Introduction
Alzheimer's disease (AD) is a progressive neurodegenerative disorder that affects a growing proportion of the aging population. Patients with AD manifest with gradual decline of cognitive and functional abilities and shortened lifespan [1]. Due to the complex and multifactorial nature of AD, the etiology of which remains poorly understood, effective interventional means for prevention and treatment are lacking [2, 3]. There is growing recognition that the pathological mechanisms underlying AD do not only involve the aggregation of abnormal proteins, such as amyloid beta peptide (Aβ) and tau, but also include dysfunction of immune responses in the brain [4]. Since there is a lack of adaptive immune system in human brains under normal circumstances, impaired innate immune function has been proposed to be a key mechanism in the initiation and progression of AD [5]. Although the innate immune system has been considered a potential therapeutic target and has drawn substantial attention in biological and pharmaceutical studies, it is still disputed whether innate immunity is increased or decreased in AD [6, 7].
Animal models are indispensable tools to investigate pathological mechanisms and intervention strategies for AD. Over the past few decades, many studies have been conducted in Drosophila melanogaster to gain insight into the pathophysiological processes underpinning AD, identify potentially important genes and biomarkers, and screen new drug candidates. Besides the well-known advantages of using Drosophila as a model species [8], the host defense of the fruit fly rests entirely within its complex innate immune system, which makes it a desirable model for research on innate immunity in AD [9].
Aging is generally regarded as the most important risk factor for AD. However, the effects of aging on innate immunity in Drosophila have not been fully elucidated. Therefore, we performed a comprehensive data mining of the published expressional profiles [10–23] and experimental study at the transcriptomic level to analyze the expression profiles of innate immunity genes in wild-type (WT) and Aβ transgenic Drosophila model during aging. The transcriptional levels of major differentially expressed genes, Aβ deposition, and neuronal apoptosis in the head of both control and AD flies were also assessed to evaluate the effects of dysregulation of innate immunity on disease progression.
Results
Transcriptomic data mining revealed a general upregulation of antimicrobial peptide (AMP) expression in the head of normal D. melanogaster during aging
To explore the expressive regularity of the innate immune system from a broad range of data, a total of 18 eligible experiments from GEO database comprising data of 52 young, 57 middle-aged, and 75 old healthy Drosophila head samples were obtained through data retrieval and selection of Drosophila head samples (Table 1). All expression profiles of innate immune genes were converted into occurrence of high expression. RD and OR values, which represented the difference in expression of innate immune genes, were calculated subsequently.
Table 1. Eligible datasets mined from GEO database.
| GEO accession | Sample | Age | Age group |
| GSE122470 | GSM3466957, GSM3466958, GSM3466959 | 3 | Young |
| GSM3466960, GSM3466961, GSM3466962 | 15 | Mid-aged | |
| GSM3466963, GSM3466964, GSM3466965 | 30 | Old | |
| GSM3466966, GSM3466967, GSM3466968 | 45 | ||
| GSE75216 [10] | GSM1945845, GSM1945846, GSM1945847, GSM1945855, GSM1945856 | 7 | Young |
| GSM1945843, GSM1945844, GSM1945853, GSM1945854 | 22 | Mid-aged | |
| GSE64108 | GSM1564407, GSM1564408, GSM1564409, GSM1564410 | 21 | Mid-aged |
| GSM1564415, GSM1564417, GSM1564419, GSM1564421 | 35 | Old | |
| GSM1564431, GSM1564432, GSM1564433, GSM1564434 | 49 | Old | |
| GSE38998 [11] | GSM1186462, GSM1186463 | 3 | Young |
| GSM953478, GSM953479 | 10 | Mid-aged | |
| GSE81100 [12] | GSM2143625, GSM2143626, GSM2143627, GSM2143628, GSM2143629, GSM2143630, GSM2143631, GSM2143632, GSM2143633, GSM2143634, GSM2143635, GSM2143636 | 5 | Young |
| GSM2143637, GSM2143638, GSM2143639, GSM2143640, GSM2143641, GSM2143642, GSM2143643, GSM2143644, GSM2143645, GSM2143646, GSM2143647, GSM2143648 | 55 | Old | |
| GSE110135 [13] | GSM2978238, GSM2978239, GSM2978240 | 3 | Young |
