Introduction
Cholangiocarcinoma (CCA) is a digestive carcinoma with a high mortality rate across the world [1,2]. It is originated from the neoplastic transformation of biliary epithelial cells with high malignancy. Based on public data, the 5-year overall survival of this devastating disease is less than 40% [3]. Due to the limit approaches for CCA early detection, many patients lost their chance for radical surgery. Although great advancements in surgical technique and neoadjuvant chemoradiotherapy have been achieved, the prognosis of CCA patients remains unfavorable [4]. Therefore, it is urgent to identify the underlying mechanisms in regulating CCA initiation and progression.
Studies have documented that more than 90% of the human genome is transcribed into non-coding RNAs (ncRNAs), indicating that ncRNAs might play crucial roles in cellular physiology and pathology processes [5,6]. Circular RNAs (circRNAs) is a class of ncRNAs which has limited protein coding potential and formed by a circular structure [7,8]. Accumulating evidence documented that circRNAs are closely involved in carcinogenesis and cancer progression [9,10]. Mechanistically, the studies demonstrated that circRNAs could sponge miRNAs or function as molecular scaffolds to guide RNA binding proteins (RBPs) [11]. For example, a recent study indicated that decreased circRNA_100269 in gastric cancer could suppress cancer cell growth by targeting miR-630 [12]. Nevertheless, the study relevant to circRNAs and CCA is still on the way. Previously, we identified circ_0001649 was downregulated in CCA and ectopic expression of circ_0001649 exerts tumor suppressive role in cell progression [13]. CircRNAs exploration might provide the researches a novel insight into the pathogenesis of this fatal disease.
Previously, we identified circ_0005230 was elevated in breast cancer tissue samples and was linked to breast cancer patients’ unfavorable prognosis. Furthermore, the biological functions of circ_0005230 was partially attributed to its modulation on miR-618/CBX8 signaling [14]. Circ_0005230 is located at chr1:172109619-172113577 and is spliced from DNM3OS (DNM3 opposite strand/antisense RNA) gene (Fig. 1A). In this work, we further determined circ_0005230 expression in CCA tissue samples and cell lines. Additionally, loss/gain of-function experiments were conducted to investigate cell growth, apoptosis, migration, and invasion potential affected by circ_0005230. Moreover, in vivo study was performed to validate the in vitro data. Importantly, the underlying mechanism was explored by dual luciferase reporter assays. Collectively, this work may help to form a novel effective therapeutics target for treating this devastating disease.
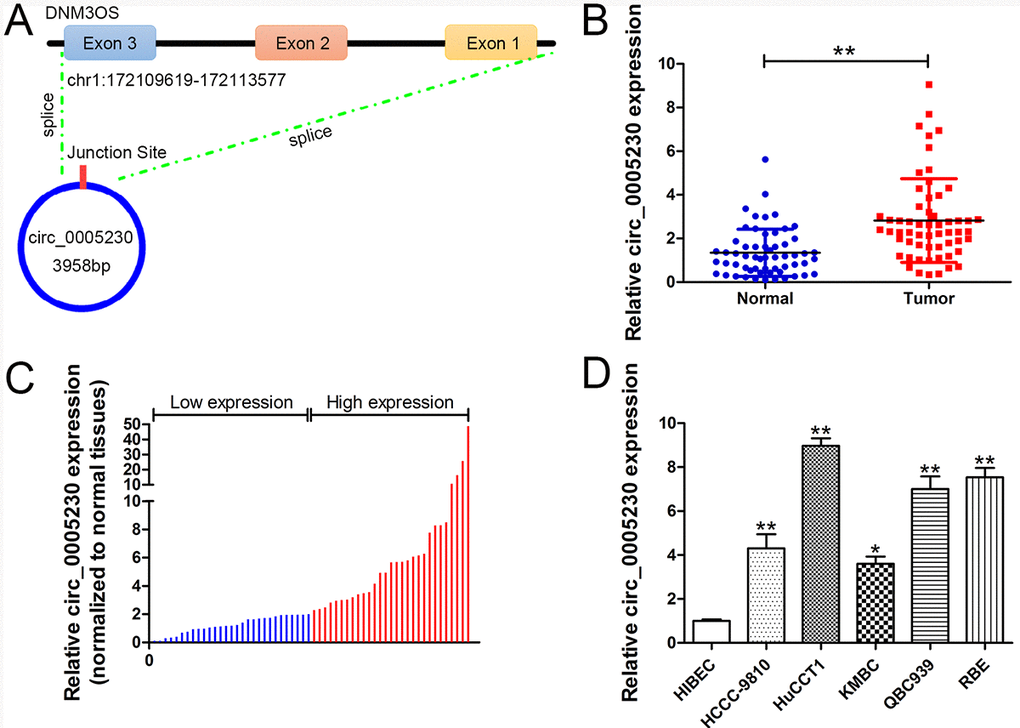
Figure 1. Relative expression of circ_0005230 in CCA tissues and cell lines and its clinical significance. (A) A schematic diagram of the genomic location and splicing pattern of circ_0005230. (B) Relative expression of circ_0005230 in CCA tissue samples and their paired non-cancerous tissue samples measured by qRT-PCR. (C) The patients were classified into two groups according to circ_0005230 expression. (D) Relative expression of circ_0005230 in CCA cell lines and normal cell line measured by qRT-PCR. *p<0.05, **p<0.01.
Results
Circ_0005230 expression is enhanced in CCA tissues and cells and correlates with patients’ clinical severity
The data from qRT-PCR uncovered that circ_0005230 expression was evidently enhanced in CCA specimens compared with the healthy tissues (2.10-fold, Fig. 1B). Therefore, fifty eight recruited CCA patients could be divided into high (>2.10) and low (<2.10) expression groups (Fig. 1C). Furthermore, similar results were identified in CCA cells relative to the normal biliary epithelial cells (HIBEC, Fig. 1D). After validating the elevation of circ_0005230 in CCA specimens and cell lines, its clinical implication was investigated. As analysis indicated, circ_0005230 expression is related to larger tumor size (p=0.028), positive lymph node invasion (p=0.003), and advanced TNM stages (p=0.020) for CCA patients. The other features were not linked to circ_0005230 expression (gender, age, tumor site, and differentiation grade, Table 1).
