Introduction
Lung cancer remains one of the most common cancers in men worldwide [1, 2]. In the United States, between 2008 and 2014, the 5-year relative survival rate for lung cancer was only 19% [2]. Lung cancer includes multiple subtypes, and the proportion of lung adenocarcinoma (LUAD) has increased in recent years. Despite significant improvements in chemotherapy and molecular targeted therapy, the survival rate of LUAD is still unsatisfactory. Tumor recurrence and metastasis are major problems in the clinical treatment of LUAD [3]. Drug resistance to common gene targeted drugs and therapeutic effect differences between different patients can occur [4]. Pabla, Conroy [5] found that many genes represented by KIAA0101 are immune related genes of adenocarcinoma, and KIAA0101 and other genes might affect the presentation of tumor immune antigens. However, the specific relationship between KIAA0101 and immune escape is not clear. In this study, we found that the expression of KIAA0101 correlated negatively with a proportion of tumor infiltrating lymphocytes, and the expression of KIAA0101 was correlated negatively with MHC molecule expression. Therefore, we carried out a study on the relationship between the expression of KIAA0101 and immune escape in lung adenocarcinoma, and further studied the effect of KIAA0101 expression on immunotherapy of lung cancer. Our study showed that overexpression of KIAA0101 promotes immune escape in lung adenocarcinoma. Therefore, searching for novel molecular biomarkers and improving immunotherapy of tumors in the diagnosis and treatment of LUAD are important [6].
KIAA0101 is a proliferating cell nuclear antigen (PCNA) binding factor discovered by Yu et al. in 2001 [7] using yeast two-hybrid experiments. It has different functions to other binding factors of PCNA, such as p21 and p57, in that KIAA0101 does not inhibit DNA replication and cell cycle process. PCNA-binding proteins act as regulators of DNA repair during DNA replication. Following DNA damage, they also facilitate the bypass of replication-fork-blocking lesions. KIAA0101 also acts as a regulator of centrosome number. However, the role of KIAA0101 in lung adenocarcinoma has not been determined. The protein encoded by KIAA0101 contains eight domain chains (Supplementary Table 1) [8]. We noted that KIAA0101 expression in lung adenocarcinoma was confirmed using an antibody in The Human Protein Atlas (Supplementary Figure 1) [9]. The expression of KIAA0101 in the nucleus was significantly increased and slightly enhanced in the cytoplasm. The expression of KIAA0101 increased in a variety of cancer tissues and cells, especially in liver cancer [10], breast cancer [11, 12], gastric cancer [13], and other cancers.
The present study aimed to explore the relationship between KIAA0101 expression and the prognosis and immune infiltration of LUAD. We also searched for KIAA0101-related gene regulatory networks. KIAA0101 expression was analyzed via the Oncomine, GEPIA, and Ualcan databases. The prognosis of LUAD related to KIAA0101 was evaluated using prognostan and other databases. Analysis of tumor immune infiltration related to KIAA0101 was analyzed using TISIDB. Gene set enrichment analysis was performed using Linkedmics. We hoped to provide new biomarkers related to KIAA0101 for future targeted therapy of LUAD. In addition, we assessed whether we could predict the survival of patients with LUAD based on the expression of KIAA0101. The discovery of the gene regulatory network of KIAA0101 will open new avenues for further studies on the epigenetics of KIAA0101. The study of immune infiltration related to KIAA0101 will provide new directions for immunotherapy of lung adenocarcinoma.
Results
Differential expression of KIAA0101 in lung adenocarcinoma
Figure 1A shows a heat map of multiple genes that were overexpressed in LUAD. The top 23 ranked differentially expressed genes were obtained based on the log2 median centered ratio. After a genome-wide analysis, we identified these 23 genes, including KIAA0101, which were overexpressed in LUAD (Figure 1A, red text genes). We used Oncomine to analyze the expression distribution of KIAA0101 mRNA in different tumors (Figure 1B). Oncomine collected these datasets from public repositories such as Gene Expression Omnibus (GEO) and Array Express. The results showed that when comparing cancer samples with normal samples, KIAA0101 was overexpressed in 17 LUAD datasets, but was not under-expressed in any of the LUAD datasets (Figure 1B). The thresholds were a P value less than 0.001, fold change greater than two, and gene rank in the top 10%. All results were statistically significant. The results were analyzed statistically using Student's t test. The mRNA expression level of KIAA0101 was significantly higher in LUAD than in normal tissues. Figure 1C1–1C8 show the data from Oncomine, GEPIA, and Ualcan comparisons of KIAA0101 mRNA levels between eight different gene sets of adenocarcinoma and normal tissues. All the results showed that the expression level of KIAA0101 in LUAD tissues was significantly higher than that in normal lung tissues, as assessed using Student's t test. Further analysis of various clinical features of LUAD in the TCGA showed that KIAA0101 was highly transcribed. In subgroup analysis based on stage, race, sex, age, smoking habit, and metastasis in the UALCAN database, the transcription level of KIAA0101 in patients with LUAD was significantly higher than that in healthy people (Supplementary Figure 14A–14F). Therefore, the expression of KIAA0101 could be used as a potential diagnostic marker for LUAD [14].
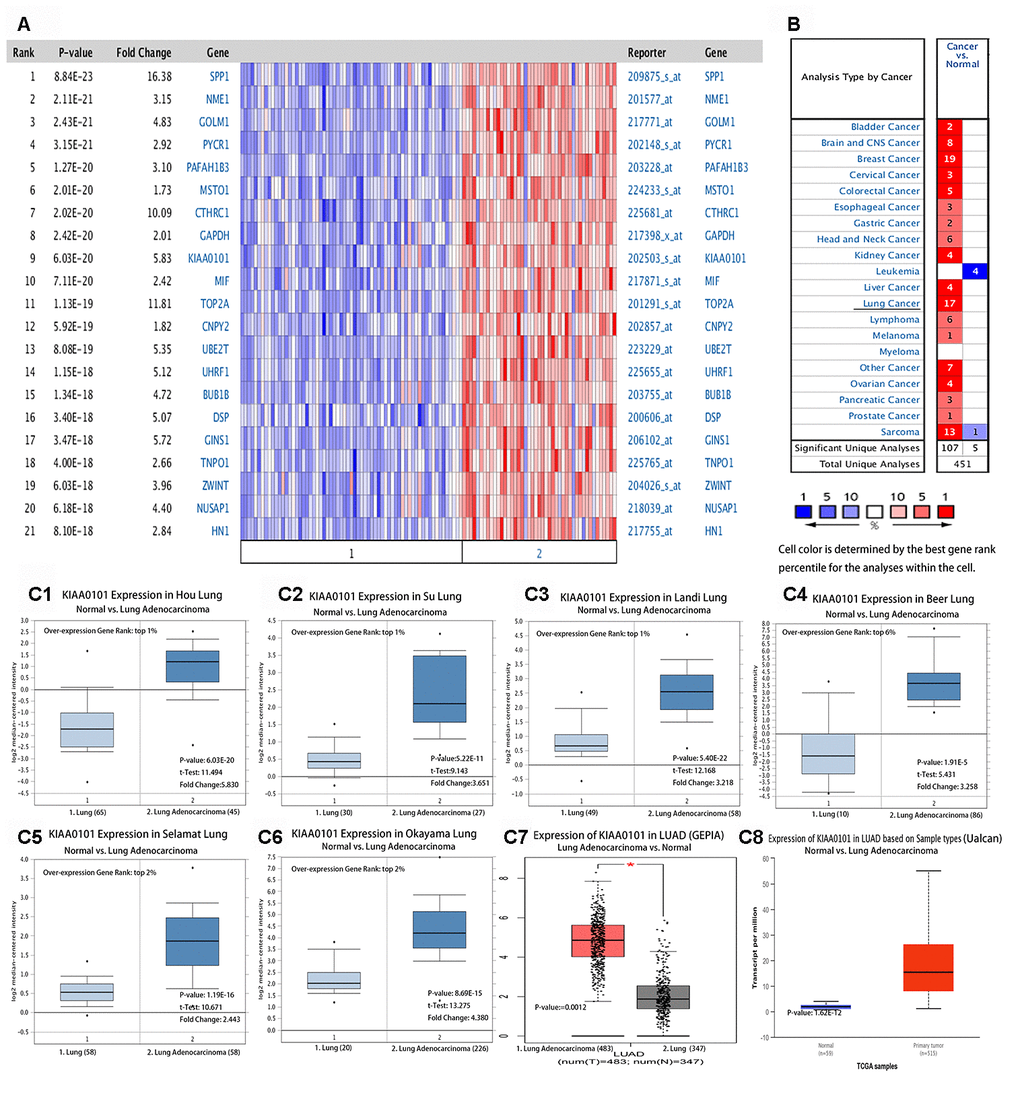
Figure 1. (A) Multigene view of lung adenocarcinoma Heat Maps Comparison of all Genes in the study of Hou lung [50]. The expression of KIAA0101 in lung adenocarcinoma was higher than that in the normal control group. (B) Summary view of KIAA0101. The transcription level of KIAA0101 in different types of cancer. Parameter setting: gene: KIAA0101, threshold (P-value): 0.001, threshold (fold change):2, threshold (gene rank): 10%, data type: DNA and mRNA. Note: The color is standardized by the Z-score to describe the relative value in the row. They cannot be used to compare values between rows. Among them, Red signifies gene overexpression or copy gain in the analyses represented by that cell in the table; blue represents the gene's underexpression or copy loss in those analyses. Datasets comprised samples represented as microarray data measuring either mRNA expression on primary tumors, cell lines, or xenografts. (C) Transcription of KIAA0101 in lung adenocarcinoma (from Oncomine, GEPIA, and Ualcan). mRNA expression levels of KIAA0101 were significantly higher in lung adenocarcinoma than in normal tissue. (C1–C6) Shown are the fold change, associated p-values, and overexpression Gene Rank, based on Oncomine 4.5 analysis. Box plot showing KIAA0101 mRNA levels in, respectively, the Hou Lung, Su Lung, Landi Lung, Beer Lung, Selamat Lung and Okayama Lung datasets [50]. (C7) Expression of KIAA0101 in LUAD based on GEPIA analysis; the p-value was 0.0012. (C8) Shows the expression of KIAA0101 in LUAD based on Ualcan analysis; the p-value was 1.62E-12.
