5 was set for all samples.
Sequencing, preprocessing and alignment
Total RNA was sequenced at Novogene Corporation, Chula Vista, CA on an Illumina HiSeq 4000 system, using a standard paired-end 150 bp (PE150) dual indexing protocol. The two sets of samples (University of Kentucky and University of Alabama), each containing samples from all timepoints of both placebo and metformin arm as well as young, were sequenced in different batches that were corrected for in the downstream analyses. Raw fastq sequence reads were passed through quality control using FastQC (0.11.4) [54] and the QC results were compiled for all samples using MultiQC (1.7) [55]. Due to adapter contamination, the raw fastq files were trimmed for adapter sequences, filtered for low quality reads and too short reads, using the default parameters in fastp (0.19.4) - an all-in-one preprocessing tool for fastq files [56]. After checking for QC using the same steps as before, RNA-Sequencing by Expectation Maximization- RSEM (1.3.0) in conjunction with the STAR aligner (2.6.1b) was used to align the raw reads to the GRCh38 primary assembly build of the reference human genome, with transcript annotations (gencode.v29.annotation.gtf) downloaded from GENCODE [57–59].
Differential gene expression analysis
All statistical analysis for the gene expression data were carried out using the R statistical software (R-3.6.0). The raw counts were filtered for low expression using a counts-per-million (cpm) threshold of 0.6 (10/minimum library size) in at least 24 (number of samples in the smallest group of comparison). Since the principal component analyses revealed a distinct sequencing batch effect, the raw count data was corrected for it using batch as a covariate in the generalized linear model in limma (3.4.0) [60]. The corrected data using the removeBatchEffect function in limma was deemed to have minimal sequencing batch effects after a visual inspection of PCA and used for all downstream analyses. Raw data were normalized using the trimmed mean of M-values (TMM) normalization. To minimize heteroscedasticity from the count data and incorporate precision weights to account for the mean-variance relationship, the voom function was applied on the normalized data [60]. A linear model was fit on the voom-normalized data, while adjusting for the study arm (placebo vs metformin) and biopsy time (baseline, 14 weeks of PRT with treatment and young). Due to the paired nature of the study, the participant id was used as a blocking variable. Differential gene expression was measured using an Empirical Bayes statistic in limma for the following comparisons –plaPRT vs Baseline, metPRT vs Baseline, Young vs Old Baseline, Young vs plaPRT (16 weeks), and Young vs metPRT (16 weeks). The raw P-values were adjusted for multiple comparisons using the Benjamini-Hochberg correction. The adjusted P-value threshold of 0.01 was used to characterize statistically significant differentially expressed genes (DEG).
Pathway overrepresentation analysis
Genes that were deemed to be differentially expressed with statistical significance (FDR-adj P-value < 0.01), were exported to ConsensusPathdb (http://cpdb.molgen.mpg.de/CPDB) database [61]. The pathways interrogated were Reactome, KEGG, Biocarta, Wikipathways and PharmGKB. A hypergeometric test was run and pathways with a gene overlap threshold of 10% of all input genes and the p-value cutoff of 0.01 were included in the output. Duplicated pathways were excluded from the output.
Data availability statement
The raw RNA-Seq and count data from this experiment will be accessible in the Gene Expression Omnibus database (GEO Accession code- GSE157585).
Supplementary Materials
Author Contributions
A.S.K., B.P., N.B. and C.A.P. conceived the project. B.P. and R.G.W. performed RNA isolation and analyses. B.P., R.G.W., P.A.K., S.T.W., M.M.B. and C.A.P. designed and executed the MASTERS Trial. A.S.K. and J.C.M. carried out the RNA-seq computational analysis. A.S.K, B.P., N.B. and C.A.P. interpreted the data and wrote the manuscript.
Conflicts of Interest
There are no conflicts of interest in any aspect of this study. We declare that the results of the study are presented clearly, honestly, and without fabrication, falsification, or inappropriate data manipulation.
Funding
This work is supported by the Nathan Shock Center of Excellence in the basic Biology of Aging (P30AG038072) (N.B.), the Einstein-Paul Glenn Foundation for Medical Research Center for the Biology of Human Aging (N.B.), a Paul Glenn Investigator Award (C.A.P.), and NIH grant R01AG046920 (P.A.K., M.M.B. and C.A.P.) and the University of Kentucky (UL1TR001998) and the University of Alabama, Birmingham (UL1TR003096) NIH CTSAs.