Table 1. Relationship between circ_0005230 expression and clinicopathologic features of CCA patients.
| Clinicopathologic | patients | circ_0005230 expression | p-value | |
| features | High | Low | ||
| Gender | ||||
| Male | 27 | 15(25.86%) | 12(20.69%) | 0.599 |
| Female | 31 | 14(24.14%) | 17(29.31%) | |
| Age | ||||
| <60 | 30 | 17(29.31%) | 13(22.41%) | 0.431 |
| ≥60 | 28 | 12(20.69%) | 16(27.59%) | |
| Tumor site | ||||
| Intrahepatic | 15 | 6(10.34%) | 9(15.52%) | 0.550 |
| Extrahepatic | 43 | 23(39.66%) | 20(34.48%) | |
| Tumor size | ||||
| <3cm | 37 | 14(24.14%) | 23(39.66%) | 0.028 |
| ≥3cm | 21 | 15(25.86%) | 6(10.34%) | |
| Lymph node invasion | ||||
| Present | 36 | 24(41.38%) | 12(20.69%) | 0.003 |
| Absent | 22 | 5(8.62%) | 17(29.31%) | |
| TNM stage | ||||
| I-II | 17 | 4(6.90%) | 13(22.41%) | 0.020 |
| III-IV | 41 | 25(43.10%) | 16(27.59%) | |
| Differentiation grade | ||||
| Well/moderately | 20 | 8(13.79%) | 12(20.69%) | 0.408 |
| Poorly/undifferentiated | 38 | 21(36.21%) | 17(29.31%) | |
Circ_0005230 facilitates HuCCT1 and KMBC cell progression in vitro
SiRNAs were used for transfection to silence circ_0005230 expression in HuCCT1 cells, (Fig. 2A). In addition, circ_0005230 was ectopically expressed in KMBC cells for the reason that it has the lowest level of circ_0005230 (Fig. 2B). Cell growth was evaluated by CCK-8 and clonogenic experiments and the data revealed that cell proliferation was attenuated in the si-circ-1/-2 groups relative to the negative control group. After circ_0005230 overexpression, cell viability and clone forming ability were both enhanced (Fig. 2C and D). We next investigated the effect of circ_0005230 on CCA cell apoptosis by flow cytometry and AO/EB staining assays. Apoptotic cells were measured at 48h post-transfection. Compared to si-NC group, cell apoptotic rate was elevated in si-circ-1 and si-circ-2 groups. On the contrary, cell apoptosis was inhibited after circ_0005230 upregulated in KMBC cells (Fig. 2E). Similar results were found in AO/EB double fluorescence staining assay (Fig. 2F). Cell migratory and invasive assays confirmed that cell metastatic ability was decreased after circ_0005230 knockdown in HuCCT1 cells. Whereas, in comparison with empty vector group, the cells with elevated circ_0005230 had a stronger migratory and invasive capacities (Fig. 2G).
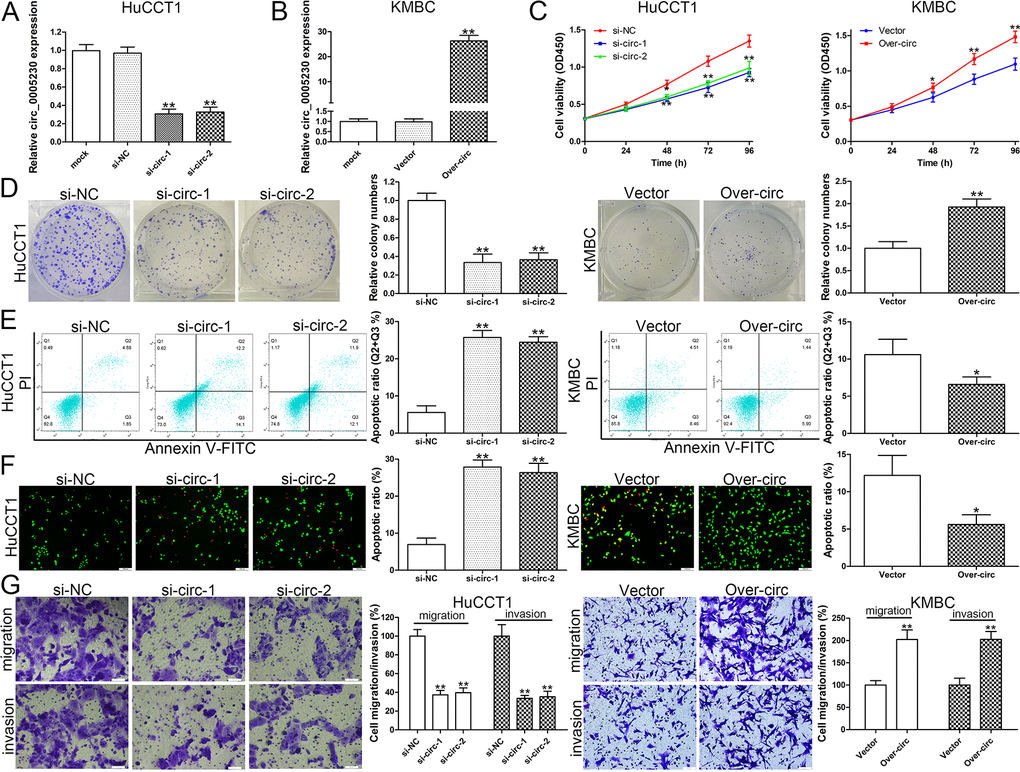
Figure 2. Circ_0005230 regulates cell growth, apoptosis, migration and invasion. (A) Circ_0005230 expression was detected after transfection in HuCCT1 cells by qRT-PCR. (B) Circ_0005230 expression was detected after transfection in KMBC cells by qRT-PCR. (C) CCK-8 assays were used to detect cell viability of HuCCT1 and KMBC cells after transfection. (D) Colony formation assays were used to detect the clone ability of HuCCT1 and KMBC cells after transfection. (E) Flow cytometric analysis were used to detect cell apoptosis of HuCCT1 and KMBC cells after transfection. (F) AO/EB staining was used to detect cell apoptosis of HuCCT1 and KMBC cells after transfection. (G) Transwell assays were used to detect cell migration and invasion capacities of HuCCT1 and KMBC cells after transfection. *p<0.05, **p<0.01.
