Introduction
Chronic lymphocytic leukemia (CLL), a heterogeneous disease characterized by the monoclonal expansion of B cells, is the commonest leukemia in the western hemisphere [1, 2]. However, the pathogenesis mechanism of CLL remains elusive. MicroRNAs (miRNAs) are a large class of non-coding RNAs (ncRNAs) involved in tumorigenesis and have been reported to exert effects in the progression of multiple diseases [3–5]. Several current studies have shed lights on the aberrantly expressed miRNAs in CLL and demonstrated their potential roles as novel biomarkers and therapeutic targets [6, 7]. For example, the upregulated miR-92a-3p in CLL patients predicts a favorable prognostic outcome [8]. miR-125a-5p and miR-34a-5p are considered as promising indicators of CLL Richter syndrome transformation [9]. miR-34a is revealed to participate in B-cell receptor (BCR) signaling and thus affects CLL patients’ treatment [10].
miR-337-3p which has been detected to be downregulated in clear cell renal cell carcinoma [11], gastric cancer [12] and hepatocellular carcinoma [13, 14], could function as a suppressive factor in cell proliferation, tumor growth and metastasis through directly regulating target genes. Contrary to these previous reports, in our research, we accidentally found that miR-337-3p exhibited obviously high expression in peripheral blood mononuclear cell (PBMCs) of CLL patients compared with normal controls. The current results inspired us to further explore its regulatory mechanism.
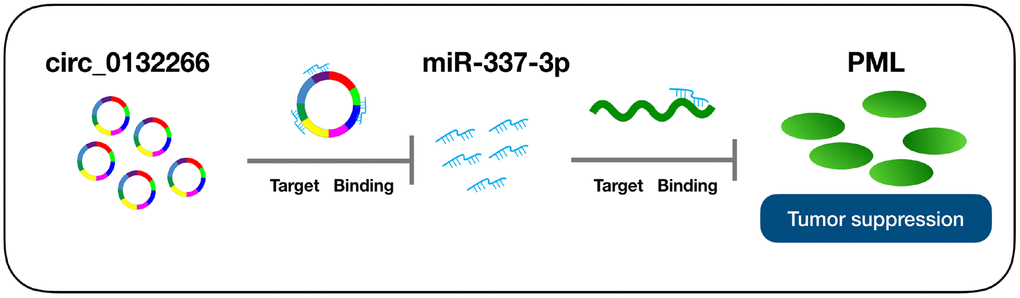
Circular RNAs (circRNAs), widely expressed in human transcriptome, have been disclosed to function as a competitive endogenous RNA (ceRNA) via sponging miRNAs and further responsible for the expression of mRNA and the occurrence and development of diseases [15, 16]. Herein, we identified a CLL-related circRNA, circ_0132266, with an obviously low expression level and was negatively associated with miR-337-3p. Our research, for the first time, uncovered the functions of circ_0132266, which could compete for miR-337-3p and further influence the expression abundance of promyelocytic leukemia protein (PML) and thus promote CLL progression. Our results may provide a novel diagnostic and therapeutic candidate for CLL.
Results
miR-337-3p upregulated in CLL, promoted cell proliferation and inhibited cell apoptosis
Our previous studies occasionally noted that miR-337-3p was upregulated in CLL patients compared with healthy individuals. To further explore the expression pattern of miR-337-3p, we collected PBMC samples from thirty newly diagnosed CLL patients and thirty healthy volunteers. An increased expression of miR-337-3p was validated through qRT-PCR analysis, confirming the observations of our previous study (Figure 1A).
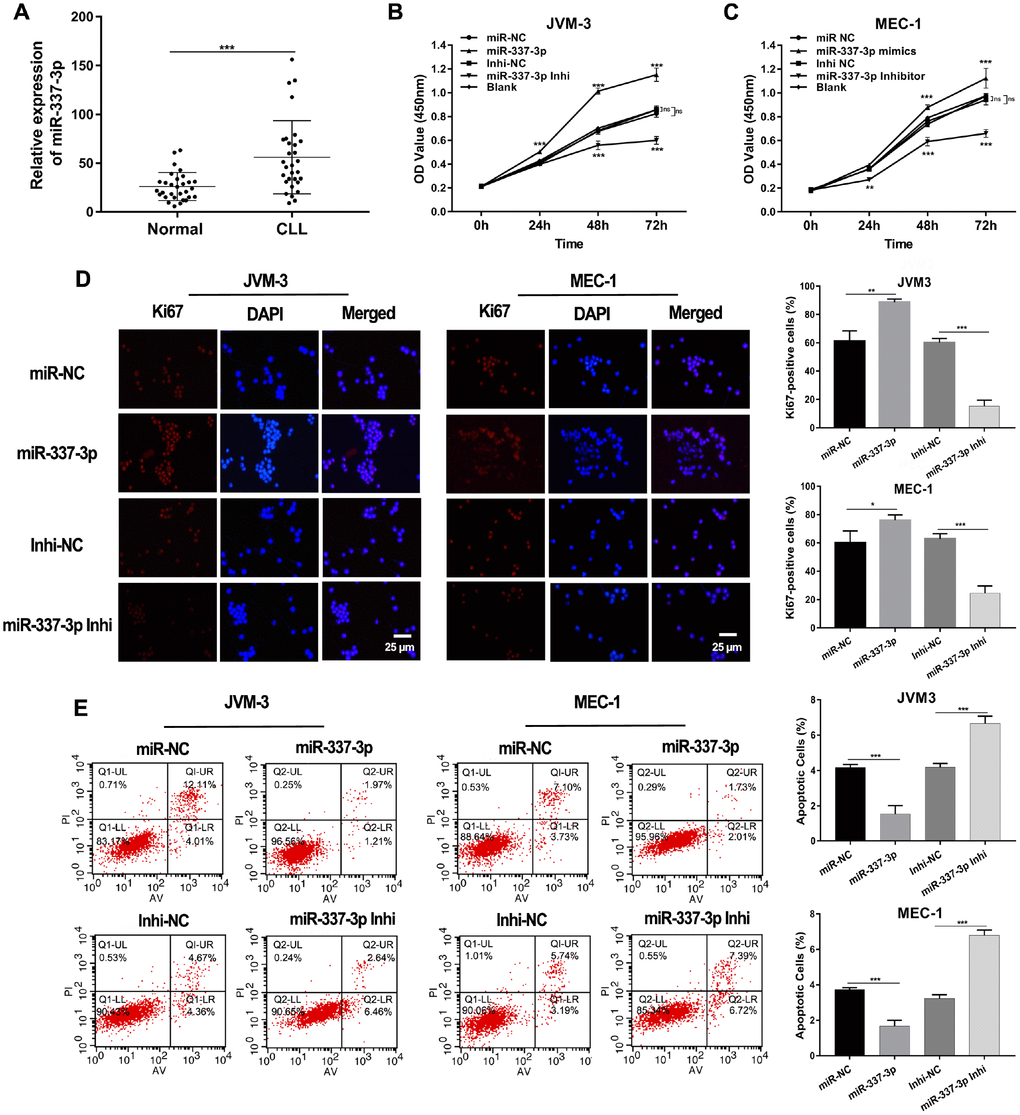