References
- 1. Larsson L, Degens H, Li M, Salviati L, Lee YI, Thompson W, Kirkland JL, Sandri M. Sarcopenia: aging-related loss of muscle mass and function. Physiol Rev. 2019; 99:427–511. https://doi.org/10.1152/physrev.00061.2017 [PubMed]
- 2. Han A, Bokshan SL, Marcaccio SE, DePasse JM, Daniels AH. Diagnostic Criteria and Clinical Outcomes in Sarcopenia Research: A Literature Review. J Clin Med. 2018; 7:70. https://doi.org/10.3390/jcm7040070 [PubMed]
- 3. Rosenberg IH. Sarcopenia: origins and clinical relevance. Clin Geriatr Med. 2011; 27:337–39. https://doi.org/10.1016/j.cger.2011.03.003 [PubMed]
- 4. Cruz-Jentoft AJ, Sayer AA. Sarcopenia. Lancet. 2019; 393:2636–46. https://doi.org/10.1016/S0140-6736(19)31138-9 [PubMed]
- 5. Tieland M, Trouwborst I, Clark BC. Skeletal muscle performance and ageing. J Cachexia Sarcopenia Muscle. 2018; 9:3–19. https://doi.org/10.1002/jcsm.12238 [PubMed]
- 6. Hughes VA, Frontera WR, Roubenoff R, Evans WJ, Singh MA. Longitudinal changes in body composition in older men and women: role of body weight change and physical activity. Am J Clin Nutr. 2002; 76:473–81. https://doi.org/10.1093/ajcn/76.2.473 [PubMed]
- 7. Kim TN, Choi KM. Sarcopenia: definition, epidemiology, and pathophysiology. J Bone Metab. 2013; 20:1–10. https://doi.org/10.11005/jbm.2013.20.1.1 [PubMed]
- 8. Distefano G, Goodpaster BH. Effects of exercise and aging on skeletal muscle. Cold Spring Harb Perspect Med. 2018; 8:a029785. https://doi.org/10.1101/cshperspect.a029785 [PubMed]
- 9. Gheller BJ, Riddle ES, Lem MR, Thalacker-Mercer AE. Understanding age-related changes in skeletal muscle metabolism: differences between females and males. Annu Rev Nutr. 2016; 36:129–56. https://doi.org/10.1146/annurev-nutr-071715-050901 [PubMed]
- 10. Petrella JK, Kim JS, Cross JM, Kosek DJ, Bamman MM. Efficacy of myonuclear addition may explain differential myofiber growth among resistance-trained young and older men and women. Am J Physiol Endocrinol Metab. 2006; 291:E937–46. https://doi.org/10.1152/ajpendo.00190.2006 [PubMed]
- 11. Lavin KM, Roberts BM, Fry CS, Moro T, Rasmussen BB, Bamman MM. The importance of resistance exercise training to combat neuromuscular aging. Physiology (Bethesda). 2019; 34:112–22. https://doi.org/10.1152/physiol.00044.2018 [PubMed]
- 12. Gharahdaghi N, Rudrappa S, Brook MS, Idris I, Crossland H, Hamrock C, Abdul Aziz MH, Kadi F, Tarum J, Greenhaff PL, Constantin-Teodosiu D, Cegielski J, Phillips BE, et al. Testosterone therapy induces molecular programming augmenting physiological adaptations to resistance exercise in older men. J Cachexia Sarcopenia Muscle. 2019; 10:1276–94. https://doi.org/10.1002/jcsm.12472 [PubMed]
- 13. Jakubowski JS, Wong EP, Nunes EA, Noguchi KS, Vandeweerd JK, Murphy KT, Morton RW, McGlory C, Phillips SM. Equivalent hypertrophy and strength gains in β-hydroxy-β-methylbutyrate- or leucine-supplemented men. Med Sci Sports Exerc. 2019; 51:65–74. https://doi.org/10.1249/MSS.0000000000001752 [PubMed]
- 14. Sakashita M, Nakamura U, Horie N, Yokoyama Y, Kim M, Fujita S. Oral supplementation using gamma-aminobutyric acid and whey protein improves whole body fat-free mass in men after resistance training. J Clin Med Res. 2019; 11:428–34. https://doi.org/10.14740/jocmr3817 [PubMed]
- 15. McKee A, Morley JE, Matsumoto AM, Vinik A. Sarcopenia: an endocrine disorder? Endocr Pract. 2017; 23:1140–49. https://doi.org/10.4158/EP171795.RA [PubMed]
- 16. Boulé NG, Robert C, Bell GJ, Johnson ST, Bell RC, Lewanczuk RZ, Gabr RQ, Brocks DR. Metformin and exercise in type 2 diabetes: examining treatment modality interactions. Diabetes Care. 2011; 34:1469–74. https://doi.org/10.2337/dc10-2207 [PubMed]