| GSM2978241, GSM2978242, GSM2978243 | 20 | Mid-aged | |
| GSE6430 | GSM12770 | 3 | Young |
| GSM12772 | 47 | Old | |
| GSE97493 [14] | GSM2570129, GSM2570130, GSM2570131, GSM2570132, GSM2570133 | 3 | Young |
| GSM2570134, GSM2570135, GSM2570136, GSM2570137, GSM2570138, GSM2570159, GSM2570160, GSM2570161, GSM2570162, GSM2570163, GSM2570164 | 10 | Mid-aged | |
| GSM2570149, GSM2570150, GSM2570151, GSM2570152, GSM2570153 | 30 | Old | |
| GSM2570154, GSM2570155, GSM2570156, GSM2570157, GSM2570158 | 45 | Old | |
| GSE98554 [15] | GSM2599109, GSM2599110, GSM2599111 | 2 | Young |
| GSM2599112, GSM2599113, GSM2599114 | 25 | Mid-aged | |
| GSE48681 [16] | GSM1183416, GSM1183417, GSM1183418, GSM1183419 | 3 | Young |
| GSM1183420, GSM1183421, GSM1183422, GSM1183423 | 10 | Mid-aged | |
| GSM1183424, GSM1183425, GSM1183426 | 20 | ||
| GSM1183427, GSM1183428, GSM1183429, GSM1183430 | 56 | Old | |
| GSM1183435, GSM1183436, GSM1183437, GSM1183438 | 68 | ||
| GSE25009 [17] | GSM614349, GSM614350, GSM614351 | 3 | Young |
| GSM614352, GSM614353, GSM614354 | 30 | Old | |
| GSM614355, GSM614356, GSM614357 | 60 | ||
| GSE26246 [18] | GSM644354, GSM644355, GSM644356 | 0 | Young |
| GSM644357, GSM644358, GSM644359 | 2 | ||
| GSM644360, GSM644361, GSM644362 | 14 | Mid-aged | |
| GSE26726 [19] | GSM658027, GSM658028, GSM658029, GSM658036, GSM658037, GSM658038, GSM658060, GSM658061, GSM658062 | 10 | Mid-aged |
| GSM658042, GSM658043, GSM658044, GSM658051, GSM658052, GSM658053, GSM658066, GSM658067, GSM658068 | 40 | Old | |
| GSE22440 | GSM557543, GSM557544, GSM557545 | 10 | Mid-aged |
| GSM557546, GSM557547, GSM557548 | 40 | Old | |
| GSE21182 [20] | GSM530094 | 1 | Young |
| GSM530096 | 40 | Old | |
| GSE6314 [21] | GSM132562, GSM132563 | 15 | Mid-aged |
| GSM132564, GSM132565 | 20 | ||
| GSM132566, GSM132567 | 30 | Old | |
| GSM132568, GSM132569 | 45 | ||
| GSM132570, GSM132571 | 60 | ||
| GSE826 [22] | GSM12770 | 3 | Young |
| GSM12772 | 47 | Old | |
| GSE37148 [23] | GSM912518, GSM912519, GSM912520 | 5 | Young |
| GSM912521, GSM912522, GSM912523 | 45 | Old |
Among all categories of innate immunity relevant genes (Figure 1A–1C), only AMP genes exhibited a marked increase in gene expression with age. However, the degree of expression of other inducible and constitutive effector molecules in the host defense system of Drosophila, such as C-type lectins or lysozymes; pattern recognition receptors (PRRs), including peptidoglycan recognition proteins (PGRPs) and gram-negative binding proteins (GNBPs); major molecules involved in Toll, immune deficiency (IMD), and Janus kinase - signal transducer and activator of transcription (JAK-STAT) pathways, essentially remained unchanged. Meta-analysis (Figure 1D–1F, Supplementary Figure 1) further demonstrated that with the exception of drosomycin (Drs), among 14 major AMP genes, including attacin (Att) A, B, C and D; cecropin (Cec) A1, A2, B and C; diptericin (Dpt) A and B; Drs; defensin (Def); drosocin (Dro); and metchnikowin (Mtk); there was overexpression of all other genes between at least one younger group with that of at least one older group. The data mining results suggested that the upregulation of AMPs in the head of healthy aging Drosophila could be one of the most important changes in the innate immune system with age.
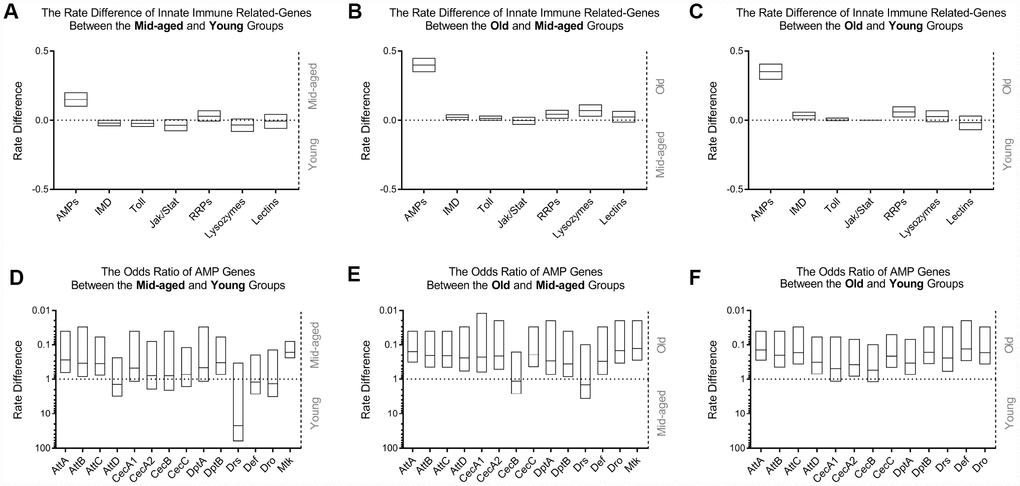
Figure 1. Comparison of innate immunity gene expression variances in WT flies with age among GEO datasets. The occurrence of high expression was introduced as a statistical parameter to compare the relative expression quantity of immune-related genes across age groups. The upper figures exhibit the rate differences among different classes of genes associated with innate immunity between the middle-aged and young (A), old and middle-aged (B), and old and young (C) groups, clearly demonstrating that AMPs had the most significant differential expression. The expression levels of other gene clusters were generally unchanged. Transcriptional differences in AMP genes were subsequently compared using meta-analysis. The odds ratio between the middle-aged and young (D), old and middle-aged (E), and old and young (F) groups, are shown in the lower graphs, indicating an increased expression of AMPs in the head of WT Drosophila during aging.
