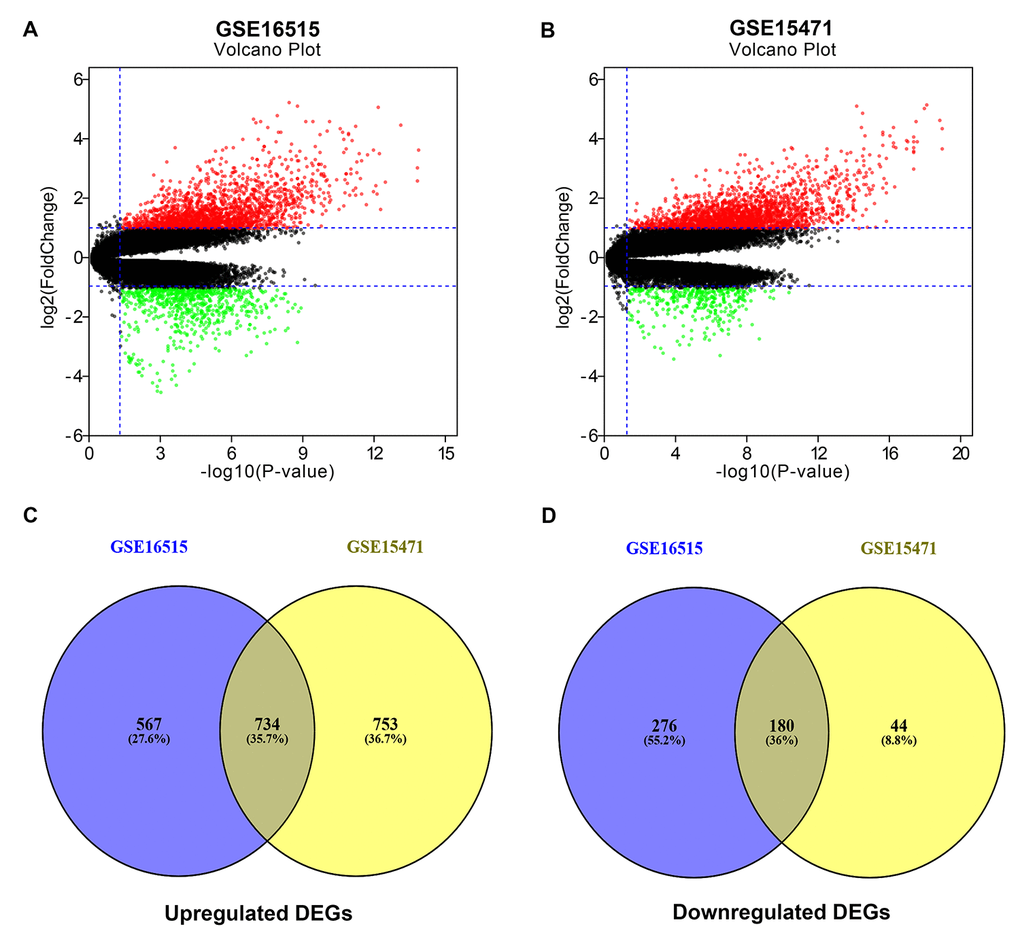
Figure 1.Identification of significant differentially expressed genes (DEGs) in pancreatic cancer. (A) Volcano plot showing the DEGs identified from GSE16515. (B) Volcano plot showing the DEGs identified from GSE15471. X axis represents log transformed P value, and Y axis indicates the mean expression differences of genes between pancreatic cancer samples and normal samples. Note: The two volcano plots showed all of the DEGs; the black dots represent genes that are not differentially expressed between pancreatic cancer samples and normal samples, and the green dots and red dots represent the downregulated and upregulated genes in pancreatic cancer samples, respectively. |log2FC| >1 and adj. p-value < 0.05 were set as the cut-off criteria. (C) The intersection of upregulated DEGs of GSE16515 and GSE15471 datasets. (D) The intersection of downregulated DEGs of GSE16515 and GSE15471 datasets. The intersected DEGs were defined as the significant DEGs.