- 17. Malin SK, Braun B. Impact of metformin on exercise-induced metabolic adaptations to lower type 2 diabetes risk. Exerc Sport Sci Rev. 2016; 44:4–11. https://doi.org/10.1249/JES.0000000000000070 [PubMed]
- 18. Baptista LC, Machado-Rodrigues AM, Martins RA. Exercise but not metformin improves health-related quality of life and mood states in older adults with type 2 diabetes. Eur J Sport Sci. 2017; 17:794–804. https://doi.org/10.1080/17461391.2017.1310933 [PubMed]
- 19. Flory J, Lipska K. Metformin in 2019. JAMA. 2019; 321:1926–27. https://doi.org/10.1001/jama.2019.3805 [PubMed]
- 20. Fontaine E. Metformin-induced mitochondrial complex I inhibition: facts, uncertainties, and consequences. Front Endocrinol (Lausanne). 2018; 9:753. https://doi.org/10.3389/fendo.2018.00753 [PubMed]
- 21. Hawley SA, Gadalla AE, Olsen GS, Hardie DG. The antidiabetic drug metformin activates the AMP-activated protein kinase cascade via an adenine nucleotide-independent mechanism. Diabetes. 2002; 51:2420–25. https://doi.org/10.2337/diabetes.51.8.2420 [PubMed]
- 22. Cameron AR, Morrison VL, Levin D, Mohan M, Forteath C, Beall C, McNeilly AD, Balfour DJ, Savinko T, Wong AK, Viollet B, Sakamoto K, Fagerholm SC, et al. Anti-inflammatory effects of metformin irrespective of diabetes status. Circ Res. 2016; 119:652–65. https://doi.org/10.1161/CIRCRESAHA.116.308445 [PubMed]
- 23. Rena G, Hardie DG, Pearson ER. The mechanisms of action of metformin. Diabetologia. 2017; 60:1577–85. https://doi.org/10.1007/s00125-017-4342-z [PubMed]
- 24. Valencia WM, Palacio A, Tamariz L, Florez H. Metformin and ageing: improving ageing outcomes beyond glycaemic control. Diabetologia. 2017; 60:1630–38. https://doi.org/10.1007/s00125-017-4349-5 [PubMed]
- 25. Rena G, Lang CC. Repurposing metformin for cardiovascular disease. Circulation. 2018; 137:422–24. https://doi.org/10.1161/CIRCULATIONAHA.117.031735 [PubMed]
- 26. Heckman-Stoddard BM, DeCensi A, Sahasrabuddhe VV, Ford LG. Repurposing metformin for the prevention of cancer and cancer recurrence. Diabetologia. 2017; 60:1639–47. https://doi.org/10.1007/s00125-017-4372-6 [PubMed]
- 27. Rotermund C, Machetanz G, Fitzgerald JC. The therapeutic potential of metformin in neurodegenerative diseases. Front Endocrinol (Lausanne). 2018; 9:400. https://doi.org/10.3389/fendo.2018.00400 [PubMed]
- 28. Barzilai N, Crandall JP, Kritchevsky SB, Espeland MA. Metformin as a tool to target aging. Cell Metab. 2016; 23:1060–65. https://doi.org/10.1016/j.cmet.2016.05.011 [PubMed]
- 29. Farr SA, Roesler E, Niehoff ML, Roby DA, McKee A, Morley JE. Metformin improves learning and memory in the SAMP8 mouse model of Alzheimer’s disease. J Alzheimers Dis. 2019; 68:1699–710. https://doi.org/10.3233/JAD-181240 [PubMed]
- 30. Malin SK, Gerber R, Chipkin SR, Braun B. Independent and combined effects of exercise training and metformin on insulin sensitivity in individuals with prediabetes. Diabetes Care. 2012; 35:131–36. https://doi.org/10.2337/dc11-0925 [PubMed]
- 31. Sharoff CG, Hagobian TA, Malin SK, Chipkin SR, Yu H, Hirshman MF, Goodyear LJ, Braun B. Combining short-term metformin treatment and one bout of exercise does not increase insulin action in insulin-resistant individuals. Am J Physiol Endocrinol Metab. 2010; 298:E815–23. https://doi.org/10.1152/ajpendo.00517.2009 [PubMed]
- 32. Braun B, Eze P, Stephens BR, Hagobian TA, Sharoff CG, Chipkin SR, Goldstein B. Impact of metformin on peak aerobic capacity. Appl Physiol Nutr Metab. 2008; 33:61–67. https://doi.org/10.1139/H07-144 [PubMed]