KIAA0101 mRNA levels predict prognosis in patients with lung adenocarcinoma
Our next analysis showed that the expression of KIAA0101 had a significant effect on the prognosis of LUAD (Table 1). Figure 2A1–2A6 show the overall survival curve of five datasets (Jacob-00182-CANDF, MICHIGAN-LC, Jacob-00182-MSK, GSE13213, GSE31210) and the relapse free survival curve of GSE31210 datasets. (Figure 2A1–2A6) show six survival curves representing the five different data sets in Table 1 (from the PrognoScan[15] database). We could observe the blue curves (low expression) were higher than the red curves (high expression) in the Kaplan-Meier plots (Figure 2A1–2A6). In addition, we could see from Table 1 that all six Hazard Ratios (HR) were greater than 1.4, the HR [95% confidence interval (CI)-low CI-up] range was greater than one. The Cox p-values were less than 0.05, indicating statistical significance. The results showed that low expression of KIAA0101 was associated with a good prognosis in patients with LUAD. Figure 2B1–2B6 show the overall survival curves of patients using data from the GEPIA [16], LinkedOmics [17], Ualcan [14], TISIDB [18], OncoLnc [19], and TCGA portal [20] databases. We found that patients with LUAD with low KIAA0101 expression had a better prognosis than those with high KIAA0101 expression in six the overall survival (OS) curves (P-value < 0.01). Figure 2C1 and 2C2 show the disease free survival curves (DFS) of patients using data from the GEPIA database. Figure 2D1 and 2D2 show the progression free survival curves (PFS) of patients using data from Kaplan Meier-plotter. We found that patients with LUAD with low KIAA0101 expression had a better prognosis than those with high KIAA0101 expression in four survival curves (P-value < 0.01). These curves reflect the fact that high expression of KIAA0101 is associated negatively with the prognosis of patients with LUAD.
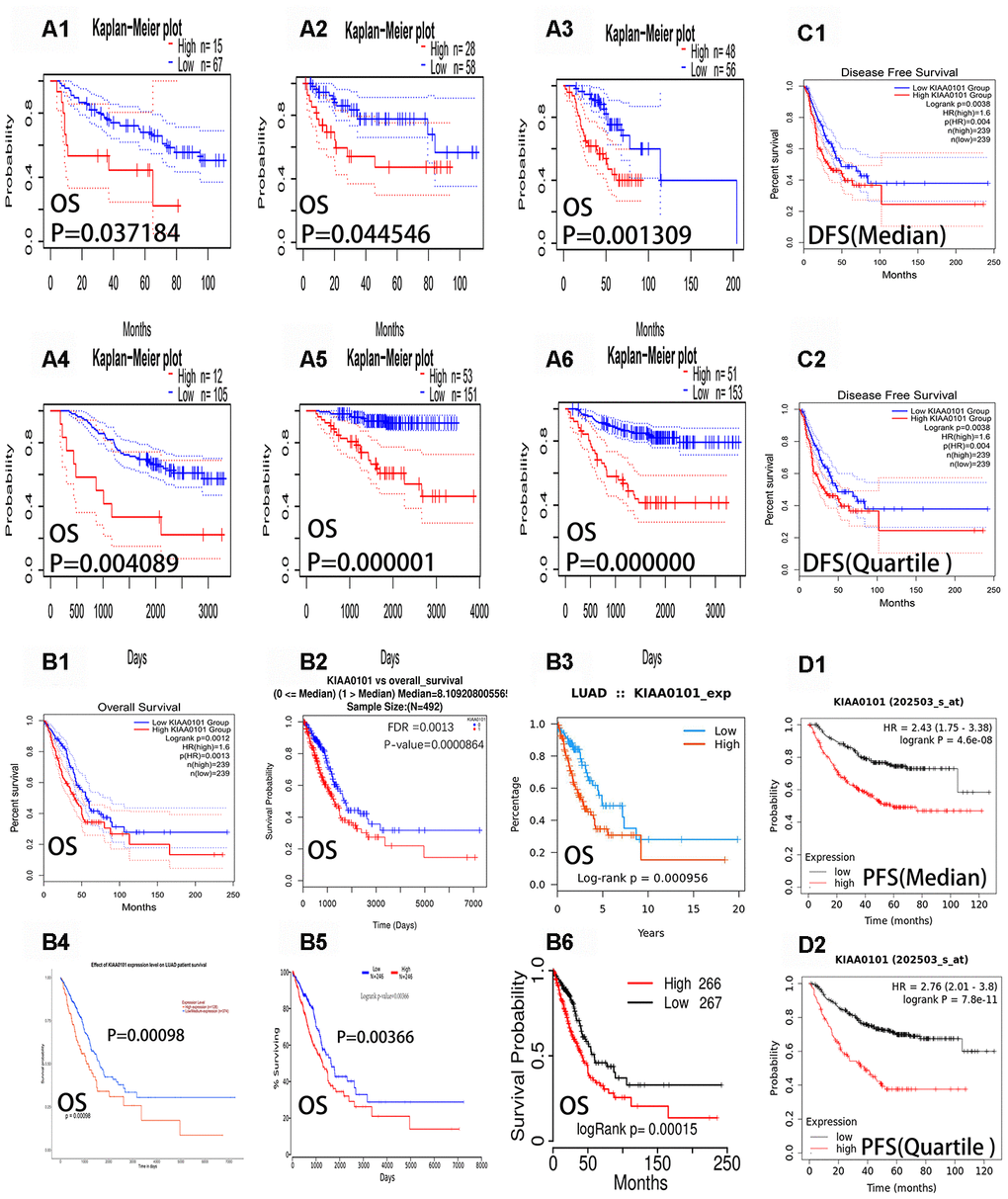
Figure 2. Overall survival curves, progression-free survival curves and disease-free survival curves of KIAA0101 in lung adenocarcinoma. The blue curves represent patients with lung adenocarcinoma with low expression of KIAA0101, and the red curves represent patients with lung adenocarcinoma with high expression of KIAA0101. (A1–A6) Six survival curves representing the six different data sets in Table 1 (from PrognoScan databases), respectively. (B1–B6) The six overall survival curves from the GEPIA, Linkedmics, Ualcan, TISIDB, Oncolnc, and TCGA portal databases, respectively. (C1–C2) Disease free survival curves (DFS) of KIAA0101 from the GEPIA database. (D1–D2) Progression free survival curves (PFS) of KIAA0101 from Kaplan Meier-plotter.