Age-associated overexpression of AMP genes in normal Drosophila was observed in RNA-Seq
To validate the gene expression variation of the innate immune system with age, whole transcriptome profiles of normal Drosophila heads at 3, 10, 20, and 30 days post eclosion were analyzed by high throughput RNA sequencing. The raw data had been deposited to the GEO database with an accession number of GSE109489. The relative quantity of gene expression and the significance value between groups were calculated by DESeq2 software. The results revealed that 12 out of 14 AMP genes, including AttA, AttB, AttC, AttD, CecA2, CecC, DptA, DptB, Drs, Def, Dro, and Mtk, had upregulated expression in at least one older group compared to that of at least one younger group (Figure 2B). Despite of slight variations on individual genes, the expression levels of these AMP genes generally presented a gradual tendency to increase with age, consistent with results from the big data analysis (Figure 2A). Lysozyme (Lys) S, and three PGRP genes, including -LC, -SA, and -SD, were overexpressed in at least one pair of age groups. However, there was no significantly different expression among other AMP-related genes.
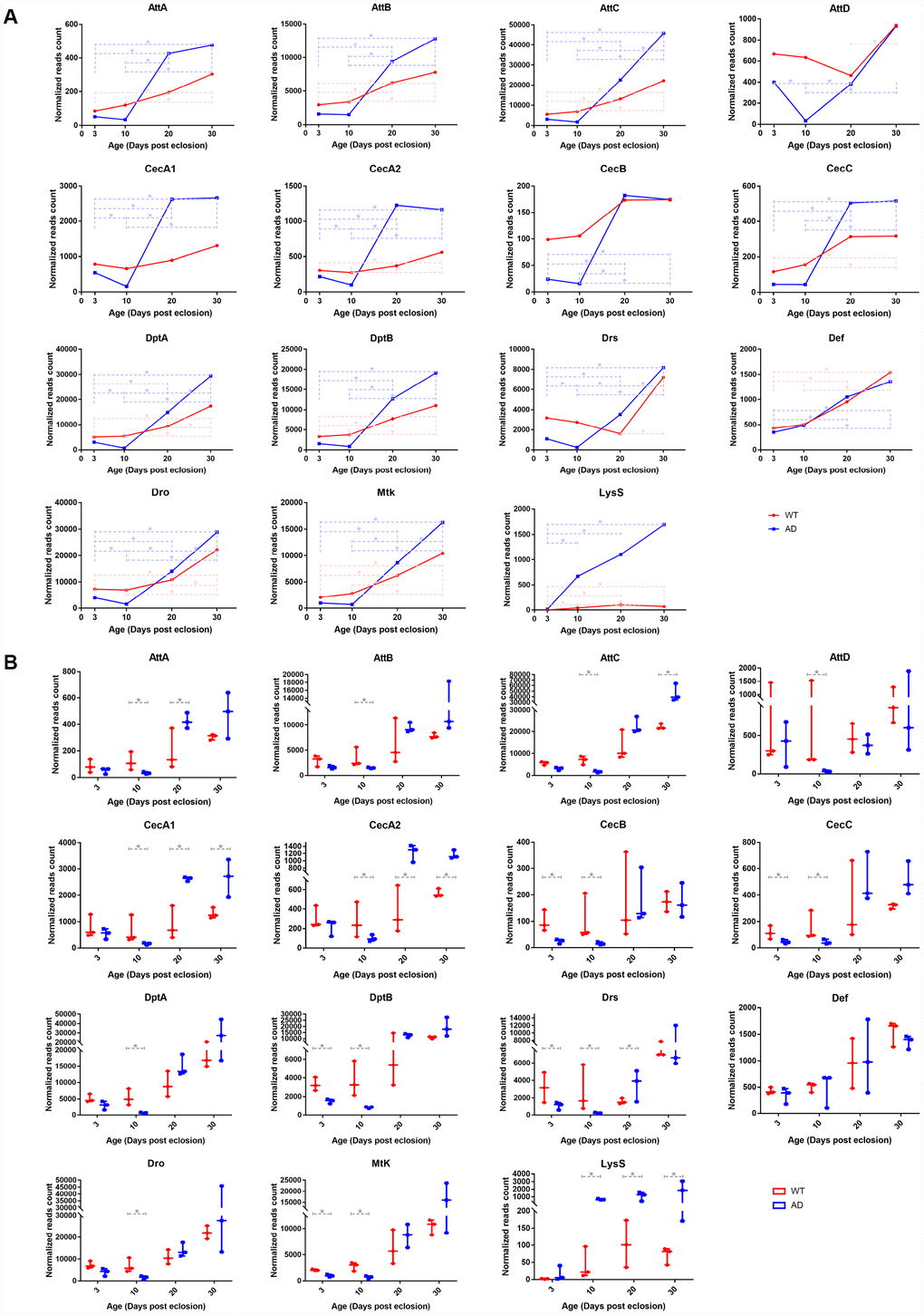
Figure 2. RNA-seq analysis of differentially expressed genes associated with the innate immune system in control and AD Drosophila model. There were significant differences in transcriptional levels of AMPs and LysS between the two groups. The line chart (A) illustrates different expression patterns of differentially expressed immune-related genes between WT (red, round dots) and AD (blue, square dots) groups during aging. The box plot (B) presents differences in expression therein between the normal (red, left) and disease (blue, right) groups at each time point (3-, 10-, 20-, 30-days post eclosion).