Figure 1. Expression and functions of miR-337-3p in CLL. (A) Relative expression levels of miR-337-3p in PBMC samples from thirty treatment-naïve CLL patients and thirty normal donors. (B, C) CCK8 assay was performed to assess the influence of miR-337-3p on CLL cells proliferation. (D) IF assay analyzed the proliferative viability of JVM-3 and MEC-1 after transfected with miR-337-3p mimics and inhibitor and ki67-positive cells rate were calculated. (E) Cell apoptosis was detected by FCM to verify the effects of miR-337-3p and the percentage of apoptotic cells was quantified. Three biological replicates were performed per condition and mean values ± SD are displayed (*P < 0.05; **P < 0.01; ***P < 0.001; ns, not significant).
Subsequent cell proliferation and apoptosis assays were performed to dissect functions of miR-337-3p in CLL. Two CLL cell lines (MEC-1 and JVM-3) were used in our research. The growth curves of cells were detected via CCK8 assay. The results showed that ectopic expression of miR-337-3p promoted cell growth and this could be rescued by inhibition of miR-337-3p (Figure 1B, 1C, Supplementary Figure 1). The proliferation potential detected by immunofluorescence (IF) also assured the functions induced by miR-337-3p (Figure 1D). Flow cytometry examined the cell apoptosis level (Figure 1E), indicating that miR-337-3p could inhibit cell apoptosis and the quantification of apoptotic cell was shown in Figure 1G.
Annotation for target genes of miR-337-3p
To address how miR-337-3p functioned, we used bioinformatic analysis to forecast the target genes directly modulated by miR-337-3p. Three genes, WD repeat domain 26 (WDR26), promyelocytic leukemia protein (PML), poly(A) polymerase beta (PAPOLB) were preliminarily screened through overlapping the prediction results of miRNA recognition elements by Targetscan, miRDB and Pictar (Figure 2A). And the postulated binding sites between miR-337-3p and the candidate genes were shown in Figure 2B. Luciferase reporter vectors containing the wild type (WT) or mutant genes 3′UTR and miR-337-3p mimics were co-transfected into HEK-293T cells. Intriguingly, the luciferase activity of wild type PML 3′UTR evidently decreased in miR-337-3p mimics group while the activity of WDR26 and PAPOLB exhibited no difference (Figure 2C). Simultaneously, we detected the mRNA expression levels of WDR26, PML, PAPOLB and noted that transfection of miR-337-3p mimics downregulated the expression of PML (Figure 2D), these results were confirmed by western blot (Figure 2E). PML, originally identified to be involved in the rise of PML–RARα and RARα–PML fusion proteins, is a well-known tumor suppressor by regulating cell apoptosis and cell cycle [17–20]. Through TCGA database, we noticed that PML was obviously downregulated in CLL patients (Figure 2F). Our results further verified the low expression of PML and the negative correlation between PML and miR-337-3p expression detected by qRT-PCR in a newly diagnosed CLL cohort (Figure 2G, 2H). These results collectively indicated that PML was a direct target of miR-337-3p and miR-337-3p exerted functions of promoting cell proliferation and protecting cells from apoptosis.