- 33. Viskochil R, Malin SK, Blankenship JM, Braun B. Exercise training and metformin, but not exercise training alone, decreases insulin production and increases insulin clearance in adults with prediabetes. J Appl Physiol (1985). 2017; 123:243–48. https://doi.org/10.1152/japplphysiol.00790.2016 [PubMed]
- 34. Konopka AR, Laurin JL, Schoenberg HM, Reid JJ, Castor WM, Wolff CA, Musci RV, Safairad OD, Linden MA, Biela LM, Bailey SM, Hamilton KL, Miller BF. Metformin inhibits mitochondrial adaptations to aerobic exercise training in older adults. Aging Cell. 2019; 18:e12880. https://doi.org/10.1111/acel.12880 [PubMed]
- 35. Walton RG, Dungan CM, Long DE, Tuggle SC, Kosmac K, Peck BD, Bush HM, Villasante Tezanos AG, McGwin G, Windham ST, Ovalle F, Bamman MM, Kern PA, Peterson CA. Metformin blunts muscle hypertrophy in response to progressive resistance exercise training in older adults: a randomized, double-blind, placebo-controlled, multicenter trial: the MASTERS trial. Aging Cell. 2019; 18:e13039. https://doi.org/10.1111/acel.13039 [PubMed]
- 36. Zahn JM, Sonu R, Vogel H, Crane E, Mazan-Mamczarz K, Rabkin R, Davis RW, Becker KG, Owen AB, Kim SK. Transcriptional profiling of aging in human muscle reveals a common aging signature. PLoS Genet. 2006; 2:e115. https://doi.org/10.1371/journal.pgen.0020115.eor [PubMed]
- 37. Su J, Ekman C, Oskolkov N, Lahti L, Ström K, Brazma A, Groop L, Rung J, Hansson O. A novel atlas of gene expression in human skeletal muscle reveals molecular changes associated with aging. Skelet Muscle. 2015; 5:35. https://doi.org/10.1186/s13395-015-0059-1 [PubMed]
- 38. López-Otín C, Blasco MA, Partridge L, Serrano M, Kroemer G. The hallmarks of aging. Cell. 2013; 153:1194–217. https://doi.org/10.1016/j.cell.2013.05.039 [PubMed]
- 39. Kulkarni AS, Gubbi S, Barzilai N. Benefits of metformin in attenuating the hallmarks of aging. Cell Metab. 2020; 32:15–30. https://doi.org/10.1016/j.cmet.2020.04.001 [PubMed]
- 40. Rayagiri SS, Ranaldi D, Raven A, Mohamad Azhar NI, Lefebvre O, Zammit PS, Borycki AG. Basal lamina remodeling at the skeletal muscle stem cell niche mediates stem cell self-renewal. Nat Commun. 2018; 9:1075. https://doi.org/10.1038/s41467-018-03425-3 [PubMed]
- 41. Kjaer M. Role of extracellular matrix in adaptation of tendon and skeletal muscle to mechanical loading. Physiol Rev. 2004; 84:649–98. https://doi.org/10.1152/physrev.00031.2003 [PubMed]
- 42. Fry CS, Kirby TJ, Kosmac K, McCarthy JJ, Peterson CA. Myogenic progenitor cells control extracellular matrix production by fibroblasts during skeletal muscle hypertrophy. Cell Stem Cell. 2017; 20:56–69. https://doi.org/10.1016/j.stem.2016.09.010 [PubMed]
- 43. Hjorth M, Norheim F, Meen AJ, Pourteymour S, Lee S, Holen T, Jensen J, Birkeland KI, Martinov VN, Langleite TM, Eckardt K, Drevon CA, Kolset SO. The effect of acute and long-term physical activity on extracellular matrix and serglycin in human skeletal muscle. Physiol Rep. 2015; 3:e12473. https://doi.org/10.14814/phy2.12473 [PubMed]
- 44. Zhang Q, Joshi SK, Lovett DH, Zhang B, Bodine S, Kim HT, Liu X. Matrix metalloproteinase-2 plays a critical role in overload induced skeletal muscle hypertrophy. Muscles Ligaments Tendons J. 2015; 4:446–54. [PubMed]
- 45. Welle S, Brooks AI, Delehanty JM, Needler N, Bhatt K, Shah B, Thornton CA. Skeletal muscle gene expression profiles in 20-29 year old and 65-71 year old women. Exp Gerontol. 2004; 39:369–77. https://doi.org/10.1016/j.exger.2003.11.011 [PubMed]
- 46. Ubaida-Mohien C, Lyashkov A, Gonzalez-Freire M, Tharakan R, Shardell M, Moaddel R, Semba RD, Chia CW, Gorospe M, Sen R, Ferrucci L. Discovery proteomics in aging human skeletal muscle finds change in spliceosome, immunity, proteostasis and mitochondria. Elife. 2019; 8:e49874. https://doi.org/10.7554/eLife.49874 [PubMed]