High throughput sequencing indicated differential AMP expression pattern in normal and AD flies
Transcriptomic profiling of the Drosophila model of AD at the time points corresponding to those of the WT group was performed using RNA-Seq. Raw transcriptomic data was deposited to the same database with an identical accession number. Time series analysis illustrated that the expression of most AMPs declined to the lowest on day 10, elevated sharply on day 20, and continuously increased or remained unchanged on day 30. The expression levels of AMP genes displayed a rough trend of initial decrease followed by an increase, which was distinct to that of healthy controls (Figure 2A). The most significant differences in expression between WT and AD groups appeared on day 10, when 13 out of 14 AMPs, including AttA, AttB, AttC, AttD, CecA1, CecA2, CecB, CecC, DptA, DptB, Drs, Def, Dro, and Mtk, showed a remarkable decline in the AD group, with log2 fold changes of -1.816, -1.182, -2.011, -4.250, -2.065, -1.455, -2.735, -1.813, -3.011, -2.184, -3.525, -2.173, and -1.975, respectively (Figure 2B). mRNA levels of LysS, PGRP-LC, -SA, and -SD genes presented a rising trend with age in the AD group, but there were no significant differences among PGRP genes when compared with those of the WT group. However, the expression of LysS exhibited a comparable increase in Drosophila with AD. The log2-fold change values of LysS between the WT and AD groups at day 3, 10, 20, and 30 were 2.891, 3.929, 3.412, and 4.576, respectively (Figure 2).
Validation of AMP gene expression profiles by quantitative real-time PCR (qPCR)
Transcript levels of key differentially expressed innate immunity genes were investigated using RNA-Seq, including 14 AMP genes and LysS, and were validated using qPCR assays. The qPCR result generally recapitulated RNA-Seq data. Expression trends of the aforementioned genes in different age groups were generally consistent with those revealed by transcriptomic analysis (Figure 3A). Quantitative analysis confirmed that expression levels of nearly all AMP genes declined to some extent in the AD group at day 3 and 10. Among these, six AMPs, including AttC, CecB, CecC, DptB, Drs, and Dro, had the most significant differential expression, with log2-fold changes of -1.344, -2.097, -2.750, -2.398, -1.973, and -1.791, respectively. After day 10, AMP expression levels in the AD group rapidly increased. The expression of most AMP genes remained at similar levels to those of the control group, of which CecA and AttA were found elevated on day 20 and 30 with log2-fold changes of 1.250 and 1.255, respectively (Figure 3B). There was a continuously increasing trend in the mRNA level of LysS from day 3 to 30, which was markedly increased relative to control levels, with log2-fold change values of 1.413, 3.636, 3.555, and 3.331, respectively (Figure 3).
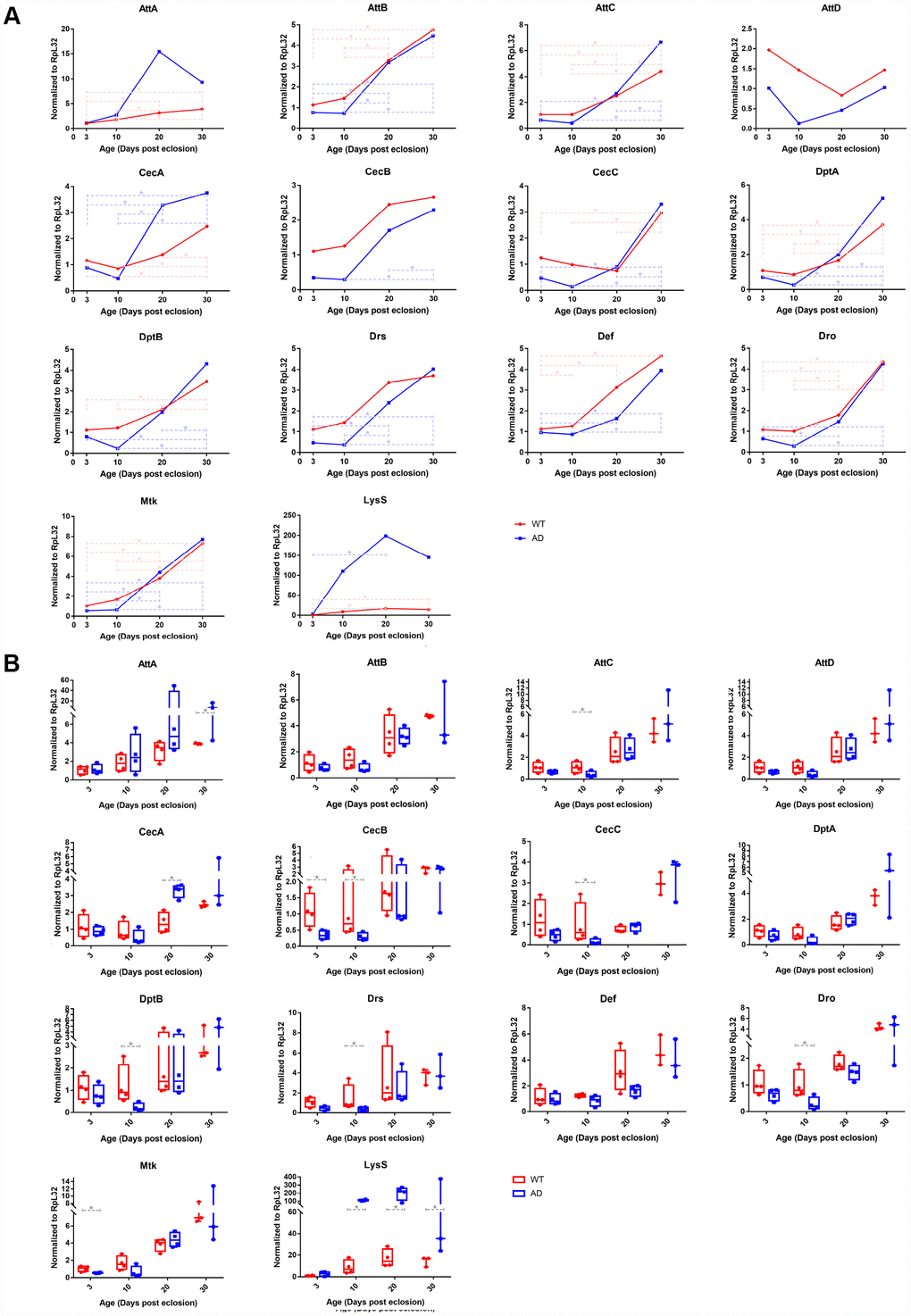
Figure 3. Quantitative PCR validation of differentially expressed immune-related genes in control and Aβ transgenic flies. The results confirmed age-associated alterations in expression trends and transcriptional regulatory levels among the AMP and LysS genes between healthy control and disease model flies. The line chart (A) displays the time series (3-, 10-, 20-, and 30-days post eclosion) gene expression in the head tissue of WT (red, round dots) and AD (blue, square dots) flies. The box plot (B) exhibits the comparison of mRNA levels between normal (red, left) and disease (blue, right) model flies among the age groups.
Discussion