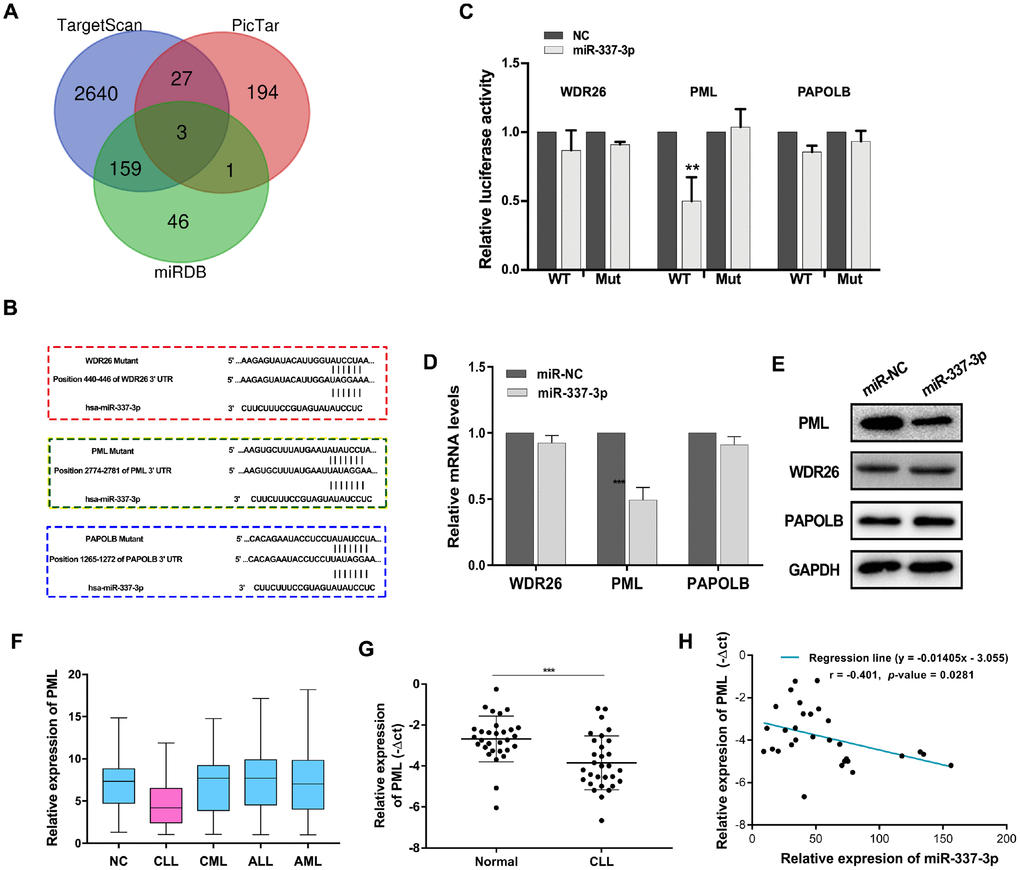
Figure 2. PML is a target gene of miR-337-3p. (A) Target genes were screened through overlapping websites of Targetscan (http://www.targetscan.org/vert_71/), miRDB (http://mirdb.org/) and Pictar (https://pictar.mdc-berlin.de/). (B) Potential binding sites between miR-337-3p and candidate genes. (C) Dual-luciferase reporter assay to verify predicted bindings. (D, E) The change of mRNAs and proteins expression after transfection with miR-337-3p mimics. (F) PML relative expression in leukemia and patients from TCGA database. (G) PML relative expression in CLL and normal patients PBMCs detected by qRT-PCR (30 CLL patients vs 30 healthy volunteers). (H) Correlation analysis between miR-337-3p and PML in CLL PBMCs.
Circ_0132266, identified as a novel downregulated circRNA in CLL, was a potential modulator of miR-337-3p
CircRNAs, a novel type of ncRNAs, have been proposed to regulate mRNAs expression and ultimately participate in the progression of diseases through sponging miRNAs [15, 21]. Since miR-337-3p was upregulated in CLL whereas previous reports have elucidated that downregulated miR-337-3p in solid tumors could promote tumor growth, invasion, and metastasis [12, 13], it inspired our interest in exploring whether miR-337-3p was modulated by specific circRNA(s).
To determine the circRNA(s) responsible for the regulation of miR-337-3p, we screened circBank database (http://www.circbank.cn) through a detailed screening system and finally screened out three most promising candidates, hsa_circ_0004731, hsa_circ_0132266 (circ_0132266), hsa_circ_0029937 (Figure 3A). The abundance of the three circRNAs was then examined in CLL and normal patients. Circ_0132266 was remarkably decreased in CLL patients while the other two had no difference (Figure 3B). Meanwhile, the expression of miR-337-3p was found to be negatively associated with circ_0132266 (Figure 3C).