Table 1. Survival analysis of KIAA0101 mRNA in lung adenocarcinoma patients (From PrognoScan).
| DATASET | PROBE ID | ENDPOINT | Number | Ln (HR high/HR low) | COXP-VALUE | Ln (HR) | HR [95% CI-low CI-up] | |
| A1 | Jacob-00182-CANDF | 202503_s_at | Overall Survival | 82 | 1.03 | 0.037184 | 0.57 | 1.76 [1.03 - 2.99] |
| A2 | MICHIGAN-LC | D14657_at | Overall Survival | 86 | 0.96 | 0.044546 | 0.43 | 1.54 [1.01 - 2.35] |
| A3 | Jacob-00182-MSK | 202503_s_at | Overall Survival | 104 | 1.12 | 0.001309 | 0.7 | 2.01 [1.31 - 3.07] |
| A4 | GSE13213 | A_23_P117852 | Overall Survival | 117 | 1.24 | 0.004089 | 0.36 | 1.44 [1.12 - 1.84] |
| A5 | GSE31210 | 202503_s_at | Overall Survival | 204 | 1.93 | 0.000001 | 1.29 | 3.65 [2.16 - 6.17] |
| A6 | GSE31210 | 202503_s_at | Relapse Free Survival | 204 | 1.5 | 0 | 1 | 2.73 [1.90 - 3.91] |
Gene set enrichment analysis of KIAA0101 functional networks
Gene set enrichment analysis (GSEA) of KIAA0101 functional networks
(using LinkedOmics [17]). Table 2 shows the top five most significant of the seven GSEA results. All the results were statistically significant; the normalized enrichment scores (NES) were all greater than one; the P-values were all less than 0.05, and the false discovery rates (FDR) were all less than 0.05. Figure 3A shows a bar graph of the Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis. Supplementary Figure 2A–2P display the enrichment plot of pathways in the enrichment results from Figure 3A, with the top 8 and the bottom 8 according to the normalized enrichment score. We observed that the top five pathways of the KEGG [21] enrichment analysis were the cell cycle (Figure 3B), ribosome (Supplementary Figure 3), proteasome (Supplementary Figure 4), spliceosome (Supplementary Figure 5), and DNA replication (Supplementary Figure 6). The gene ontology (GO) results for the gene sets (Cellular Component) included condensed chromosome, chromosomal region, mitochondrial protein complex, ribosome, and spindle. The GO results for the gene sets (Biological Process) included chromosome segregation, DNA replication, cell cycle checkpoint, double-strand break repair, and spindle organization. The GO results for the gene sets (Molecular Function) included structural constituent of ribosome, catalytic activity, acting on DNA, single-stranded DNA binding, helicase activity and catalytic activity, and acting on RNA. Kinase targets included CDK1 (cyclin-dependent kinase 1), PLK1 (polo-like kinase 1), AURKB (aurora kinase B), CDK2 (cyclin-dependent kinase 2), and ATR (ATR serine/threonine kinase). MicroRNA targets included MIR-484, MIR-512-3P, MIR-19A, MIR-19B, MIR-219, and MIR-326. Transcription factor targets included E2F Q6, E2F Q4, E2F4DP1 01, E2F1 Q6, and E2F 02. Supplementary Tables 2–8 show the details of all Leading Edge Genes specifically contained in each gene set.
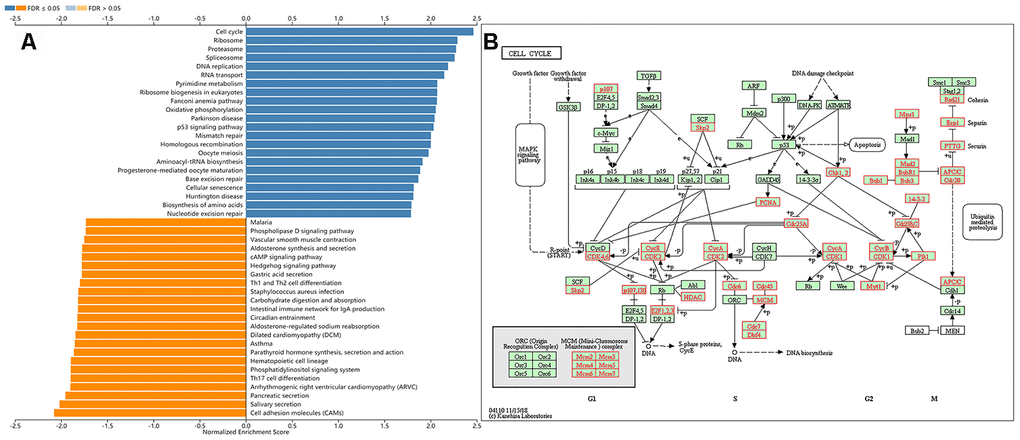
Figure 3. Gene set enrichment analysis of the KIAA0101 via the KEGG pathway. (A) Bar chart for gene set enrichment analysis of the KIAA0101 via the KEGG pathway. (B) KEGG pathway annotations of the cell cycle pathway (hsa04110). Red denotes leading edge genes; green denotes the remaining genes.
Table 2. Gene Ontology (GO) (Cellular Component), GO (Biological Process), GO (Molecular Function), Kyoto Encyclopedia of Genes and Genomes (KEGG pathway), Kinase, miRNA, and transcription factor-target networks of KIAA0101 in lung adenocarcinoma (LinkedOmics).
| Enriched Category | Geneset | LeadingEdgeNum | NES | P Value | FDR | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| GO (Cellular Component) | condensed chromosome | 66 | 2.6425 | 0 | 0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| chromosomal region | 94 | 2.5825 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| mitochondrial protein complex | 152 | 2.3976 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ribosome | 143 | 2.3858 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| spindle | 70 | 2.3837 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| GO (Biological_Process) | chromosome segregation | 97 | 2.6361 | 0 | 0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| DNA replication | 96 | 2.4830 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| cell cycle checkpoint | 60 | 2.3720 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| double-strand break repair | 61 | 2.3053 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| spindle organization | 44 | 2.2998 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| GO (Molecular_Function) | structural constituent of ribosome | 109 | 2.3582 | 0 | 0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| catalytic activity, acting on DNA | 69 | 2.2380 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| single-stranded DNA binding | 43 | 2.1594 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| helicase activity | 43 | 2.0895 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| catalytic activity, acting on RNA | 122 | 2.0460 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG pathway | Cell cycle | 48 | 2.4639 | 0 | 0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Ribosome | 100 | 2.2904 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Proteasome | 40 | 2.2761 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spliceosome | 63 | 2.2601 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| DNA replication | 27 | 2.1887 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kinase Target | Kinase_CDK1 | 74 | 2.3361 | 0 | 0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kinase_PLK1 | 32 | 2.3154 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kinase_AURKB | 35 | 2.1660 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kinase_CDK2 | 73 | 2.1657 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kinase_ATR | 20 | 2.0601 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| miRNA Target | GAGCCTG,MIR-484 | 40 | -(1.7728 | 0 | 0.025589 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAGCACT,MIR-512-3P | 53 | (1.6953 | 0 | 0.029684 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| TTTGCAC,MIR-19A,MIR-19B | 148 | (1.7059 | 0 | 0.034546 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| GACAATC,MIR-219 | 60 | (1.7191 | 0 | 0.039237 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CCCAGAG,MIR-326 | 58 | (1.6503 | 0 | 0.039481 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Transcription Factor Target | V$E2F_Q6 | 87 | 2.2583 | 0 | 0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| V$E2F_Q4 | 86 | 2.2538 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| V$E2F4DP1_01 | 94 | 2.2226 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| V$E2F1_Q6 | 96 | 2.2207 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| V$E2F_02 | 93 | 2.1958 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Abbreviations: Leading EdgeNum, the number of leading edge genes; FDR, false discovery rate from Benjamini and Hochberg from gene set enrichment analysis (GSEA); NES, Normalized Enrichment Score. It is generally considered that the absolute value of NES ≥ 1.0, NOM p-val ≤ 0.05, FDR q-val ≤ 0.25 are significant gene sets. The annotation was found in the Molecular Signatures Database (MSigDB) for transcription factors (TF). | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
The relationship between immune infiltration and the expression of KIAA0101 in lung adenocarcinoma
Figure 7A–7F show heat maps of the correlation between the expression of KIAA0101 in multiple cancers and different tumor infiltrating lymphocytes (TILs), immunoinhibitors, immunostimulators, MHC molecules, chemokines, and receptors. Figure 7G1–7G18 show scatter diagrams of tumor infiltrating lymphocytes that correlated negatively with KIAA0101 expression in LUAD. Figure 7H1–7H10 show scatter diagrams of TILs that correlated positively with KIAA0101 expression in LUAD. When the expression of KIAA0101 increased, the number of activated CD4 T cells increased. The numbers of Th1 cell, Th17 cells, and follicular helper cell were negatively related to the expression of KIAA0101, and Th2 cell numbers were positively related to the expression of KIAA0101 (P < 0.05). KIAA0101 expression correlated positively with the numbers of effector memory CD8 T cells, natural killer cells, regulatory T cells, activated B cells, immature B cells, plasmacytoid dendritic cells, macrophages, eosinophil, mast cells, and neutrophils (all P < 0.05). Supplementary Figures 9–13 show scatter plots of KIAA0101 expression related to immunoinhibitors, immunostimulators, MHC molecules, chemokines, and chemokine receptors, respectively. Most chemokine receptors correlated negatively with the expression of KIAA0101 (Supplementary Figure 13). The majority of MHC molecules correlated negatively with the expression of KIAA0101 (Supplementary Figure 11).