Drosophila has proven to be an excellent model system for studies of aging and age-related neurodegenerative diseases, not only due to its high genetic conservation to humans, but also due to its short lifespan and simple central nervous system (CNS) anatomy and physiology. In humans, aging and neurodegenerative diseases are accompanied with altered immunity [24]. Although there is a lack of adaptive immunity in Drosophila, immune defense in the human brain under normal circumstances also relies entirely on the innate immune system [5]. The innate immune systems of humans and Drosophila are highly analogous: they are activated by PGRPs and GNBPs; transcriptionally regulated through Toll, IMD, and JAK-STAT signaling pathways; and function by expressing effector molecules through NF-κB transcription factors [25, 26].
Innate immunity is the first line defender of the host based on efficient response mechanisms and potent immune molecules that are expressed, such as AMPs, lysozymes, and lectins. The innate immune system has been proposed to be upregulated during aging to withstand growing susceptibility to infections [27]. Conversely, overexpression of antimicrobial peptide genes could contribute to age-related diseases through cytotoxic effects [28]. Although the interplay between immunity and aging in Drosophila is well established, despite that the mechanism that underlie such reciprocity remain unknown, there is a lack of literature specific to the head tissue. In this study, gradually increasing levels of AMP expression in the head of healthy aging Drosophila were observed by mining transcriptome sequencing data from the GEO database, which were subsequently verified by RNA-seq and qPCR analyses, consistent with the available literature [29–31]. However, there were no significant differences in expression of major genes of Toll, IMD, or JAK-STAT signaling pathways, indicating that these classical immune relevant pathways may not be involved in the overexpression of AMP genes.
Prolonged inflammation is associated with the progression of AD in humans, which results from the accumulation of aberrant Aβ aggregates. Activated immunity in the CNS has been suggested to be responsible for the onset of neurodegeneration in Drosophila [32]. Experimental evidence has illustrated that overexpression of a particular AMP gene in the CNS of WT Drosophila was sufficient to induce pathogenesis of neurodegeneration and shortened lifespan. The pathological mechanism was attributed to the cytotoxicity of AMPs on neurons and glia in the CNS [28, 33]. Neuron loss, behavioral impairment, and shortened lifespan are the most characteristic pathological changes of Drosophila with AD. Therefore, the expression of AMP genes were deduced to be upregulated in the AD Drosophila model. In this study, LysS, which was the only differentially expressed disease-associated immune response gene other than AMPs, was strongly overexpressed in the AD group. Overexpression of lysozymes inhibits Aβ aggregation and cell apoptosis, making it a potential target for the diagnosis and treatment of AD [34]. Interestingly, to our surprise, the expression of AMPs were downregulated in the early stage of adult AD flies based on our results, especially on day 10, indicating that overexpression of AMPs is not an indispensable condition for the pathogenesis of AD. In contrast, the downregulation of AMPs occurred prior to commencement of substantial Aβ deposition and neuronal apoptosis on day 20, suggesting that the suppressed expression of AMPs may initiate the development of AD and may be a novel hallmark for early diagnosis of AD. Nevertheless, the expression levels of most AMP genes were generally increased compared to those of controls at the later stage of AD. Under such circumstances, AMPs may exert their cytotoxic effects on CNS cells and contribute to disease progression.