- 47. Pan Q, Shai O, Lee LJ, Frey BJ, Blencowe BJ. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat Genet. 2008; 40:1413–15. https://doi.org/10.1038/ng.259 [PubMed]
- 48. Ubaida-Mohien C, Gonzalez-Freire M, Lyashkov A, Moaddel R, Chia CW, Simonsick EM, Sen R, Ferrucci L. Physical activity associated proteomics of skeletal muscle: being physically active in daily life may protect skeletal muscle from aging. Front Physiol. 2019; 10:312. https://doi.org/10.3389/fphys.2019.00312 [PubMed]
- 49. Rodríguez SA, Grochová D, McKenna T, Borate B, Trivedi NS, Erdos MR, Eriksson M. Global genome splicing analysis reveals an increased number of alternatively spliced genes with aging. Aging Cell. 2016; 15:267–78. https://doi.org/10.1111/acel.12433 [PubMed]
- 50. Melov S, Tarnopolsky MA, Beckman K, Felkey K, Hubbard A. Resistance exercise reverses aging in human skeletal muscle. PLoS One. 2007; 2:e465. https://doi.org/10.1371/journal.pone.0000465 [PubMed]
- 51. Kulkarni AS, Brutsaert EF, Anghel V, Zhang K, Bloomgarden N, Pollak M, Mar JC, Hawkins M, Crandall JP, Barzilai N. Metformin regulates metabolic and nonmetabolic pathways in skeletal muscle and subcutaneous adipose tissues of older adults. Aging Cell. 2018; 17:e12723. https://doi.org/10.1111/acel.12723 [PubMed]
- 52. Long DE, Peck BD, Martz JL, Tuggle SC, Bush HM, McGwin G, Kern PA, Bamman MM, Peterson CA. Metformin to augment strength training effective response in seniors (MASTERS): study protocol for a randomized controlled trial. Trials. 2017; 18:192. https://doi.org/10.1186/s13063-017-1932-5 [PubMed]
- 53. Stec MJ, Thalacker-Mercer A, Mayhew DL, Kelly NA, Tuggle SC, Merritt EK, Brown CJ, Windham ST, Dell’Italia LJ, Bickel CS, Roberts BM, Vaughn KM, Isakova-Donahue I, et al. Randomized, four-arm, dose-response clinical trial to optimize resistance exercise training for older adults with age-related muscle atrophy. Exp Gerontol. 2017; 99:98–109. https://doi.org/10.1016/j.exger.2017.09.018 [PubMed]
- 54. Andrews S. FastQC: a quality control tool for high throughput sequence data. 2010. https://www.bioinformatics.babraham.ac.uk/projects/fastqc/.
- 55. Ewels P, Magnusson M, Lundin S, Käller M. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics. 2016; 32:3047–48. https://doi.org/10.1093/bioinformatics/btw354 [PubMed]
- 56. Chen S, Zhou Y, Chen Y, Gu J. Fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics. 2018; 34:i884–90. https://doi.org/10.1093/bioinformatics/bty560 [PubMed]
- 57. Li B, Dewey CN. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinformatics. 2011; 12:323. https://doi.org/10.1186/1471-2105-12-323 [PubMed]
- 58. Harrow J, Frankish A, Gonzalez JM, Tapanari E, Diekhans M, Kokocinski F, Aken BL, Barrell D, Zadissa A, Searle S, Barnes I, Bignell A, Boychenko V, et al. GENCODE: the reference human genome annotation for the ENCODE project. Genome Res. 2012; 22:1760–74. https://doi.org/10.1101/gr.135350.111 [PubMed]
- 59. Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, Batut P, Chaisson M, Gingeras TR. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013; 29:15–21. https://doi.org/10.1093/bioinformatics/bts635 [PubMed]
- 60. Law CW, Chen Y, Shi W, Smyth GK. Voom: precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 2014; 15:R29. https://doi.org/10.1186/gb-2014-15-2-r29 [PubMed]
- 61. Herwig R, Hardt C, Lienhard M, Kamburov A. Analyzing and interpreting genome data at the network level with ConsensusPathDB. Nat Protoc. 2016; 11:1889–907. https://doi.org/10.1038/nprot.2016.117 [PubMed]