Knockdown of circ_0005230 expression does not affect HIBEC cell growth and apoptosis
In order to clear whether silencing of circ_0005230 has side effect on normal biliary epithelial cells, we silenced circ_0005230 expression in HIBEC cells. As Fig. 3A showed, both of the two siRNAs had effective knockdown efficiencies. However, the cell viability curves based on CCK-8 assay indicated that the growth of HIBEC cells was not altered after circ_0005230 silenced (Fig. 3B). Additionally, apoptotic rate was not statistically significance affected in different transfected groups (Fig. 3C).
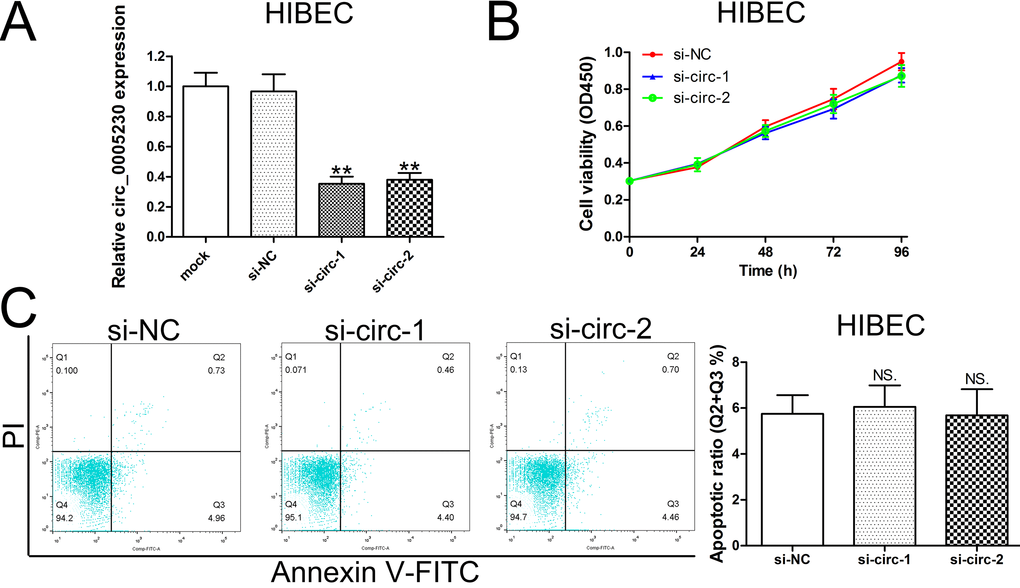
Figure 3. Decreased expression of circ_0005230 does not affect HIBEC cell proliferation and apoptosis. (A) HIBEC cells were transfected with siRNAs and siRNA-depletion efficiency was detected by qRT-PCR. (B) The proliferation of HIBEC cells after transfection was detected by CCK-8 assays. (C) The apoptosis of HIBEC cells after transfection was detected by flow cytometry.
Circ_0005230 directly sponges miR-1238 and miR-1299 in CCA cells
Various reports illustrated that circRNAs participates in post-transcriptional regulation by sponging several miRNAs [15,16]. To further evaluate the mechanisms of circ_0005230 exert in CCA, bioinformatics analysis was used to predict miRNAs that may be sponged by circ_0005230. Among all the predicted miRNAs, only miR-1238 and miR-1299 could be regulated by si-circ-1 and si-circ-2. The other recruited miRNAs were not altered statistically significant (Fig. 4A). The binding sites of miR-1238 and miR-1299 within circ_0005230 were shown in Fig. 4B. There are two potential binding sites of miR-1238 within circ_0005230 (94-100 and 214-220). Afterwards, the relative expression of miR-1238 and miR-1299 in CCA cancerous tissues was detected. Correlation analysis documented that either miR-1238 or miR-1299 was negatively correlated with circ_0005230 level in CCA tissue samples (Fig. 4C). For the enrolled CCA and HIBEC cells, we found miR-1238 and miR-1299 were both downregulated in CCA cell lines (Fig. 4D). Finally, we conducted luciferase reporter assay to confirm the binding capacity between miR-1238/1299 and circ_0005230. As Fig. 4E demonstrated, both of the binding sites of miR-1238 on circ_0005230 were functional. In addition, the luciferase activities of HuCCT1 and KMBC cells transfected with miR-1299 and circ_0005230 WT were decreased compared with the scramble control group (Fig. 4F).