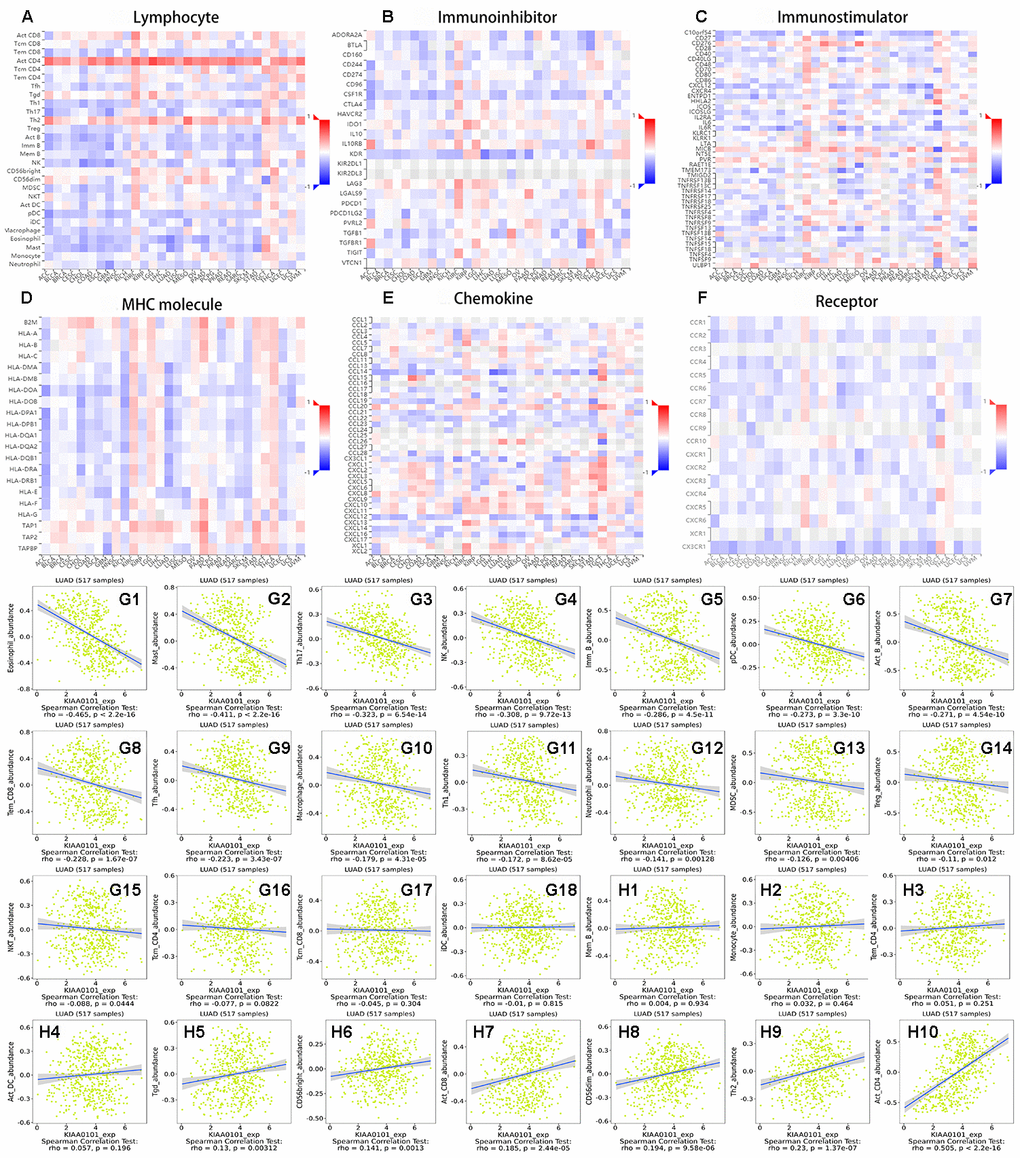
Figure 7. The relationship between immune infiltration and expression of KIAA0101 in lung adenocarcinoma. (A–F) Heat maps of KIAA0101 expression and lymphocytes, immunoinhibitors, immunostimulators, MHC molecules, chemokines, and receptors in different cancers. (G1–G18) are scatter plots of the negative correlation between KIAA0101 expression and lymphocytes in the treatment of lung adenocarcinoma. (H1–H10) are scatter plots of the positive correlation between KIAA0101 expression and lymphocytes in the treatment of lung adenocarcinoma.
We found that the metabolism of seven drugs upregulated the expression of KIAA0101 (Table 3). They included Acetaminophen [22], Estradiol [23], Genistein [23], Progesterone [24], Resveratrol [25], Rifampicin [26], and Valproic acid [27]. The metabolism of 24 drugs downregulated the expression of KIAA0101. They included amoxicifene, estradiol, yolk, copper, and cyclosporine (Table 3). The metabolism of Estradiol [23, 28], Genistein [23, 29], and Progesterone [24, 30] could both upregulate and downregulate the expression of KIAA0101 (Table 3).
Table 3. The Pharmaco-transcriptomics of KIAA0101 (Up or downregulation of genes caused by the metabolism of pharmaceutical compounds).