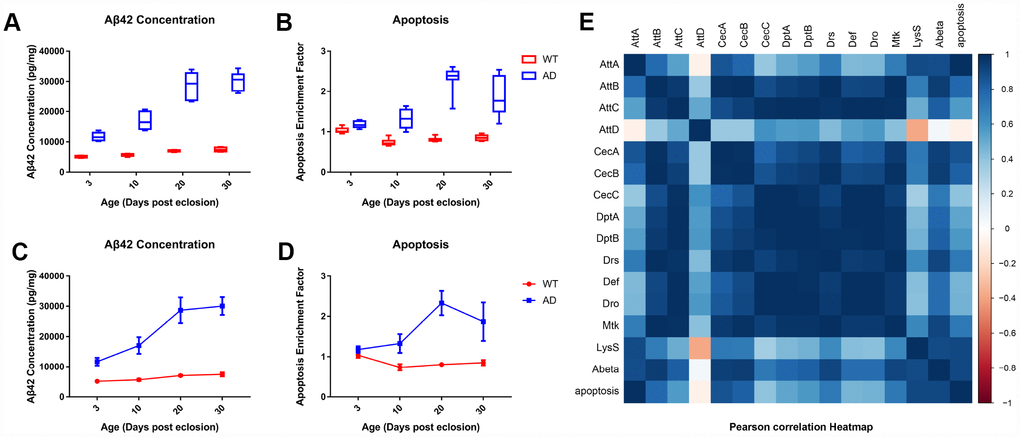
Available transcriptomic profiles from the GEO database include three studies (Supplementary Figure 2) on Drosophila models of AD (GSE48681), amyotrophic lateral sclerosis (ALS, GSE37148) and Parkinson’s disease (PD, GSE74247) [16, 23, 35]. The experiment revealed a generally decreased expression of AMPs in the neurodegenerative disease group compared with the relative controls at early ages, although only a few of the differences among individual AMPs were statistically significant. Low levels of AMP gene expression may be a common phenomenon in the early stages of neurodegenerative diseases. However, the time points set after 20 days in the AD study were not matched by age between the disease and control groups, so it is difficult to determine the variation in AMP expression in Drosophila with advanced AD, although generally elevated AMP mRNA levels could be observed at the late stage of ALS. A microarray assay of AD also demonstrated that AMP genes were downregulated when the mortality of Drosophila with AD started to increase. Our study further illustrated that there was no significant association between AMP expression and neuronal apoptosis, according to correlation coefficient analysis. Therefore, decreased AMP expression is unlikely to be directly responsible for neuronal loss or mortality in Drosophila models of AD. Notably, correlation analysis pointed to trends towards positive correlations between expression of several antimicrobials and neurotoxic Aβ42 concentration. These findings suggest that downregulation of AMP expression presumptively lead to Aβ deposition, resulting in neuronal apoptosis and mortality.
AMP gene expression relies predominantly on the Toll and IMD signaling pathways. The JAK-STAT pathway is also involved in the regulation of innate immune responses [36]. Activation of Toll and IMD pathways leads to AMP expression, while absence of both pathways results in loss of AMP production [37]. It is noteworthy that Aβ is increasingly being recognized as an AMP that protects the host against pathogenic microorganism infection [38]. Therefore, downregulated expression of Aβ peptide could be mediated by negative feedback mechanisms. The expression of Aβ in the transgenic AD Drosophila model that was used in this study was transcriptionally regulated by the GAL4 protein, hence a competitive transcription factor binding mechanism could not be responsible for the low expression of AMP genes. Moreover, in this study, increasing PGRP expression with age was detected in both the AD and WT groups. However, no significant differences in expression among the major genes of the Toll, IMD, or JAK-STAT pathways between the disease and control groups were detected in either in silico or experimental analyses; the mechanisms involved in aberrant AMP expression therefore remain unclear.
In conclusions, the innate immune systems of AD and WT Drosophila were systematically analyzed using an integrative strategy of transcriptomics and experimental validation in this study. The degree of Aβ production and neuronal apoptosis was also investigated. The expression of AMPs in the WT group increased gradually with time, but the increasing trend of AMP expression was disrupted in the AD group, which exhibited an initial downward trend followed by an upward trend during aging. The occurrence of disordered AMP expression, massive aberrant Aβ aggregates, and significant neuronal apoptosis appeared in sequential order, and correlation analysis further indicated a possible causal relationship among the variables. In conclusion, our study revealed dysregulation of AMP expression in an AD Drosophila model with age, distinct from normal aging. Disordered AMP expression may contribute to AD progression by inducing Aβ deposition. However, the physiological and pathological mechanisms of aberrant AMP regulation and the effects on AD and healthy aging are yet to be discovered.