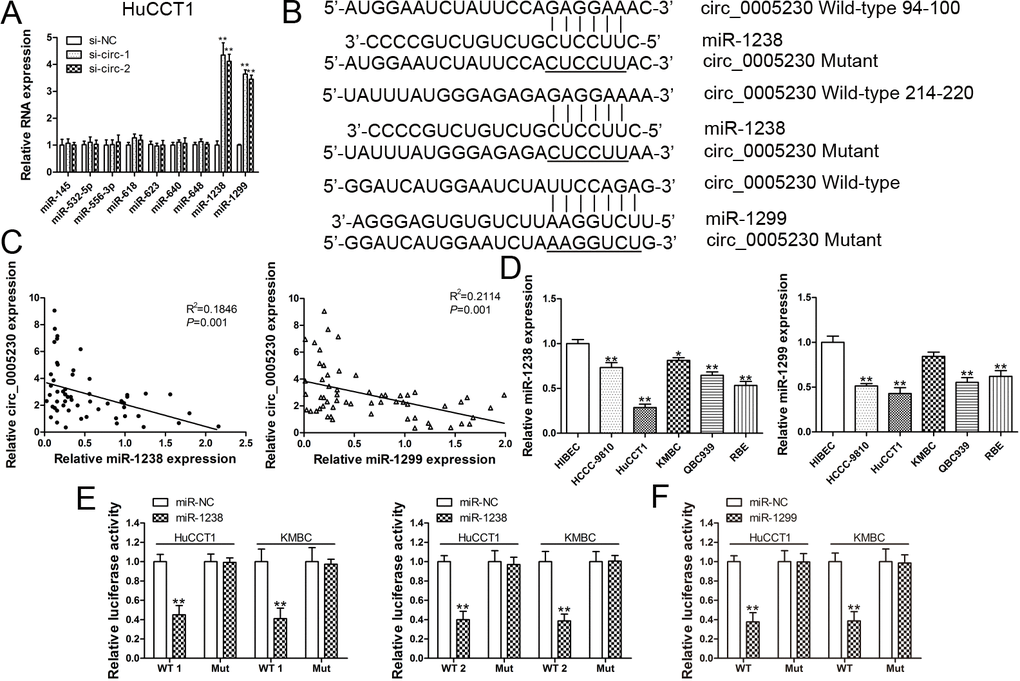
Figure 4. Circ_0005230 is a miRNA sponge for miR-1238 and miR-1299. (A) MiRNAs expression was detected after silencing of circ_0005230 in HuCCT1 cells. (B) Diagrammatic sketch of the binding sites for miR-1238 and miR-1299 in circ_0005230. (C) Correlation analysis of circ_0005230 and miR-1238/miR-1299 in CCA patients’ tissues. (D) qRT-PCR was performed to detect the expression of miR-1238/miR-1299 CCA cells and HIBEC cells. (E, F) Luciferase reporter assay showed that ectopic expression of miR-1238 and miR-1299 suppressed the activity of circ_0005230-WT in HuCCT1 and KMBC cells. *p<0.05, **p<0.01.
Circ_0005230 facilitates cell progression by sponging miR-1238 and miR-1299
After confirming that circ_0005230 could interact with miR-1238 and miR-1299 in HuCCT1 and KMBC cells, it is imperative to evaluate whether the oncogenic activities of circ_0005230 were attributed to its modulation on miR-1238/1299. As Fig. 5A showed, HuCCT1 cells were cotransfected with si-circ_0005230 and indicated inhibitor miRNA to reverse elevated miR-1238 and miR-1299. In KMBC cell line, upregulation of circ_0005230 evidently attenuated miR-1238 and miR-1299 levels. Further cotransfected with miR-1238 or miR-1299 mimics remarkably restored its level (Fig. 5B). For the part of functional assays, si-circ transfected with either miR-1238 or miR-1299 inhibitor partly rescued the tumor suppressing functions caused by si-circ_0005230 in HuCCT1 cells. Furthermore, silencing of miR-1238 and miR-1299 at the same time almost totally reversed the tumor suppressing activities of si-circ_0005230 (Fig. 5C and D). In KMBC cells, cell growth and invasion potential were partly attenuated after cotransfected with indicated miRNA mimics. In the over-circ_0005230, miR-1238 and miR-1299 co-transfection group, cell growth and invasion capacities were further decreased (Fig. 5E and F).
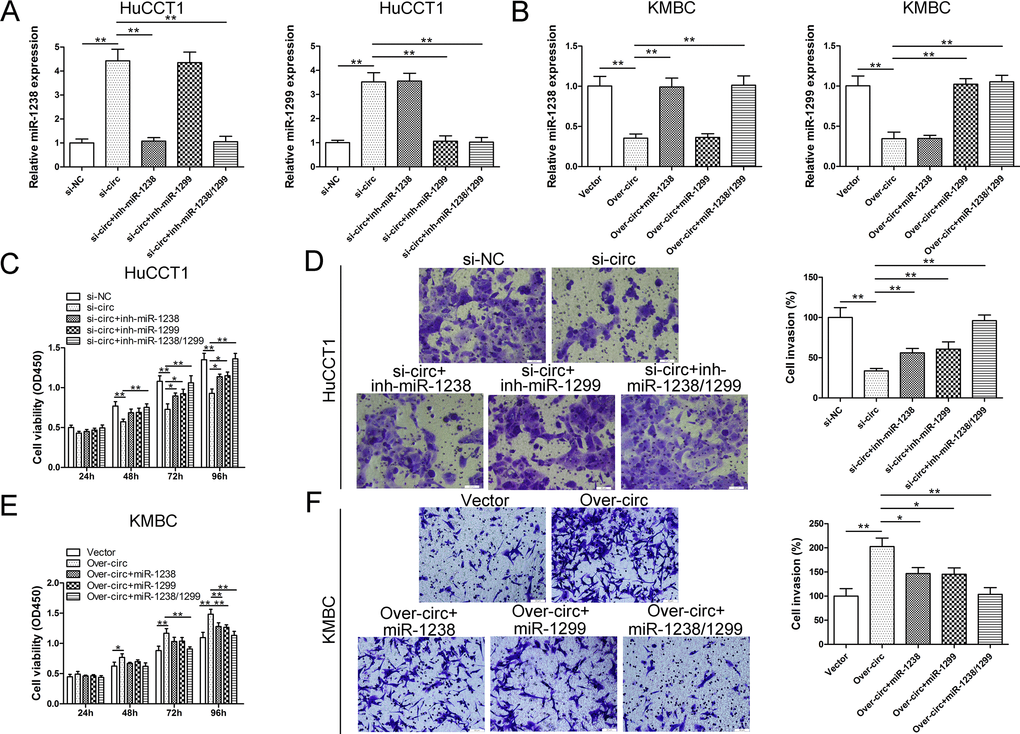
Figure 5. The oncogenic role of circ_0005230 is partly dependent on its regulation on miR-1238 and miR-1299. (A) Transfection with miR-1238 and miR-1299 inhibitor led to decreased expression of miR-1238 and miR-1299 in circ_0005230-downregulated HuCCT1 cells. (B) Transfection with miR-1238 and miR-1299 mimics led to elevated expression of miR-1238 and miR-1299 in circ_0005230-upregulated KMBC cells. (C, D) Inhibition of miR-1238 and/or miR-1299 rescued the proliferation and invasion of circ_0005230-downregulated HuCCT1 cells. (E, F) Ectopic expression of miR-1238 and/or miR-1299 rescued the proliferation and invasion of circ_0005230-upregulated KMBC cells.*p<0.05, **p<0.01.