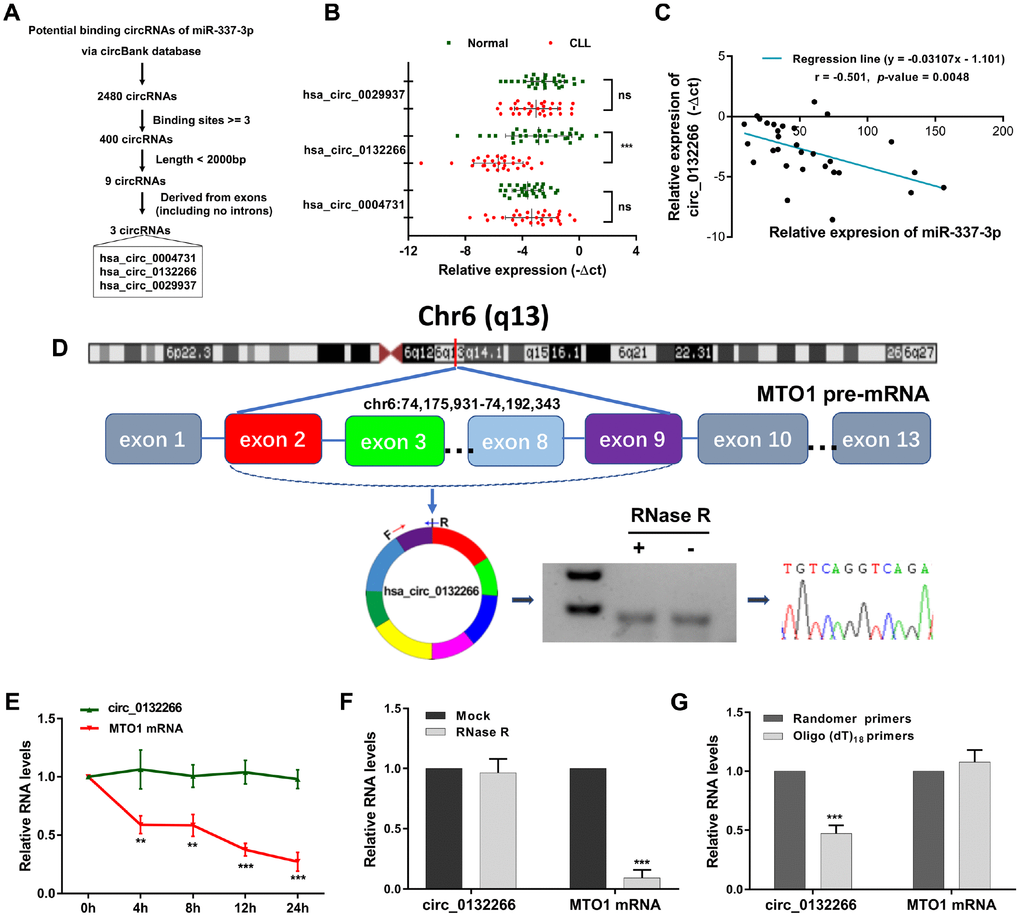
Figure 3. Characteristics of circ_0132266 in CLL patients and cell lines. (A) The schematic showed the screening rules of circRNAs that might regulate miR-337-3p. (B) Relative expression of the three candidate circRNAs in CLL patients and healthy individuals (n=30). (C) Expression correlation between miR-337-3p and circ_0132266 in CLL PBMCs. (D) The genomic loci of the MTO1 gene and circ_0132266. The expression of circ_0132266 was detected by qRT-PCR and was validated by Sanger sequencing. The relative expression of circ_0132266 and MTO1 mRNA after treatment of (E) actinomycin D and (F) RNase R. (G) qRT-PCR detected the levels of circ_0132266 and MTO1 mRNA after reverse transcribed with random primers and oligo (dt)18 primers.
Circ_0132266 was derived from exon2-9 of the MTO1 gene on chromosome 6. The spanning junction primers were designed and expected size of product was amplified and verified by Sanger sequencing (Figure 3D). RNase R and actinomycin D treatment manifested the stability of circ_0132266 (Figure 3E, 3F). qRT-PCR analysis following the use of oligo (dt)18 primers and random primers in reverse transcription illustrated that circ_0132266 was a closed loop structure (Figure 3G). The results suggested the characteristics of circ_0132266 which was downregulated in CLL.
Circ_0132266 was validated to be miR-337-3p sponge
Given to the expression correlation between circ_0132266 and miR-337-3p, we hypothesized that circ_0132266 functioned as miR-337-3p sponge. FISH assay and nuclear and cytoplasmic fraction experiments showed circ_0132266 was abundant in cytoplasm (Figure 4A, 4B). RIP assay showed the enrichment of circ_0132266 by AGO2 antibodies compared with IgG (Figure 4C). To further assure the binding of predicted putative binding sites of circ_0132266 and miR-337-3p (Figure 4D), one of the most potential bindings with highest score was detected through dual luciferase assay. As expected, there was a significant decrease of the luciferase activity mediated by wild-type circ_0132266 sequence when co-transfecting miR-337-3p mimics and Sanger sequencing assured the binding sites of conducted plasmids (Figure 4D, 4E).