Materials and Methods
Analysis of innate immune gene expression in the head of normal D. melanogaster during aging based on data mining strategies
To analyze the gene expression of innate immunity in Drosophila with age, transcriptome profiles, including microarray and high-throughput sequencing data, were retrieved from the GEO database by organism-specific keyword searching using terms of aging, age, and “time course.” Eligible experiments had to include head tissue of normal D. melanogaster, which were reared under routine culture conditions without any stimulation and contain at least two age groups (young: < 10 days; middle-aged: 10-29 days; old: ≥ 30 days) in a single experiment. The gene expression value was extracted from the original published datasets. The log2-fold change of gene expression relative to control was calculated. Values greater than 1 (2 folds) were considered significant. We compared the statistical parameters of differential expression obtained from individual datasets instead of comparing gene expression signals between experiments to avoid the difficulties in comparing data from different conditions. Each sample from an individual dataset was compared with other age groups. If the log2-fold change exceeded the cutoff threshold, it was marked as one occurrence of high expression. The number of occurrences (k) and number of samples in a single experiment (n) were counted separately.
For comparison of a certain class of genes, the frequency of high expression genes (response rate, P) was calculated (P = k / n), and the overall expression differences were presented as rate difference (RD) = response rate of the older group (Po) - response rate of the younger group (Py). For comparison of the expression of an individual gene among different age groups, the occurrence of high expression genes and total events were counted separately. The analysis was performed using Review Manager Version 5.0. The odds ratios (OR) were the principal measurements of the effects and were presented with 95% confidence intervals (CI). Differences with p < 0.05 were considered statistically significant.
Drosophila stocks
The Drosophila model of AD that expresses human Aβ42 peptide in the brain was constructed using a cross of the w1118 genetic background UAS-Aβ42 flies driven by elav-GAL4c155 line, while WT (w1118) flies which also outcrossed with the elav-GAL4c155 line were used as controls [39]. All flies were reared at 23°C and 42% relative humidity, and fed with standard corn meal food under 12 h/12 h light/dark cycles. Head tissues from 3-, 10-, 20, and 30-day-old male WT and AD Drosophila were collected for further experimentation.
RNA isolation and gene expression analysis
An approximate 200 fly heads for each of triplicates for each age group were collected, and total RNAs were extracted with Trizol reagent (Thermo Fisher, USA), from which mRNAs were purified by poly-T oligo-attached magnetic beads. Libraries were constructed for sequencing using a Superscript Double-Stranded cDNA Synthesis kit (Thermo Fisher, USA) according to manufacturer's specifications.
The prepared library was sequenced on the HiSeq X Ten platform (Illumina, USA) based on a 2 × 150 bp paired-end (PE150) sequencing protocol. The raw sequencing data were processed with SOAPnuke 1.5.2 with parameters of -l 15 -q 0.2 -n 0.05 -i, to remove adaptor sequences, poly-N reads, and low quality reads [40]. The obtained clean data were mapped to Drosophila melanogaster whole genome dmel_r6.11 using HISAT 2.0.4 with parameters of --phred64 --sensitive --no-discordant --no-mixed -I 1 -X 1000 [41, 42]. The relative quantities of gene expression were calculated using RSEM software.
The levels of differentially expressed innate immunity genes were further elucidated through standard qPCR experiment subsequently. Specific primers used in qPCR analysis are listed as Supplementary Table 1. The assays were performed in triplicate on ABI ViiA 7 Real-time PCR system (Applied Biosystems, USA) using QuantiTect SYBR Green PCR kit (Qiagen, Germany) as the fluorescent reporter. Relative expression was estimated with ribosomal protein L32 (RpL32) as the reference gene using the 2−∆∆Ct method.
Protein extraction and ELISA assays
Aβ expression in the brains of Drosophila was examined using a High Sensitivity Human Amyloid β42 ELISA kit (Merck Millipore, USA) according to the manufacturer’s instruction. Briefly, 20 fly heads of each of three replicates for each age group were homogenized then diluted in Standard and Sample Diluent followed by centrifugal purification. The supernatant was transferred into a capture-antibody-coated ELISA plate for overnight incubation at 4°C. After careful rinsing with washing solution, the plate was incubated with the biotinylated detection antibody to form an antibody-amyloid-antibody-complex, which was visualized using the streptavidin-HRP method and measured at 450 nm with a microplate reader (Pulang, China).
Neuronal apoptosis in Drosophila was determined with a Cell Death Detection ELISA Plus kit (Roche, Switzerland). In brief, homogenate samples were prepared using the same method described above. The supernatant containing cytoplasmic histone-associated DNA fragments was added to a streptavidin-coated microplate and incubated with a mixture of anti-histone (biotin-labeled) and anti-DNA (peroxidase-conjugated) antibodies that included in the ELISA kit. The DNA-histone-complex was colored with the ABTS Substrate, then measured at 405 nm with a microplate reader (Pulang, China).
Statistical analysis
Transcriptomic analysis for in silico data was conducted using the DESeq2 package of R software [43]. Transcripts with absolute values of log2-fold change greater than 1 and false discovery rates less than 5% were considered to be significantly differently expressed. Statistical analyses for ELISA and qPCR studies were performed using SPSS Statistics 19.0 software and presented as mean ± SD. Dynamic changes in gene expression were statistically analyzed using one-way analysis of variance (ANOVA), with the Bonferroni post-hoc multiple comparison tests. Significance of qPCR validation among age groups was examined using the Mann-Whitney U test. Correlations between differential gene expression, Aβ42 concentration, and neuronal apoptosis were examined using Pearson's correlation analysis. Results were plotted using the corrplot package of R software. A p-value <0.05 was considered statistically significant.