Circ_0005230 facilitates HuCCT1 cell growth and metastasis in vivo
To validate the oncogenic role of circ_0005230 in CCA, in vivo study was performed. The xenografts formed from sh-circ_0005230-1 were significantly smaller in comparison with the shCtrl group tumors. In addition, sh-circ_0005230-1 cotransfected with miR-1238/1299 inhibitor could reverse the tumor suppressing role of sh-circ_0005230-1 (Fig. 6A and B). What’s more, the proliferative marker, Ki67 expression was weaker in the tumors formed from sh-circ_0005230-1 compared with control group. After co-silencing of circ_0005230, miR-1238 and miR-1299, Ki67 expression was elevated proved by IHC assay (Fig. 6C). Further lung metastatic tumor model demonstrated that downregulation of circ_0005230 resulted in less metastatic lung nodules, which is in accordance with the in vitro results (Fig. 6D-F).
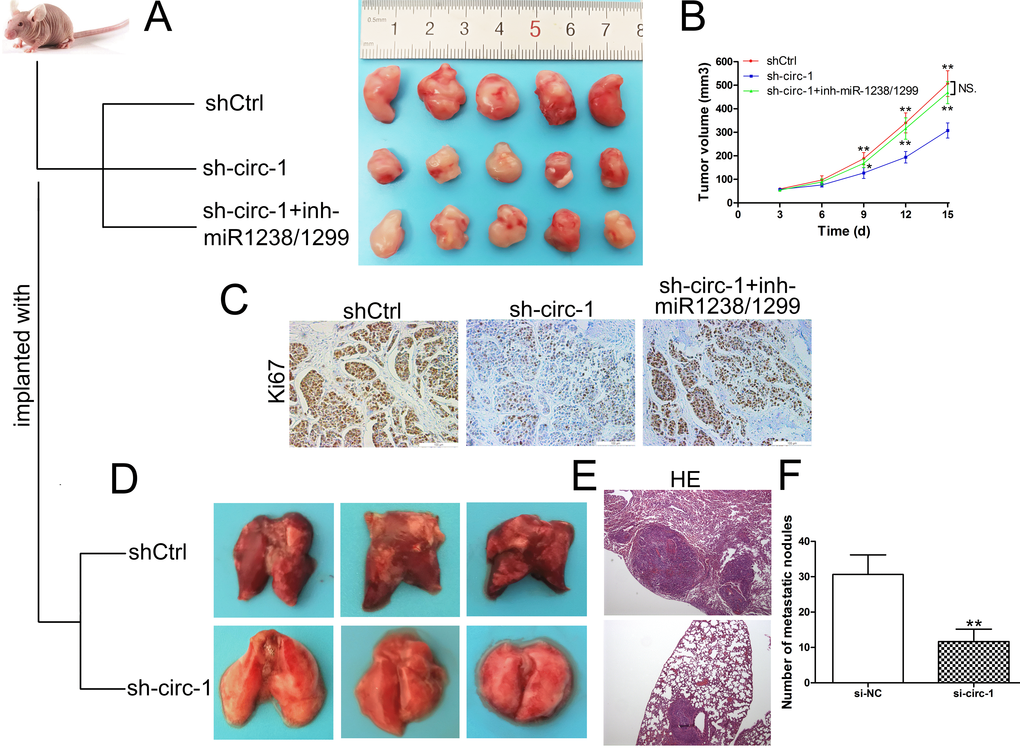
Figure 6. Circ_0005230 promotes cell growth and metastasis in vivo. (A) Tumors from nude mice after injection of transfected HuCCT1 cells. (B) Tumor volume was monitored every 3 days. (C) Ki67 expression and location were determined by IHC. (D) Lungs from experimental metastasis animal model of each group are shown. (E) HE staining was used to stain the resected lungs. (F) The number of tumor nodules on lung surfaces from two groups. *p<0.05, **p<0.01.
Discussion
CCA is a devastating malignancy with increasing incidence across the world. Due to its insensitive to conventional chemotherapy, the outcomes of clinical therapy are dismal [1,2]. Emerging evidence verified that circRNAs can play critical roles in the pathogenesis of malignancies. For example, Li et al. revealed that upregulated hsa_circ_0007534 could regulate osteosarcoma cell progression by affecting AKT/GSK-3β signal [17]. Several studies also demonstrated the diagnostic or prognostic role of circRNAs in malignancies [18,19]. For CCA, we previous found Cdr1as might be regarded as a potential biomarker to predict tumor progression and poor prognosis [20]. In this study, the elevation of circ_0005230 was confirmed in CCA tissues specimens and cells in comparison with their matched non-cancerous samples and HIBEC, respectively. Moreover, the enhanced expression of circ_0005230 is related to larger tumor size (p=0.028), positive lymph node invasion (p=0.003), and higher TNM stages (p=0.020) for CCA patients.