| DRUG | CHANGE | INTERACTION | REFERENCES (PMID) |
| Acetaminophen | upregulated | Acetaminophen results in increased expression of KIAA0101 mRNA | 25704631 |
| Afimoxifene | downregulated | Afimoxifene results in decreased expression of KIAA0101 mRNA | 16514628 |
| Estradiol | downregulated | Estradiol results in decreased expression of KIAA0101 mRNA | 23019147 |
| Estradiol | upregulated | Estradiol results in increased expression of KIAA0101 mRNA | 16474171 |
| Custirsen | downregulated | CC-8490 results in decreased expression of KIAA0101 mRNA | 15604281 |
| Copper | downregulated | Binding with copper resulted in decreased expression of KIAA0101 mRNA | 20971185 |
| Cyclosporine | downregulated | Cyclosporine results in decreased expression of KIAA0101 mRNA | 20106945 |
| Dasatinib | downregulated | Dasatinib results in decreased expression of KIAA0101 mRNA | 20579391 |
| Dronabinol | downregulated | Dronabinol results in decreased expression of KIAA0101 mRNA | 18454173 |
| Calcitriol | downregulated | Calcitriol results in decreased expression of KIAA0101 mRNA | 21592394 |
| Fluorouracil | downregulated | Fluorouracil results in decreased expression of KIAA0101 mRNA | 17039268 |
| Fulvestrant | downregulated | fulvestrant results in decreased expression of KIAA0101 mRNA | 16514628 |
| Genistein | downregulated | Genistein results in decreased expression of KIAA0101 mRNA | 19371625 |
| Genistein | upregulated | Genistein results in increased expression of KIAA0101 mRNA | 16474171 |
| Lucanthone | downregulated | Lucanthone results in decreased expression of KIAA0101 mRNA | 21148553 |
| Metamfetamine | downregulated | Methamphetamine results in decreased expression of KIAA0101 mRNA | 25290377 |
| Methotrexate | downregulated | Methotrexate results in decreased expression of KIAA0101 mRNA | 24657277 |
| Palbociclib | downregulated | Palbociclib results in decreased expression of KIAA0101 mRNA | 22869556 |
| Phenethyl Isothiocyanate | downregulated | Phenethyl isothiocyanate results in decreased expression of KIAA0101 mRNA | 26678675 |
| Piroxicam | downregulated | Piroxicam results in decreased expression of KIAA0101 mRNA | 21858171 |
| Progesterone | downregulated | Progesterone results in decreased expression of KIAA0101 mRNA | 21795739 |
| Progesterone | upregulated | Progesterone results in increased expression of KIAA0101 mRNA | 18070364 |
| Raloxifene | downregulated | Raloxifene Hydrochloride results in decreased expression of KIAA0101 mRNA | 16514628 |
| Resveratrol | upregulated | Resveratrol results in increased expression of KIAA0101 mRNA | 19167446 |
| Rifampicin | upregulated | Rifampin results in increased expression of KIAA0101 mRNA | 24552687 |
| Silicon dioxide | downregulated | Silicon Dioxide analog results in decreased expression of KIAA0101 mRNA | 25895662 |
| Tamoxifen | downregulated | Tamoxifen results in decreased expression of KIAA0101 mRNA | 15604281 |
| Testosterone | downregulated | Testosterone results in decreased expression of KIAA0101 mRNA | 21592394 |
| Valproic acid | upregulated | Valproic Acid results in increased expression of KIAA0101 mRNA | 23179753 |
| Vitamin E | downregulated | Vitamin E results in decreased expression of KIAA0101 mRNA | 19244175 |
| Zoledronic acid | downregulated | Zoledronic acid results in decreased expression of KIAA0101 mRNA | 24714768 |
Discussion and conclusions
Early detection of LUAD is a challenge for clinicians. Low dose computed tomography (LDCT) has the advantages of simplicity and high sensitivity. Compared with the traditional chest film and tumor markers, the false negative rate of LDCT was reduced [31]. However, more accurate and timely biomarkers are needed to support a diagnosis of LUAD. It has been reported that KIAA0101 is differentially expressed in ovarian cancer [32], adrenal cancer [33], renal cell carcinoma [10], hepatocellular carcinoma [10], and breast cancer [12]. However, there are few studies about KIAA0101 in LUAD [34]. The results of the present study showed that KIAA0101 was significantly differentially expressed in all lung adenocarcinomas (Figure 1). This is very interesting; however, it requires verification in further comparative studies.
We further explored the mechanism by which KIAA0101 can act as a marker of LUAD. Hosokawa et al. [35] found that if KIAA0101 was knocked down using small interfering RNA (siRNA), the DNA replication rate was greatly reduced in pancreatic cancer cell lines. The exogenous overexpression of KIAA0101 enhanced the growth of cancer cells, suggesting the carcinogenicity of KIAA0101. They also found that inhibition of the KIAA0101-PCNA interaction significantly inhibited the growth of cancer cells. Lv et al. [11] found that knockdown of KIAA0101 could promote the formation of p53 / SP1 complexes in breast cancer, and inhibited the proliferation and cell cycle process of breast cancer cells. Many studies have shown that KIAA0101 is involved in the regulation of DNA replication and repair, cell cycle progression, and cell proliferation, and can reduce the apoptosis induced by UV [35–37]. Our results are consistent with these previous reports. KEGG enrichment analysis of KIAA0101 identified the cell cycle, ribosome, proteasome, spliceosome, and DNA replication pathways as enriched. In LUAD, KIAA0101 was related to kinase networks including CDK1, PLK1, AURKB, CDK2, and ATR. These kinases regulate cell cycle stability, chromosome separation, and the cell cycle during mitosis and meiosis [38, 39]. In fact, ATR is one of the key kinase regulators for genomic stability; it directly phosphorylates a variety of important substrates, including p53 protein and cyclin [40]. ATR kinase inhibitors can reduce the growth of tumor cells [41]. In addition, differentially expressed genes highly related to KIAA0101 in lung adenocarcinoma were associated with the cell cycle, DNA replication and modification, mitosis, and meiosis. Thus, KIAA0101 might regulate cell cycle progression and DNA replication and repair via the above kinases in LUAD [42]. Cell cycle progression and DNA replication occur mainly in the nucleus, which might also be the main reason for the markedly increased expression of KIAA0101 in the nucleus and slightly enhanced expression in the cytoplasm.
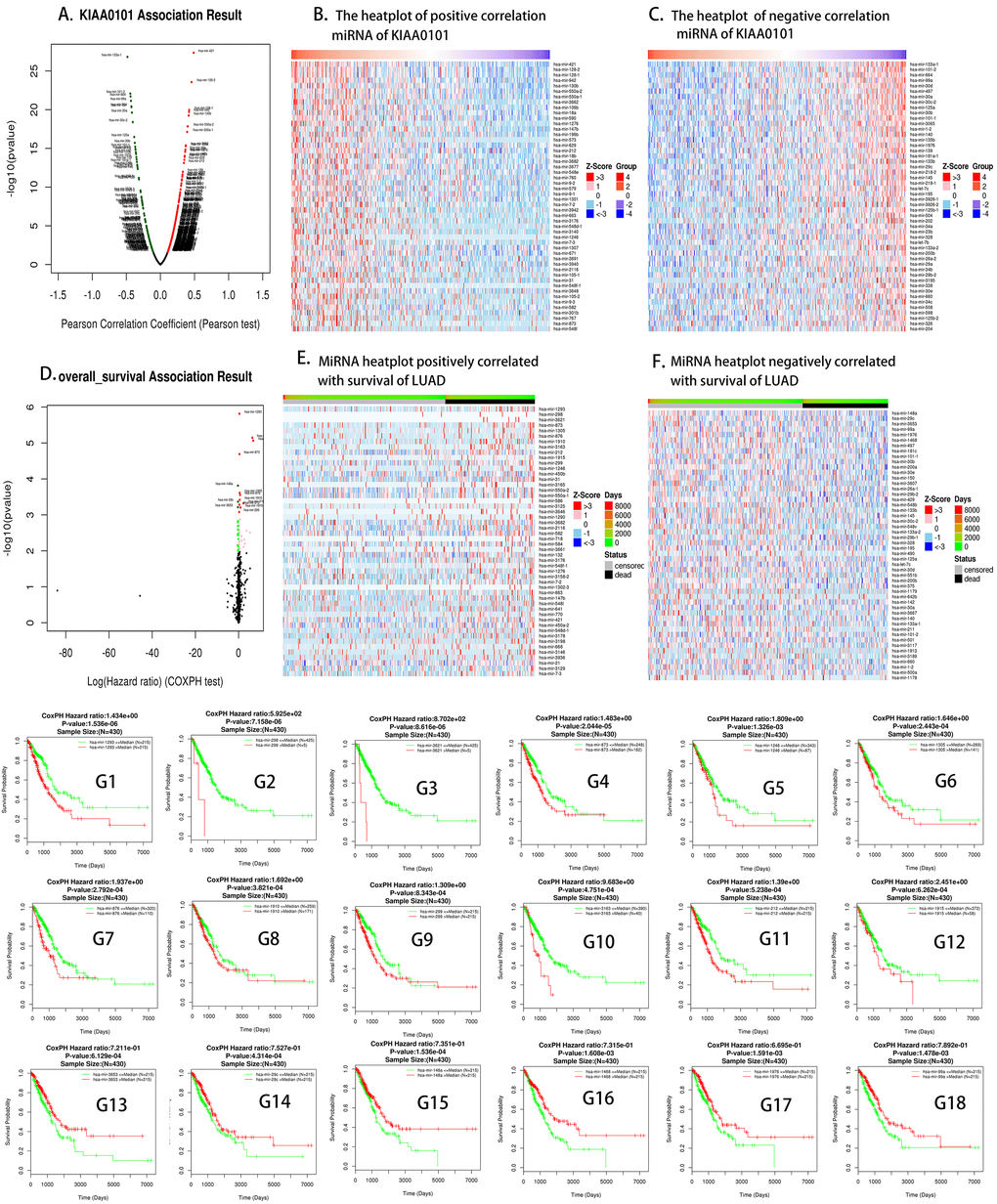
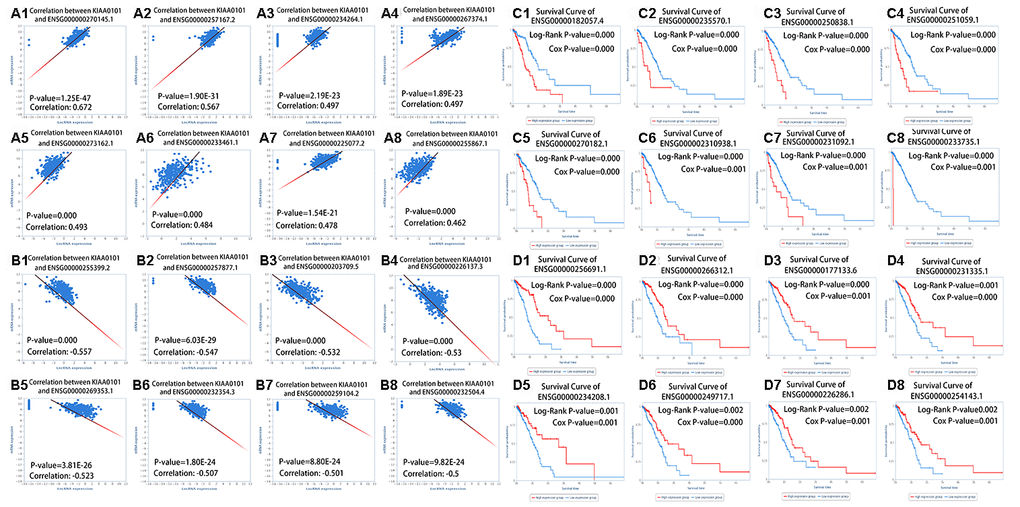
Based on the above results (Figure 2), we found that the prognosis of patients with LUAD could be predicted by the expression of KIAA0101. We found that some drugs [43] could reduce the expression of KIAA0101 (Table 3). The decrease in gene expression by these drugs [22] might be transient or concentration-dependent, and we will further evaluate the effect of different concentrations and times on KIAA0101 expression in the future. Some drugs that upregulated the expression of KIAA0101 might be potential carcinogens of LUAD. We also identified miRNAs and lncRNAs that are related to the expression of KIAA0101 (Figures 5 and 6). The expression of these miRNAs and lncRNAs was related to the survival of patients with lung adenocarcinoma. Thus, these miRNAs and lncRNAs could also be used as potential biomarkers of lung adenocarcinoma [17, 44]. These miRNAs and lncRNAs could be further studied to improve the epigenetics of KIAA0101 in lung adenocarcinoma.