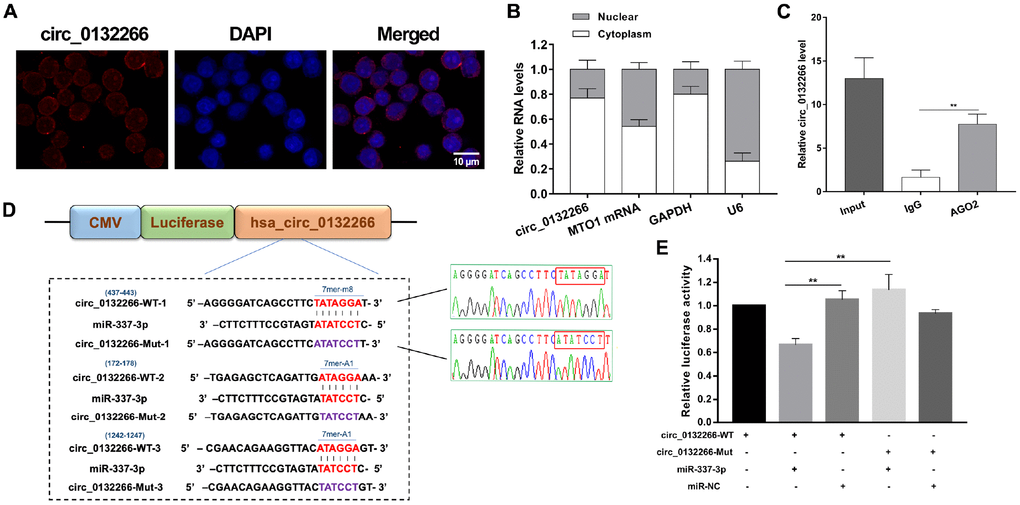
Figure 4. Circ_0132266 serves as a sponge for miR-337-3p. (A, B) FISH and nuclear and cytoplasmic fraction experiments were performed to detect the location of circ_0132266. (C) RIP assay performed with AGO2 antibody to assure the ability of circ_0132266 to serve as miRNA sponge. (D) Schematic showed the predicted binding sites between circ_0132266 and miR-337-3p and Sanger sequencing confirmed the accuracy of conducted vectors which will be used in the dual-luciferase assay. (E) The binding between circ_0132266 and miR-337-3p was verified through dual-luciferase activity assay.
Taken together, our data revealed that circ_0132266 possessed miRNA-binding sites and acted as a sponge of miR-337-3p.
Discussion
Though CLL is the commonest leukemia in western countries and shows an increasing tendency in China, its pathogenesis has not yet been clear. It is therefore in urgent need of exploring the molecular mechanism of CLL.
In the present study, we noted an obviously upregulated miR-337-3p which has been reported to be aberrantly regulated in solid tumors and possessed tumor suppressing activity [11, 13, 22]. It is becoming increasingly evident that circRNAs as emerging ncRNAs have been uncovered preferential roles in mediating miRNAs and contributing to disease progression [23–25]. Therefore, we speculated a modulation mechanism of miR-337-3p mediated by circRNAs. Through a screening system of the genomic region of miR-337-3p, we identified and characterized the existence of a CLL-related circRNA, circ_0132266, and defined its expression profile and functions. Our results revealed a significant negative correlation between the expression levels of miR-337-3p and circ_0132266. And the dual luciferase reporter assay facilitated to support our propose. It is a reasonable speculation that circ_0132266 could accelerated CLL cells apoptosis and impaired proliferation through miR-337-3p/PML axis. Our research model not only opens a new avenue for understanding the regulatory mechanism underlying miR-337-3p but also highlights the possibility of considering circ_0132266 as a potential therapeutic target.
However, limitations cannot not be neglected in our research. Our future work will focus on the following explorations: Firstly, validate the expression of miR-337-3p in a larger cohort of CLL patients. Retrospective analysis is needed to be carried out by following up patients, aimed at finding out the relationship between miR-337-3p and the diagnosis, treatment and prognosis of CLL patients. Considering that our screen system might rule out circRNAs that participated in the process, the next step would be to add the whole transcriptome which will enable the full disclosure of regulatory mechanism of miR-337-3p. A body of evidence indicated that PML is a tumor suppressor and promising therapeutic targets, might be associated with chemo-resistant as well [26–28]. Nextly, gain and loss function assay of PML, a target gene presumably involved in CLL progression, will be performed to further elucidate its functions in order to provide a novel strategy to treat CLL.
Mounting evidence has shown that circRNAs sponging miRNAs might regulate gene transcription and play predominately roles in tumorigenesis. Our findings may enrich the knowledge of circ_0132266 induced mechanism leading to altered miR-337-3p and thus influence the proto-oncogene PML. In summary, the present study defines a novel mechanism of CLL progression and have important therapeutic implications for better treatment in the way of clinical therapy. The prospect of developing miRNA and circRNA-based interventions is undoubtedly an exciting one in the future.
Materials and Methods
Patient samples
For the expression study, samples from thirty treatment-naïve patients diagnosed with CLL and thirty healthy volunteers were obtained after informed consent at the First Affiliated Hospital of Nanjing Medical University, Jiangsu Province Hospital. PBMCs were isolated by density gradient centrifugation with Lymphoprep™ (Stemcell technologies, Canada). The study was approved by the clinical investigation ethical committee from Jiangsu Province Hospital.