Based on these observations, a series of functional assays were conducted on HuCCT1 (highest circ_0005230 expression) and KMBC cells (lowest circ_0005230 expression). The data documented that circ_0005230 could markedly enhance cell proliferation, migration and invasion potential and decrease cell apoptosis proved by CCK-8, clonogenic, flow cytmetric, AO/EB staining and transwell assays. Further animal study validated the in vitro data that circ_0005230 could facilitate cell growth and metastasis in CCA. These findings preliminary documented circ_0005230 as an oncogenic circRNA and a potential therapeutic target for CCA treatment. HIBEC cell line was used to explore whether knockdown of circ_0005230 may have side effect on normal biliary epithelial cells. The data indicated that silenced circ_0005230 did not affect the growth and apoptosis of HIBEC cells, which implied that the regulatory network of circ_0005230 in HIBEC is different from CCA cells and circ_0005230 does not play a major role in HIBEC cell growth. The gene expression regulated by circ_0005230 in HIBEC is different from CCA cells, indicating the mechanism of circ_0005230 is tissue specific.
CircRNAs could act as molecular sponges to bind to miRNAs or scaffold RBPs to exert their biological functions in carcinogenesis and tumor progression. For instance, Xiong et al. reported that circRNA ZNF609 upregulates FOXP4 expression to regulate renal carcinoma cell progression via sponging miR-138-5p [21]. Previously, we found increased expression of circ_0005230 could function as a competitive endogenous RNA to increase CBX8 expression by sponging miR-618 in BC [14]. Interestingly, miR-618 was almost not affected by circ_0005230 in the two selected CCA cells, which suggested that the mechanism of circ_0005230 maybe tissue specific. Among all the predicted miRNAs, only miR-1238 and miR-1299 were negatively correlated with circ_0005230 level. To clear whether circ_0005230 could directly interact with miR-1238/miR-1299, luciferase reporter assay was induced. In line with our expectation, the predicted binding sites were functional. The study relevant to the role of miR-1238 in cancer progression is rare. Only one report by Shi et al. identified miR-1238 as a tumor suppressor in cancer initiation and development. They uncovered that miR-1238 inhibits non-small cell lung cancer (NSCLC) cell growth partly by repressing LHX2 [22]. In the current study, we first proved miR-1238 as a tumor suppressive miRNA in CCA. MiR-1299 is a well-studied miRNA in some kind of malignancies and functions as a tumor suppressor. For example, Zhu et al. found miR-1299 could suppress hepatocellular carcinoma (HCC) cell growth by targeting CDK6 [23]. Another study indicated that ectopic expression of miR-1299 could downregulate STAT3 pathway to inhibit colon cancer cell growth [24]. Sang et al. demonstrated that Cdr1as maintains metastatic phenotypes of triple-negative breast cancer as a ceRNA of miR-1299 to target MMPs [25]. In this study, rescue experiments were used to uncover that the oncogenic role of circ_0005230 was partly attributed to its inhibition on miR-1238 and miR-1299. Previously, we have documented that Cdr1as is elevated in CCA, whether miR-1299 could also be sponged by Cdr1as in CCA is needed to further investigation. In addition, how miR-1238 and miR-1299 suppress CCA cell progression and the downstream targets of miR-1238 and miR-1299 are still needed to investigate. Along with future studies, circ_0005230 may be a rational therapeutic target for CCA.
Taken together, we identified circ_0005230 elevation in CCA tissues and cells and related to patients’ clinical severity. Furthermore, circ_0005230 plays an oncogenic role partly by sponging miR-1238 and miR-1299. Collectively, this work may help to form a novel effective therapeutics target for treating this devastating disease.
Materials and Methods
Patient selection and tissue samples
The current study was reviewed by the Ethics Review Committees of the Second Affiliated Hospital of Harbin Medical University. Fifty-eight paired tissues and matched noncancerous specimens were acquired from the CCA patients who underwent operation from April 2012 to March 2014. Preoperative intervention was not adopted before the surgery and the samples were stored in liquid nitrogen.
Cell culture
HCCC-9810 and RBE were purchused from Type Culture of Chinese Academy of Sciences (Shanghai, China). Normal biliary cell line, HIBEC, and the other CCA cells (KMBC, HuCCT1 and QBC939) were stored in our laboratory. Above five CCA cell lines and normal biliary epithelial cells were cultured in RPMI-1640 with 10% FBS (Invitrogen Life Technologies, USA) in a humidified air with 5% CO2 at 37°C.
RNA isolation and qRT-PCR
Total RNA was isolated by Trizol (Thermo Fisher Scientific, USA). The Spectrophotometer (Implen, Germany) was applied to assess the concentration and quantity of the extracted RNA. Quantitative PCR was performed through Power SYBR Green (Takara Biotechnology, China). Relative expression of the genes were determined by 2−ΔΔCt.
Cell transfection
Two siRNAs specifically targeting to the back-spliced site of circ_0005230 (si-circ-1/-2) and the negative control were acquired from GenePharma (Shanghai, China). The target sequences of si-circ_0005230 are shown below: si-circ-1: 5’-CAATATACTCTTTGTTTTGCA-3’. si-circ-2: 5’-CTCTTTGTTTTGCACACTAGG-3’. A pcDNA3.1 (+) circRNA mini vector was applied to clone the whole circRNA sequence and artificial inverted repeats. MiR-1238/1299 mimics/inhibitor were purchased from GenePharma (Shanghai, China). Lipofectamine 3000 was used for transfection according to the manufacturer’s protocols.
Cell proliferation assays
Cell vitality was evaluated by CCK-8 (Beyotime, Beijing, China). The transfected cells in each 96-well plate were cultured overnight prior to adding with CCK-8 reagent in dark environment. OD values was quantified by a microplate reader at 450nm.
To detect the clone-forming ability of CCA cells, monoplast suspension of HuCCT1 and KMBC was seeded in 6-well plates at a same number of cells per well and incubated in medium containing 10% FBS. After 12 days, the macroscopic colonies were stained, photographed and calculated.
Cell apoptosis detection
Apoptosis was measured by FACScan flow cytometer (BD Biosciences, USA). Transfected CCA cells were collected and incubated with 5μL of AnnexinV-FITC and 5μL of PI (BD, CA, USA) for 10min. The treated cells were subjected to flow cytometric analysis to detect apoptotic cells.