The interaction between tumor cells and their surrounding matrix not only affects the occurrence and development of the disease, but also is closely related to the prognosis of patients [45]. Evidence suggests that TILs infiltrate tumor tissue and play a role in the disease by regulating anti-tumor immunity [46]. With the increase in KIAA0101 expression, the numbers of Th1 cells, Th17 cells, and follicular helper cells decreased, and the number of Th2 cells increased. Type 1 T helper cells are positive immunoregulatory cells, and type 2 T helper cells are negative immunoregulatory cells. Thus, variation in KIAA0101 expression leads to a shift of the Th1/Th2 ratio. The immunosuppressive state will seriously affect the anti-tumor immunity of the body, and eventually lead to the occurrence and development of tumors [47]. With the increase in KIAA0101 expression, the number of effector memory CD8 T cells, natural killer cells, regulatory T cells, activated B cells, immature B cells, plasmacytoid dendritic cells, macrophages, eosinophil, mast cells, and neutrophils will decrease. This finding confirms the view that increased expression of KIAA0101 reduces survival in patients with LUAD. Chemokine receptors are seven transmembrane G protein-coupled receptors. Their main function is to receive chemokine signals and further conduct signals, which plays an important positive role in promoting tumor peripheral immune infiltration [48]. The main function of MHC molecules is to participate in antigen presentation. MHC molecules play an important role in the differentiation and maturation of T cells, and play an important positive role in tumor peripheral immune infiltration [49]. When the expression of KIAA0101 increased, the levels of most chemokine receptors and MHC molecules in patients decreased (Supplementary Figures 11, 12), which would further reduce the tumor peripheral immune function. This finding again confirmed that an increase of KIAA0101 expression reduces the survival rate of patients with LUAD. Therefore, our research will contribute to the improvement of immunotherapy of LUAD and the development of new immunotherapy targets.
In conclusion, we speculated that KIAA0101 is a new LUAD marker. The prognosis of patients with LUAD with high expression of KIAA0101 was poor. KIAA0101 is related to the cell cycle, ribosome, protocol, splicing, and DNA replication. The results showing the immune cell associations of KIAA0101 could lead to new methods of immunotherapy of LUAD. The study of drug transcription and metabolism associated with KIAA0101 might provide alternative drugs for the targeted treatment of LUAD. This study will provide new directions for the diagnosis, prognostic evaluation, immunotherapy, and targeted therapy of LUAD.
We must acknowledge the potential limitations of our analysis. Further experimental verification is needed. In addition, our study is limited to lung adenocarcinoma, and there was no analysis in other types of lung cancer. We hope that our next study will assess the association of KIAA0101 with other types of lung cancer besides adenocarcinoma, to further compare the specific role of KIAA0101 expression in the diagnosis and treatment of different types of lung cancer.
Materials and Methods
Oncomine, GEPIA, and Ualcan analysis
All genes in the study of Hou Lung were analyzed and compared using Oncomine [50]. Overexpression of the genes in lung adenocarcinoma was compared with that in normal control tissue (log2 median-centered intensity). We used Oncomine to analyze the expression of KIAA0101 in different tumor studies. In the Oncomine dataset, the naming convention lists the primary author's last name first, followed by the tissue type used in the study. We then used GEPIA [16] to analyze the expression of KIAA0101 in a variety of tumors compared with that in normal tissue (Figure 1C). KIAA0101 transcription in subgroups of patients with lung adenocarcinoma, stratified based on sex, age, and other criteria, were analyzed using the Ualcan database [14]. We also analyzed the differential expression of KIAA0101 in LUAD tissues and normal lung tissues using Ualcan.
PrognoScan, OncoLnc, and TCGA portal analysis
The prognosis of patients with LUAD with different expression levels of KIAA0101 was evaluated using PrognoScan [15]. PrognoScan is a new database for the meta-analysis of the prognostic value of genes. We used GEPIA [16], LinkedOmics [17], Ualcan [14], TISIDB [18], OncoLnc [19], and TCGA portals [20] to construct the OS curves for patients with LUAD. We used GEPIA [16] to construct the DFS curves for patients with LUAD. We used Kaplan Meier-plotter to construct the PFS curves for patients with LUAD. The survival curves between patients with low expression of KIAA0101 and patients with high expression of KIAA0101 in lung adenocarcinoma were compared.
LinkedOmics and TANRIC analysis
We conducted Gene Set Enrichment Analysis (GSEA) of KEGG pathway data [51], gene ontology cellular component, gene ontology biological process, gene ontology molecular function, a kinase target network, an miRNA target network, and a transcription factor target network of KIAA0101. We performed an miRNA Pearson correlation test and COXPH test of KIAA0101 using Linkedomics [17]. We found the differentially expressed genes related to KIAA0101 in LUAD. We analyzed lncRNAs associated with KIAA0101 using TANRIC [44] and assessed whether the level of lncRNA expression associated with KIAA0101 had different impacts on the survival of patients.
TISIDB analysis and DrugBank analysis
We used TISIDB [18] to analyze the relationship between the expression of KIAA0101 and immune infiltration in LUAD. Transcriptome and clinical data of 30 cancer types from the Cancer Genome Atlas (TCGA) were assessed [18]. High throughput screening data were then used to detect the resistance and sensitivity of tumor cells to T-cell-mediated killing. We used Drugbank to analyze the pharmacotranscription of KIAA0101 [52]. The DrugBank database is a unique bioinformatics and cheminformatics resource that combines detailed drug data with comprehensive drug target information.
Author Contributions
Sheng Hu and Yiping Wei designed and performed the research; analyzed and interpreted the data; and drafted the manuscript. Weibiao Zeng, Wenxiong Zhang, Jianjun Xu, Dongliang Yu and Jinhua Peng conceived the study and participated in the research design and data interpretation. All authors read and approved the final manuscript.
Acknowledgments
We hereby thank the Department of Thoracic Surgery of the Second Affiliated Hospital of Nanchang University for their strong support for this study.
Conflicts of Interest
All the data in this paper are public, and the quoted data have indicated the source. There is no conflicts of interest in this paper.