Cell culture and transfection
MEC-1 purchased from Cobioer (Nanjing, China) and JVM-3 purchased from X-Y Biotechnology (Shanghai, China) were cultured in RPMI-1640 medium (Gibco®) supplemented with 10% fetal bovine serum (FBS, Gibco, Grand Island, USA), 100 μg/mL penicillin/streptomycin (PS, Gibco, Grand Island, USA) at 37 °C in a humidified atmosphere (5% CO2). Lipofectamine Stem reagent (Invitrogen, Carlsbad, USA) according to manufacturer’s instructions was used for cell transfection. HEK-293T obtained from X-Y Biotechnology (Shanghai, China) was maintained at 37°C with 5% CO2 in DMEM Medium, supplemented with PS, 10% FBS and was transfected using Lipofectamine 2000 (Invitrogen, Carlsbad, USA) according to manufacturer’s protocol.
RNA isolation and quantitative RT-PCR
Total RNA was isolated using the Trizol method (Ambion, USA) and then transcribed reversed by TaqMan reverse transcription kit and amplified using the SYBR Green Master Mix (Applied Biosystems). The primers of mRNA and circRNAs were listed in Supplementary Table 1. miRNA was reversed and amplified by miR 1st strand cDNA Synthesis Kit and miRNA Universal SYBR qPCR Master Mix (Vazyme, China), using specific miR-377 primers GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACGAAGAA (RT) and CGCGCTCCTATATGATGCCT (forward). The U6 small nuclear RNA primer: 5′-CTCGCTTCGGCAGCACA-3′ (forward) and 5′-AACGCTTCACGAATTTGCGT-3′ (reverse) were used as an internal control.
Cell counting kit-8 (CCK8) assay
Cell Counting Kit-8 (Beyotime, Shanghai, China) was used to detect cell proliferative ability. Cells were plated on 96-well plates and 10 μl/ well CCK8 solution were added at pointed time. Then the absorbance at 450 nm was measured after 2 h incubation.
Cell apoptosis assay
Cells were collected and stained by propidium iodine/Annexin V-FITC staining (BD Biosciences) and then detected by flow cytometry (FCM) to analyze the apoptosis. For each cell line triplicates were performed.
Immunofluorescence (IF) and RNA fluorescence in situ hybridization (FISH)
MEC-1 and JVM-3 were plated on slides in 48-well plates and incubated at 60°C for IF and RNA-FISH analysis. After fixation and permeabilization, cells were incubated overnight at 4°C with primary diluted antibody anti-Ki67 (#ab15580, Abcam). The secondary antibody Alexa Fluor® 594 - Conjugated Goat anti-Rabbit IgG (H+L) was purchased from ZSGB-BIO, China. Fluorescence were observed using Axio Scope.A1 (Zeiss). FISH assay was performed using Fluorescent In Situ Hybridization Kit (RiboBio, Guangzhou, China) according to the manufacturer’s guidelines. And Cy3-labeled probes from RiboBio, Guangzhou, China were measured by visualized with a confocal microscopy.
Western blotting
Western blotting was performed using anti-Bax (# ab32503, Abcam), anti-Bcl-2 (# ab32124, Abcam), anti-PML (# ab179466, Abcam), anti-WDR26 (# ab 85961, Abcam), anti-PAPOLB (# ab163420, Abcam), and anti-GAPDH (# ab8245, Abcam) antibodies. Secondary antibodies used in the experiments were from Santa Cruz (Santa Cruz, CA, USA). And the blots were visualized using an enhanced chemiluminescence kit (Pierce, Waltham, USA).
RNA immunoprecipitation (RIP)
RIP experiments were performed using the Magna RIP RNA-Binding Protein Immunoprecipitation Kit (Millipore, Bedford, MA) according to the manufacturer’s instructions. The probes were designed and synthesized by Sangon Biotech. The abundance of circ_0132266 and miR-337-3p was detected by qRT–PCR assay.
Dual luciferase reporter assay
The interactions between circ_0132266, miR-337-3p and PML were measured using pMIR-REPORTTM system (Applied Biosystems, USA). The circ_0132266 and mRNA 3′ UTR sequences containing wild-type (WT) or mutant (Mut) miR-337-3p binding sites were synthesized and respectively inserted into pMIR-REPORT vectors and then co-transfected with miR-337-3p mimics or control mimics using Lipofectamine 2000. Dual luciferase activities were tested by the dual-luciferase reporter assay kit (Promega, Madison, WI, USA) 48 h later.
Statistical analysis
GraphPad Prism 7 (GraphPad Software Inc., La Jolla, CA) was used to analyze experimental data. The expression RNA levels were performed using student’s t test and a p value less than 0.05 was considered to be statistically significant.
Conclusions
In summary, the findings of the present study define a novel mechanism of CLL deregulation and describe a new role of miR-337-3p as a carcinogen in CLL. And circ_0132266 which was demonstrated to be downregulated in CLL could sponge miR-337-3p and be involved in CLL progression through miR-337-3p/PML axis. Accordingly, our findings highlighted that circ_0132266 might have potential values for understanding the complicated molecular mechanisms of miR-337-3p in CLL and providing a new strategy for future CLL therapy.