Acridine orange/ethidium bromide (AO/EB) double fluorescence staining assay was performed to further validate the data of flow cytometric analysis. The transfected HuCCT1 and KMBC cells were stained with prepared AO/EB mixing solution (1:1) for 5min before observation and photographing.
Transwell assay
Cell metastatic properties were determined by Transwell experiments. Both migration and invasion assays were carried out through following protocols: serum-free cell suspensions were prepared and transferred to the upper chamber with 0.2ml cell suspension per well (pre-coated by Matrigel for invasion assay). The lower chamber was filled with medium supplemented with 10% FBS. Cells were cultivated for 24h and membranes were stained by crystal violet for 15min. At last, the cells were counted.
Target prediction and luciferase reporter assay
Circular RNA Interactome was applied to predict the potential miRNAs targeting to circ_0005230. Dual luciferase reporter assay was carried out followed by the previous studies [14,26].
Animal study
BALB/c nude mice (n=5 in each group) were purchased from Vital River (Beijing, China) and the study was authorized by the Animal Ethics Committee of the Second Affiliated Hospital of Harbin Medical University. Six weeks old female nude mice were received 8 × 106 transfected HuCCT1 cells to induce xenograft. The volume of tumors were detected every three days. The mice were sacrificed at 15 days after injection. Immunohistochemistry (IHC) assay was conducted on the paraffin-embedded tissue samples from the mice to detect the location and relative expression of Ki67 (Abcam, Cambridge, MA, USA).
The stably transfected HuCCT1 cells (1 × 107) were inoculated into the tail vein of each mice to construct the late stage lung metastatic model (n=3 in each group). Four weeks later, the mice were killed and their lungs were removed and subjected to hematoxylin and eosin (HE) dyeing. Finally, the number of lung metastases nodules were counted.
Data analysis
All data are expressed as means ± standard deviation and analyzed using SPSS 19.0 software (SPSS, USA). The clinical significance of circ_0005230 in CCA was evaluated by using fisher’ exact test. Statistical difference was analyzed using t-test or one-way ANOVA. A p value less than 0.05 was considered statistically significant.
Conflicts of Interest
The authors report no conflict of interests in this work.
Funding
This study was funded by National Key Research and Development Program of China (Grant No. 2017YFC1308600); China Postdoctoral Science Foundation (Grant No. 2018M641849 and 2018M640311); Heilongjiang Postdoctoral Science Foundation (Grant No. LBH-Z18107 and LBH-Z18112); The Fundamental Research Funds for the Heilongjiang Provincial Universities; Postgraduate Innovative Research Project of Harbin Medical University (Grant No. YJSCX2016-21HYD); Foundation of Key Laboratory of Myocardial Ischemia, Ministry of Education (Grant No. KF201810); Chen Xiaoping Foundation for the Development of Science and Technology of Hubei Province (Grant No. CXPJJH11800004-001 and CXPJJH11800004-003).
References
- 1. Khan SA, Davidson BR, Goldin RD, Heaton N, Karani J, Pereira SP, Rosenberg WM, Tait P, Taylor-Robinson SD, Thillainayagam AV, Thomas HC, Wasan H, and British Society of Gastroenterology. Guidelines for the diagnosis and treatment of cholangiocarcinoma: an update. Gut. 2012; 61:1657–69. https://doi.org/10.1136/gutjnl-2011-301748 [PubMed]
- 2. Bergquist A, von Seth E. Epidemiology of cholangiocarcinoma. Best Pract Res Clin Gastroenterol. 2015; 29:221–32. https://doi.org/10.1016/j.bpg.2015.02.003 [PubMed]
- 3. Rizvi S, Borad MJ, Patel T, Gores GJ. Cholangiocarcinoma: molecular pathways and therapeutic opportunities. Semin Liver Dis. 2014; 34:456–64. https://doi.org/10.1055/s-0034-1394144 [PubMed]
- 4. Skipworth JR, Olde Damink SW, Imber C, Bridgewater J, Pereira SP, Malagó M. Review article: surgical, neo-adjuvant and adjuvant management strategies in biliary tract cancer. Aliment Pharmacol Ther. 2011; 34:1063–78. https://doi.org/10.1111/j.1365-2036.2011.04851.x [PubMed]
- 5. Ponting CP, Belgard TG. Transcribed dark matter: meaning or myth? Hum Mol Genet. 2010; 19:R162–68. https://doi.org/10.1093/hmg/ddq362 [PubMed]
- 6. Derrien T, Johnson R, Bussotti G, Tanzer A, Djebali S, Tilgner H, Guernec G, Martin D, Merkel A, Knowles DG, Lagarde J, Veeravalli L, Ruan X, et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res. 2012; 22:1775–89. https://doi.org/10.1101/gr.132159.111 [PubMed]
- 7. Jeck WR, Sharpless NE. Detecting and characterizing circular RNAs. Nat Biotechnol. 2014; 32:453–61. https://doi.org/10.1038/nbt.2890 [PubMed]
- 8. Haimovich G, Medina DA, Causse SZ, Garber M, Millán-Zambrano G, Barkai O, Chávez S, Pérez-Ortín JE, Darzacq X, Choder M. Gene expression is circular: factors for mRNA degradation also foster mRNA synthesis. Cell. 2013; 153:1000–11. https://doi.org/10.1016/j.cell.2013.05.012 [PubMed]