Funding
This study was supported by grants from the National Natural Science Foundation of China (81560345 and 81860379), the Preeminence Youth Fund of Jiangxi Province (20162BCB23058), the China Postdoctoral Science Foundation Grant (2017M610401), and the Science and Technology Planning Project at the Department of Science and Technology of Jiangxi Province, China (20151BBG70165 and 20171BAB205075).
References
- 1. Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, Jemal A, Yu XQ, He J. Cancer statistics in China, 2015. CA Cancer J Clin. 2016; 66:115–32. https://doi.org/10.3322/caac.21338 [PubMed]
- 2. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2019. CA Cancer J Clin. 2019; 69:7–34. https://doi.org/10.3322/caac.21551 [PubMed]
- 3. Govindan R, Page N, Morgensztern D, Read W, Tierney R, Vlahiotis A, Spitznagel EL, Piccirillo J. Changing epidemiology of small-cell lung cancer in the United States over the last 30 years: analysis of the surveillance, epidemiologic, and end results database. J Clin Oncol. 2006; 24:4539–44. https://doi.org/10.1200/JCO.2005.04.4859 [PubMed]
- 4. Ruiz-Cordero R, Devine WP. Targeted therapy and checkpoint immunotherapy in lung cancer. Surg Pathol Clin. 2020; 13:17–33. https://doi.org/10.1016/j.path.2019.11.002 [PubMed]
- 5. Pabla S, Conroy JM, Nesline MK, Glenn ST, Papanicolau-Sengos A, Burgher B, Hagen J, Giamo V, Andreas J, Lenzo FL, Yirong W, Dy GK, Yau E, et al. Proliferative potential and resistance to immune checkpoint blockade in lung cancer patients. J Immunother Cancer. 2019; 7:27. https://doi.org/10.1186/s40425-019-0506-3 [PubMed]
- 6. Sun CC, Li SJ, Yuan ZP, Li DJ. MicroRNA-346 facilitates cell growth and metastasis, and suppresses cell apoptosis in human non-small cell lung cancer by regulation of XPC/ERK/Snail/E-cadherin pathway. Aging (Albany NY). 2016; 8:2509–24. https://doi.org/10.18632/aging.101080 [PubMed]
- 7. Moldovan GL, Pfander B, Jentsch S. PCNA, the maestro of the replication fork. Cell. 2007; 129:665–79. https://doi.org/10.1016/j.cell.2007.05.003 [PubMed]
- 8. Halling-Brown MD, Bulusu KC, Patel M, Tym JE, Al-Lazikani B. canSAR: an integrated cancer public translational research and drug discovery resource. Nucleic Acids Res. 2012; 40:D947–56. https://doi.org/10.1093/nar/gkr881 [PubMed]
- 9. Thul PJ, Åkesson L, Wiking M, Mahdessian D, Geladaki A, Ait Blal H, Alm T, Asplund A, Björk L, Breckels LM, Bäckström A, Danielsson F, Fagerberg L, et al. A subcellular map of the human proteome. Science. 2017; 356:eaal3321. https://doi.org/10.1126/science.aal3321 [PubMed]
- 10. Zhang T, Guo J, Gu J, Chen K, Wang Z, Li H, Wang G, Wang J. KIAA0101 is a novel transcriptional target of FoxM1 and is involved in the regulation of hepatocellular carcinoma microvascular invasion by regulating epithelial-mesenchymal transition. J Cancer. 2019; 10:3501–16. https://doi.org/10.7150/jca.29490 [PubMed]
- 11. Lv W, Su B, Li Y, Geng C, Chen N. KIAA0101 inhibition suppresses cell proliferation and cell cycle progression by promoting the interaction between p53 and Sp1 in breast cancer. Biochem Biophys Res Commun. 2018; 503:600–06. https://doi.org/10.1016/j.bbrc.2018.06.046 [PubMed]
- 12. Kais Z, Barsky SH, Mathsyaraja H, Zha A, Ransburgh DJ, He G, Pilarski RT, Shapiro CL, Huang K, Parvin JD. KIAA0101 interacts with BRCA1 and regulates centrosome number. Mol Cancer Res. 2011; 9:1091–99. https://doi.org/10.1158/1541-7786.MCR-10-0503 [PubMed]
- 13. Zhu K, Diao D, Dang C, Shi L, Wang J, Yan R, Yuan D, Li K. Elevated KIAA0101 expression is a marker of recurrence in human gastric cancer. Cancer Sci. 2013; 104:353–59. https://doi.org/10.1111/cas.12083 [PubMed]
- 14. Chandrashekar DS, Bashel B, Balasubramanya SA, Creighton CJ, Ponce-Rodriguez I, Chakravarthi BV, Varambally S. UALCAN: a portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia. 2017; 19:649–58. https://doi.org/10.1016/j.neo.2017.05.002 [PubMed]
- 15. Mizuno H, Kitada K, Nakai K, Sarai A. PrognoScan: a new database for meta-analysis of the prognostic value of genes. BMC Med Genomics. 2009; 2:18. https://doi.org/10.1186/1755-8794-2-18 [PubMed]
- 16. Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 2017; 45:W98–102. https://doi.org/10.1093/nar/gkx247 [PubMed]
- 17. Vasaikar SV, Straub P, Wang J, Zhang B. LinkedOmics: analyzing multi-omics data within and across 32 cancer types. Nucleic Acids Res. 2018; 46:D956–63. https://doi.org/10.1093/nar/gkx1090 [PubMed]
- 18. Ru B, Wong CN, Tong Y, Zhong JY, Zhong SS, Wu WC, Chu KC, Wong CY, Lau CY, Chen I, Chan NW, Zhang J. TISIDB: an integrated repository portal for tumor-immune system interactions. Bioinformatics. 2019; 35:4200–02. https://doi.org/10.1093/bioinformatics/btz210 [PubMed]
- 19. Anaya J. Guide to using OncoLnc, a new TCGA data portal. Figshare. 2016. https://doi.org/https://doi.org/10.6084/m9.figshare.2991640.v1
- 20. Xu S, Feng Y, Zhao S. Proteins with evolutionarily hypervariable domains are associated with immune response and better survival of basal-like breast cancer patients. Comput Struct Biotechnol J. 2019; 17:430–40. https://doi.org/10.1016/j.csbj.2019.03.008 [PubMed]
- 21. KEGG. Curr Opin Plant Biol. 2000; 3:177. https://doi.org/10.1016/S1369-5266(00)80051-5
- 22. Jiang J, Briedé JJ, Jennen DG, Van Summeren A, Saritas-Brauers K, Schaart G, Kleinjans JC, de Kok TM. Increased mitochondrial ROS formation by acetaminophen in human hepatic cells is associated with gene expression changes suggesting disruption of the mitochondrial electron transport chain. Toxicol Lett. 2015; 234:139–50. https://doi.org/10.1016/j.toxlet.2015.02.012 [PubMed]
- 23. Buterin T, Koch C, Naegeli H. Convergent transcriptional profiles induced by endogenous estrogen and distinct xenoestrogens in breast cancer cells. Carcinogenesis. 2006; 27:1567–78. https://doi.org/10.1093/carcin/bgi339 [PubMed]
- 24. Wilcox CB, Feddes GO, Willett-Brozick JE, Hsu LC, DeLoia JA, Baysal BE. Coordinate up-regulation of TMEM97 and cholesterol biosynthesis genes in normal ovarian surface epithelial cells treated with progesterone: implications for pathogenesis of ovarian cancer. BMC Cancer. 2007; 7:223. https://doi.org/10.1186/1471-2407-7-223 [PubMed]
- 25. Dip R, Lenz S, Gmuender H, Naegeli H. Pleiotropic combinatorial transcriptomes of human breast cancer cells exposed to mixtures of dietary phytoestrogens. Food Chem Toxicol. 2009; 47:787–95. https://doi.org/10.1016/j.fct.2009.01.008 [PubMed]
- 26. Takahashi K, Tatsumi N, Fukami T, Yokoi T, Nakajima M. Integrated analysis of rifampicin-induced microRNA and gene expression changes in human hepatocytes. Drug Metab Pharmacokinet. 2014; 29:333–40. https://doi.org/10.2133/dmpk.dmpk-13-rg-114 [PubMed]