Supplementary Materials
Author Contributions
Conceptualization: JYL and HJ; methodology: WW, ZJW and HJ; investigation: WW, ZJW, YX, SCQ; clinical data acquisition: ZJW, YL, JZW, JHL and LW; bioinformatical analyses: HYZ and LF; writing-original draft: WW, ZJW; writing-review and editing: JYL and HJ; visualization: JXF and WX; supervision and funding acquisition: JYL, HJ and WW and YX.
Conflicts of Interest
The authors declare no conflicts of interest.
Funding
This study was supported by National Natural Science Foundation of China (Grant no. 81700155, 81600162, 81370657, 81470328, 81600130, 81770166, 81720108002, 81700193), Jiangsu Province’s Medical Elite Programme (ZDRCA2016022), Jiangsu Provincial Special Program of Medical Science (BE2017751) and National Science and Technology Major Project (2018ZX09734-007).
References
- 1. Hallek M. Chronic lymphocytic leukemia: 2017 update on diagnosis, risk stratification, and treatment. Am J Hematol. 2017; 92:946–65. https://doi.org/10.1002/ajh.24826 [PubMed]
- 2. Chiorazzi N, Rai KR, Ferrarini M. Chronic lymphocytic leukemia. N Engl J Med. 2005; 352:804–15. https://doi.org/10.1056/NEJMra041720 [PubMed]
- 3. Ambros V. The functions of animal microRNAs. Nature. 2004; 431:350–55. https://doi.org/10.1038/nature02871 [PubMed]
- 4. Halvorsen AR, Helland Å, Gromov P, Wielenga VT, Talman MM, Brunner N, Sandhu V, Børresen-Dale AL, Gromova I, Haakensen VD. Profiling of microRNAs in tumor interstitial fluid of breast tumors - a novel resource to identify biomarkers for prognostic classification and detection of cancer. Mol Oncol. 2017; 11:220–34. https://doi.org/10.1002/1878-0261.12025 [PubMed]
- 5. Powers JT, Tsanov KM, Pearson DS, Roels F, Spina CS, Ebright R, Seligson M, de Soysa Y, Cahan P, Theißen J, Tu HC, Han A, Kurek KC, et al. Multiple mechanisms disrupt the let-7 microRNA family in neuroblastoma. Nature. 2016; 535:246–51. https://doi.org/10.1038/nature18632 [PubMed]
- 6. Musilova K, Mraz M. MicroRNAs in B-cell lymphomas: how a complex biology gets more complex. Leukemia. 2015; 29:1004–17. https://doi.org/10.1038/leu.2014.351 [PubMed]
- 7. Mraz M, Kipps TJ. MicroRNAs and B cell receptor signaling in chronic lymphocytic leukemia. Leuk Lymphoma. 2013; 54:1836–39. https://doi.org/10.3109/10428194.2013.796055 [PubMed]
- 8. Papageorgiou SG, Diamantopoulos MA, Kontos CK, Bouchla A, Vasilatou D, Bazani E, Scorilas A, Pappa V. MicroRNA-92a-3p overexpression in peripheral blood mononuclear cells is an independent predictor of prolonged overall survival of patients with chronic lymphocytic leukemia. Leuk Lymphoma. 2019; 60:658–667. https://doi.org/10.1080/10428194.2018.1461861 [PubMed]
- 9. Balatti V, Tomasello L, Rassenti LZ, Veneziano D, Nigita G, Wang HY, Thorson JA, Kipps TJ, Pekarsky Y, Croce CM. miR-125a and miR-34a expression predicts Richter syndrome in chronic lymphocytic leukemia patients. Blood. 2018; 132:2179–82. https://doi.org/10.1182/blood-2018-04-845115 [PubMed]
- 10. Cerna K, Oppelt J, Chochola V, Musilova K, Seda V, Pavlasova G, Radova L, Arigoni M, Calogero RA, Benes V, Trbusek M, Brychtova Y, Doubek M, et al. MicroRNA miR-34a downregulates FOXP1 during DNA damage response to limit BCR signalling in chronic lymphocytic leukaemia B cells. Leukemia. 2019; 33:403–414. https://doi.org/10.1038/s41375-018-0230-x [PubMed]
- 11. Zhuang Q, Shen J, Chen Z, Zhang M, Fan M, Xue D, Lu H, Xu R, He X, Hou J. MiR-337-3p suppresses the proliferation and metastasis of clear cell renal cell carcinoma cells via modulating Capn4. Cancer Biomark. 2018; 23:515–25. https://doi.org/10.3233/CBM-181645 [PubMed]
- 12. Wang Z, Wang J, Yang Y, Hao B, Wang R, Li Y, Wu Q. Loss of has-miR-337-3p expression is associated with lymph node metastasis of human gastric cancer. J Exp Clin Cancer Res. 2013; 32:76. https://doi.org/10.1186/1756-9966-32-76 [PubMed]
- 13. Zuo XL, Chen ZQ, Wang JF, Wang JG, Liang LH, Cai J. miR-337-3p suppresses the proliferation and invasion of hepatocellular carcinoma cells through targeting JAK2. Am J Cancer Res. 2018; 8:662–74. [PubMed]