- 9. Hao L, Rong W, Bai L, Cui H, Zhang S, Li Y, Chen D, Meng X. Upregulated circular RNA circ_0007534 indicates an unfavorable prognosis in pancreatic ductal adenocarcinoma and regulates cell proliferation, apoptosis, and invasion by sponging miR-625 and miR-892b. J Cell Biochem. 2019; 120:3780–89. https://doi.org/10.1002/jcb.27658 [PubMed]
- 10. Liu H, Xue L, Song C, Liu F, Jiang T, Yang X. Overexpression of circular RNA circ_001569 indicates poor prognosis in hepatocellular carcinoma and promotes cell growth and metastasis by sponging miR-411-5p and miR-432-5p. Biochem Biophys Res Commun. 2018; 503:2659–65. https://doi.org/10.1016/j.bbrc.2018.08.020 [PubMed]
- 11. Huang JL, Zheng L, Hu YW, Wang Q. Characteristics of long non-coding RNA and its relation to hepatocellular carcinoma. Carcinogenesis. 2014; 35:507–14. https://doi.org/10.1093/carcin/bgt405 [PubMed]
- 12. Zhang Y, Liu H, Li W, Yu J, Li J, Shen Z, Ye G, Qi X, Li G. CircRNA_100269 is downregulated in gastric cancer and suppresses tumor cell growth by targeting miR-630. Aging (Albany NY). 2017; 9:1585–94. https://doi.org/10.18632/aging.101254 [PubMed]
- 13. Xu Y, Yao Y, Zhong X, Leng K, Qin W, Qu L, Cui Y, Jiang X. Downregulated circular RNA hsa_circ_0001649 regulates proliferation, migration and invasion in cholangiocarcinoma cells. Biochem Biophys Res Commun. 2018; 496:455–61. https://doi.org/10.1016/j.bbrc.2018.01.077 [PubMed]
- 14. Xu Y, Yao Y, Leng K, Ji D, Qu L, Liu Y, Cui Y. Increased expression of circular RNA circ_0005230 indicates dismal prognosis in breast cancer and regulates cell proliferation and invasion via miR-618/CBX8 signal pathway. Cell Physiol Biochem. 2018; 51:1710–22. https://doi.org/10.1159/000495675 [PubMed]
- 15. Li L, Guo L, Yin G, Yu G, Zhao Y, Pan Y. Upregulation of circular RNA circ_0001721 predicts unfavorable prognosis in osteosarcoma and facilitates cell progression via sponging miR-569 and miR-599. Biomed Pharmacother. 2019; 109:226–32. https://doi.org/10.1016/j.biopha.2018.10.072 [PubMed]
- 16. Li Y, Hu J, Li L, Cai S, Zhang H, Zhu X, Guan G, Dong X. Upregulated circular RNA circ_0016760 indicates unfavorable prognosis in NSCLC and promotes cell progression through miR-1287/GAGE1 axis. Biochem Biophys Res Commun. 2018; 503:2089–94. https://doi.org/10.1016/j.bbrc.2018.07.164 [PubMed]
- 17. Li B, Li X. Overexpression of hsa_circ_0007534 predicts unfavorable prognosis for osteosarcoma and regulates cell growth and apoptosis by affecting AKT/GSK-3β signaling pathway. Biomed Pharmacother. 2018; 107:860–66. https://doi.org/10.1016/j.biopha.2018.08.086 [PubMed]
- 18. Fang S, Pan J, Zhou C, Tian H, He J, Shen W, Jin X, Meng X, Jiang N, Gong Z. Circular RNAs serve as novel biomarkers and therapeutic targets in cancers. Curr Gene Ther. 2018. https://doi.org/10.2174/1566523218666181109142756 [Epub ahead of print] [PubMed]
- 19. Shuai M, Hong J, Huang D, Zhang X, Tian Y. Upregulation of circRNA_0000285 serves as a prognostic biomarker for nasopharyngeal carcinoma and is involved in radiosensitivity. Oncol Lett. 2018; 16:6495–501. https://doi.org/10.3892/ol.2018.9471 [PubMed]
- 20. Jiang XM, Li ZL, Li JL, Xu Y, Leng KM, Cui YF, Sun DJ. A novel prognostic biomarker for cholangiocarcinoma: circRNA Cdr1as. Eur Rev Med Pharmacol Sci. 2018; 22:365–71. https://doi.org/10.26355/eurrev_201801_14182 [PubMed]
- 21. Xiong Y, Zhang J, Song C. CircRNA ZNF609 functions as a competitive endogenous RNA to regulate FOXP4 expression by sponging miR-138-5p in renal carcinoma. J Cell Physiol. 2019; 234: 10646–10654. https://doi.org/10.1002/jcp.27744 [PubMed]
- 22. Shi X, Zhan L, Xiao C, Lei Z, Yang H, Wang L, Zhao J, Zhang HT. miR-1238 inhibits cell proliferation by targeting LHX2 in non-small cell lung cancer. Oncotarget. 2015; 6:19043–54. https://doi.org/10.18632/oncotarget.4232 [PubMed]
- 23. Zhu H, Wang G, Zhou X, Song X, Gao H, Ma C, Chang H, Li H, Liu FF, Lu J, Ma J. miR-1299 suppresses cell proliferation of hepatocellular carcinoma (HCC) by targeting CDK6. Biomed Pharmacother. 2016; 83:792–97. https://doi.org/10.1016/j.biopha.2016.07.037 [PubMed]
- 24. Wang Y, Lu Z, Wang N, Zhang M, Zeng X, Zhao W. MicroRNA-1299 is a negative regulator of STAT3 in colon cancer. Oncol Rep. 2017; 37:3227–34. https://doi.org/10.3892/or.2017.5605 [PubMed]
- 25. Sang M, Meng L, Liu S, Ding P, Chang S, Ju Y, Liu F, Gu L, Lian Y, Geng C. Circular RNA ciRS-7 Maintains Metastatic Phenotypes as a ceRNA of miR-1299 to Target MMPs. Mol Cancer Res. 2018; 16:1665–75. https://doi.org/10.1158/1541-7786.MCR-18-0284 [PubMed]
- 26. Xu Y, Yao Y, Jiang X, Zhong X, Wang Z, Li C, Kang P, Leng K, Ji D, Li Z, Huang L, Qin W, Cui Y. SP1-induced upregulation of lncRNA SPRY4-IT1 exerts oncogenic properties by scaffolding EZH2/LSD1/DNMT1 and sponging miR-101-3p in cholangiocarcinoma. J Exp Clin Cancer Res. 2018; 37:81. https://doi.org/10.1186/s13046-018-0747-x [PubMed]