- 27. Krug AK, Kolde R, Gaspar JA, Rempel E, Balmer NV, Meganathan K, Vojnits K, Baquié M, Waldmann T, Ensenat-Waser R, Jagtap S, Evans RM, Julien S, et al. Human embryonic stem cell-derived test systems for developmental neurotoxicity: a transcriptomics approach. Arch Toxicol. 2013; 87:123–43. https://doi.org/10.1007/s00204-012-0967-3 [PubMed]
- 28. Gertz J, Reddy TE, Varley KE, Garabedian MJ, Myers RM. Genistein and bisphenol a exposure cause estrogen receptor 1 to bind thousands of sites in a cell type-specific manner. Genome Res. 2012; 22:2153–62. https://doi.org/10.1101/gr.135681.111 [PubMed]
- 29. Boehme K, Simon S, Mueller SO. Gene expression profiling in ishikawa cells: a fingerprint for estrogen active compounds. Toxicol Appl Pharmacol. 2009; 236:85–96. https://doi.org/10.1016/j.taap.2009.01.006 [PubMed]
- 30. Soloff MS, Jeng YJ, Izban MG, Sinha M, Luxon BA, Stamnes SJ, England SK. Effects of progesterone treatment on expression of genes involved in uterine quiescence. Reprod Sci. 2011; 18:781–97. https://doi.org/10.1177/1933719111398150 [PubMed]
- 31. Seijo LM, Peled N, Ajona D, Boeri M, Field JK, Sozzi G, Pio R, Zulueta JJ, Spira A, Massion PP, Mazzone PJ, Montuenga LM. Biomarkers in lung cancer screening: achievements, promises, and challenges. J Thorac Oncol. 2019; 14:343–57. https://doi.org/10.1016/j.jtho.2018.11.023 [PubMed]
- 32. Chen H, Xia B, Liu T, Lin M, Lou G. KIAA0101, a target gene of miR-429, enhances migration and chemoresistance of epithelial ovarian cancer cells. Cancer Cell Int. 2016; 16:74. https://doi.org/10.1186/s12935-016-0353-y [PubMed]
- 33. Jain M, Zhang L, Patterson EE, Kebebew E. KIAA0101 is overexpressed, and promotes growth and invasion in adrenal cancer. PLoS One. 2011; 6:e26866. https://doi.org/10.1371/journal.pone.0026866 [PubMed]
- 34. Kato T, Daigo Y, Aragaki M, Ishikawa K, Sato M, Kaji M. Overexpression of KIAA0101 predicts poor prognosis in primary lung cancer patients. Lung Cancer. 2012; 75:110–18. https://doi.org/10.1016/j.lungcan.2011.05.024 [PubMed]
- 35. Emanuele MJ, Ciccia A, Elia AE, Elledge SJ. Proliferating cell nuclear antigen (PCNA)-associated KIAA0101/PAF15 protein is a cell cycle-regulated anaphase-promoting complex/cyclosome substrate. Proc Natl Acad Sci USA. 2011; 108:9845–50. https://doi.org/10.1073/pnas.1106136108 [PubMed]
- 36. Simpson F, Lammerts van Bueren K, Butterfield N, Bennetts JS, Bowles J, Adolphe C, Simms LA, Young J, Walsh MD, Leggett B, Fowles LF, Wicking C. The PCNA-associated factor KIAA0101/p15(PAF) binds the potential tumor suppressor product p33ING1b. Exp Cell Res. 2006; 312:73–85. https://doi.org/10.1016/j.yexcr.2005.09.020 [PubMed]
- 37. van Bueren KL, Bennetts JS, Fowles LF, Berkman JL, Simpson F, Wicking C. Murine embryonic expression of the gene for the UV-responsive protein p15(PAF). Gene Expr Patterns. 2007; 7:47–50. https://doi.org/10.1016/j.modgep.2006.05.006 [PubMed]
- 38. Fang H, Niu K, Mo D, Zhu Y, Tan Q, Wei D, Li Y, Chen Z, Yang S, Balajee AS, Zhao Y. RecQL4-aurora B kinase axis is essential for cellular proliferation, cell cycle progression, and mitotic integrity. Oncogenesis. 2018; 7:68. https://doi.org/10.1038/s41389-018-0080-4 [PubMed]
- 39. Herrera MC, Chymkowitch P, Robertson JM, Eriksson J, Bøe SO, Alseth I, Enserink JM. Corrigendum: Cdk1 gates cell cycle-dependent tRNA synthesis by regulating RNA polymerase III activity. Nucleic Acids Res. 2018; 46:12188–89. https://doi.org/10.1093/nar/gky1102 [PubMed]
- 40. Cortez D, Guntuku S, Qin J, Elledge SJ. ATR and ATRIP: partners in checkpoint signaling. Science. 2001; 294:1713–16. https://doi.org/10.1126/science.1065521 [PubMed]
- 41. Hafsi H, Dillon MT, Barker HE, Kyula JN, Schick U, Paget JT, Smith HG, Pedersen M, McLaughlin M, Harrington KJ. Combined ATR and DNA-PK inhibition radiosensitizes tumor cells independently of their p53 status. Front Oncol. 2018; 8:245. https://doi.org/10.3389/fonc.2018.00245 [PubMed]
- 42. Lin Y, Liang R, Qiu Y, Lv Y, Zhang J, Qin G, Yuan C, Liu Z, Li Y, Zou D, Mao Y. Expression and gene regulation network of RBM8A in hepatocellular carcinoma based on data mining. Aging (Albany NY). 2019; 11:423–47. https://doi.org/10.18632/aging.101749 [PubMed]
- 43. Scafoglio C, Ambrosino C, Cicatiello L, Altucci L, Ardovino M, Bontempo P, Medici N, Molinari AM, Nebbioso A, Facchiano A, Calogero RA, Elkon R, Menini N, et al. Comparative gene expression profiling reveals partially overlapping but distinct genomic actions of different antiestrogens in human breast cancer cells. J Cell Biochem. 2006; 98:1163–84. https://doi.org/10.1002/jcb.20820 [PubMed]
- 44. Li J, Han L, Roebuck P, Diao L, Liu L, Yuan Y, Weinstein JN, Liang H. TANRIC: an interactive open platform to explore the function of lncRNAs in cancer. Cancer Res. 2015; 75:3728–37. https://doi.org/10.1158/0008-5472.CAN-15-0273 [PubMed]
- 45. Joyce JA, Pollard JW. Microenvironmental regulation of metastasis. Nat Rev Cancer. 2009; 9:239–52. https://doi.org/10.1038/nrc2618 [PubMed]
- 46. Sasada T, Suekane S. Variation of tumor-infiltrating lymphocytes in human cancers: controversy on clinical significance. Immunotherapy. 2011; 3:1235–51. https://doi.org/10.2217/imt.11.106 [PubMed]
- 47. Lauerova L, Dusek L, Simickova M, Kocák I, Vagundová M, Zaloudík J, Kovarík J. Malignant melanoma associates with Th1/Th2 imbalance that coincides with disease progression and immunotherapy response. Neoplasma. 2002; 49:159–66. [PubMed]
- 48. Liu J, Li Y, Zhu X, Li Q, Liang X, Xie J, Hu S, Peng W, Li C. Increased CX3CL1 mRNA expression level is a positive prognostic factor in patients with lung adenocarcinoma. Oncol Lett. 2019; 17:4877–90. https://doi.org/10.3892/ol.2019.10211 [PubMed]
- 49. Holling TM, Schooten E, van Den Elsen PJ. Function and regulation of MHC class II molecules in T-lymphocytes: of mice and men. Hum Immunol. 2004; 65:282–90. https://doi.org/10.1016/j.humimm.2004.01.005 [PubMed]
- 50. Rhodes DR, Kalyana-Sundaram S, Mahavisno V, Varambally R, Yu J, Briggs BB, Barrette TR, Anstet MJ, Kincead-Beal C, Kulkarni P, Varambally S, Ghosh D, Chinnaiyan AM. Oncomine 3.0: genes, pathways, and networks in a collection of 18,000 cancer gene expression profiles. Neoplasia. 2007; 9:166–80. https://doi.org/10.1593/neo.07112 [PubMed]
- 51. Kanehisa M, Sato Y. KEGG mapper for inferring cellular functions from protein sequences. Protein Sci. 2020; 29:28–35. https://doi.org/10.1002/pro.3711 [PubMed]
- 52. Wishart DS, Feunang YD, Guo AC, Lo EJ, Marcu A, Grant JR, Sajed T, Johnson D, Li C, Sayeeda Z, Assempour N, Iynkkaran I, Liu Y, et al. DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Res. 2018; 46:D1074–82. https://doi.org/10.1093/nar/gkx1037 [PubMed]