- 14. Demarez C, Gérard C, Cordi S, Poncy A, Achouri Y, Dauguet N, Rosa DA, Gunning PT, Manfroid I, Lemaigre FP. MicroRNA-337-3p controls hepatobiliary gene expression and transcriptional dynamics during hepatic cell differentiation. Hepatology. 2018; 67:313–27. https://doi.org/10.1002/hep.29475 [PubMed]
- 15. Piwecka M, Glažar P, Hernandez-Miranda LR, Memczak S, Wolf SA, Rybak-Wolf A, Filipchyk A, Klironomos F, Cerda Jara CA, Fenske P, Trimbuch T, Zywitza V, Plass M, et al. Loss of a mammalian circular RNA locus causes miRNA deregulation and affects brain function. Science. 2017; 357:357. https://doi.org/10.1126/science.aam8526 [PubMed]
- 16. Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK, Kjems J. Natural RNA circles function as efficient microRNA sponges. Nature. 2013; 495:384–88. https://doi.org/10.1038/nature11993 [PubMed]
- 17. de Thé H, Chomienne C, Lanotte M, Degos L, Dejean A. The t(15;17) translocation of acute promyelocytic leukaemia fuses the retinoic acid receptor alpha gene to a novel transcribed locus. Nature. 1990; 347:558–61. https://doi.org/10.1038/347558a0 [PubMed]
- 18. Trotman LC, Alimonti A, Scaglioni PP, Koutcher JA, Cordon-Cardo C, Pandolfi PP. Identification of a tumour suppressor network opposing nuclear Akt function. Nature. 2006; 441:523–27. https://doi.org/10.1038/nature04809 [PubMed]
- 19. Wang ZG, Ruggero D, Ronchetti S, Zhong S, Gaboli M, Rivi R, Pandolfi PP. PML is essential for multiple apoptotic pathways. Nat Genet. 1998; 20:266–72. https://doi.org/10.1038/3073 [PubMed]
- 20. Gurrieri C, Capodieci P, Bernardi R, Scaglioni PP, Nafa K, Rush LJ, Verbel DA, Cordon-Cardo C, Pandolfi PP. Loss of the tumor suppressor PML in human cancers of multiple histologic origins. J Natl Cancer Inst. 2004; 96:269–79. https://doi.org/10.1093/jnci/djh043 [PubMed]
- 21. Wang K, Long B, Liu F, Wang JX, Liu CY, Zhao B, Zhou LY, Sun T, Wang M, Yu T, Gong Y, Liu J, Dong YH, et al. A circular RNA protects the heart from pathological hypertrophy and heart failure by targeting miR-223. Eur Heart J. 2016; 37:2602–11. https://doi.org/10.1093/eurheartj/ehv713 [PubMed]
- 22. Huang Z, Zhang N, Ma W, Dai X, Liu J. MiR-337-3p promotes chondrocytes proliferation and inhibits apoptosis by regulating PTEN/AKT axis in osteoarthritis. Biomed Pharmacother. 2017; 95:1194–200. https://doi.org/10.1016/j.biopha.2017.09.016 [PubMed]
- 23. Jost I, Shalamova LA, Gerresheim GK, Niepmann M, Bindereif A. Functional sequestration of microRNA-122 from Hepatitis C Virus by circular RNA sponges. RNA Biol. 2018; 15:1032–1039. https://doi.org/10.1080/15476286.2018.1435248. [PubMed]
- 24. Zheng Q, Bao C, Guo W, Li S, Chen J, Chen B, Luo Y, Lyu D, Li Y, Shi G, Liang L, Gu J, He X, Huang S. Circular RNA profiling reveals an abundant circHIPK3 that regulates cell growth by sponging multiple miRNAs. Nat Commun. 2016; 7:11215. https://doi.org/10.1038/ncomms11215 [PubMed]
- 25. Memczak S, Jens M, Elefsinioti A, Torti F, Krueger J, Rybak A, Maier L, Mackowiak SD, Gregersen LH, Munschauer M, Loewer A, Ziebold U, Landthaler M, et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013; 495:333–38. https://doi.org/10.1038/nature11928 [PubMed]
- 26. Ponente M, Campanini L, Cuttano R, Piunti A, Delledonne GA, Coltella N, Valsecchi R, Villa A, Cavallaro U, Pattini L, Doglioni C, Bernardi R. PML promotes metastasis of triple-negative breast cancer through transcriptional regulation of HIF1A target genes. JCI Insight. 2017; 2:e87380. https://doi.org/10.1172/jci.insight.87380 [PubMed]
- 27. Mazza M, Pelicci PG. Is PML a Tumor Suppressor? Front Oncol. 2013; 3:174. https://doi.org/10.3389/fonc.2013.00174 [PubMed]
- 28. Ito K, Bernardi R, Morotti A, Matsuoka S, Saglio G, Ikeda Y, Rosenblatt J, Avigan DE, Teruya-Feldstein J, Pandolfi PP. PML targeting eradicates quiescent leukaemia-initiating cells. Nature. 2008; 453:1072–78. https://doi.org/10.1038/nature07016 [PubMed]