Acknowledgments
The authors would like to thank Bloomington Drosophila Stock Center for providing experimental materials. We thank BGI Shenzhen for the technical support of transcriptomic profiling of drosophila head samples. The authors acknowledge National Natural Science Foundation of China, China Postdoctoral Science Foundation and Ministry of Finance of the People’s Republic of China for financially supporting this work.
Conflicts of Interest
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was funded by the National Natural Science Foundation of China (No. 81803848, 81801635), China Postdoctoral Science Foundation (No. 2017M610812) and the Ministry of Finance of the People’s Republic of China (No. 1981420400011).
References
- 1. Brookmeyer R, Abdalla N. Estimation of lifetime risks of Alzheimer’s disease dementia using biomarkers for preclinical disease. Alzheimers Dement. 2018; 14:981–88. https://doi.org/10.1016/j.jalz.2018.03.005 [PubMed]
- 2. Verheijen J, Sleegers K. Understanding Alzheimer Disease at the Interface between Genetics and Transcriptomics. Trends Genet. 2018; 34:434–47. https://doi.org/10.1016/j.tig.2018.02.007 [PubMed]
- 3. Hsu D, Marshall GA. Primary and Secondary Prevention Trials in Alzheimer Disease: Looking Back, Moving Forward. Curr Alzheimer Res. 2017; 14:426–40. https://doi.org/10.2174/1567205013666160930112125 [PubMed]
- 4. García-González P, Cabral-Miranda F, Hetz C, Osorio F. Interplay Between the Unfolded Protein Response and Immune Function in the Development of Neurodegenerative Diseases. Front Immunol. 2018; 9:2541. https://doi.org/10.3389/fimmu.2018.02541 [PubMed]
- 5. Heneka MT, Golenbock DT, Latz E. Innate immunity in Alzheimer’s disease. Nat Immunol. 2015; 16:229–36. https://doi.org/10.1038/ni.3102 [PubMed]
- 6. Jevtic S, Sengar AS, Salter MW, McLaurin J. The role of the immune system in Alzheimer disease: etiology and treatment. Ageing Res Rev. 2017; 40:84–94. https://doi.org/10.1016/j.arr.2017.08.005 [PubMed]
- 7. Van Eldik LJ, Carrillo MC, Cole PE, Feuerbach D, Greenberg BD, Hendrix JA, Kennedy M, Kozauer N, Margolin RA, Molinuevo JL, Mueller R, Ransohoff RM, Wilcock DM, et al. The roles of inflammation and immune mechanisms in Alzheimer’s disease. Alzheimers Dement (N Y). 2016; 2:99–109. https://doi.org/10.1016/j.trci.2016.05.001 [PubMed]
- 8. Prüßing K, Voigt A, Schulz JB. Drosophila melanogaster as a model organism for Alzheimer’s disease. Mol Neurodegener. 2013; 8:35. https://doi.org/10.1186/1750-1326-8-35 [PubMed]
- 9. Lye SH, Chtarbanova S. Drosophila as a Model to Study Brain Innate Immunity in Health and Disease. Int J Mol Sci. 2018; 19:E3922. https://doi.org/10.3390/ijms19123922 [PubMed]
- 10. Peleg S, Feller C, Forne I, Schiller E, Sévin DC, Schauer T, Regnard C, Straub T, Prestel M, Klima C, Schmitt Nogueira M, Becker L, Klopstock T, et al. Life span extension by targeting a link between metabolism and histone acetylation in Drosophila. EMBO Rep. 2016; 17:455–69. https://doi.org/10.15252/embr.201541132 [PubMed]
- 11. Gupta VK, Scheunemann L, Eisenberg T, Mertel S, Bhukel A, Koemans TS, Kramer JM, Liu KS, Schroeder S, Stunnenberg HG, Sinner F, Magnes C, Pieber TR, et al. Restoring polyamines protects from age-induced memory impairment in an autophagy-dependent manner. Nat Neurosci. 2013; 16:1453–60. https://doi.org/10.1038/nn.3512 [PubMed]
- 12. Kuintzle RC, Chow ES, Westby TN, Gvakharia BO, Giebultowicz JM, Hendrix DA. Circadian deep sequencing reveals stress-response genes that adopt robust rhythmic expression during aging. Nat Commun. 2017; 8:14529. https://doi.org/10.1038/ncomms14529 [PubMed]
- 13. Kennerdell JR, Liu N, Bonini NM. MiR-34 inhibits polycomb repressive complex 2 to modulate chaperone expression and promote healthy brain aging. Nat Commun. 2018; 9:4188. https://doi.org/10.1038/s41467-018-06592-5 [PubMed]
- 14. Spurrier J, Shukla AK, McLinden K, Johnson K, Giniger E. Altered expression of the Cdk5 activator-like protein, Cdk5α, causes neurodegeneration, in part by accelerating the rate of aging. Dis Model Mech. 2018; 11:dmm031161. https://doi.org/10.1242/dmm.031161 [PubMed]
- 15. Sekiya M, Maruko-Otake A, Hearn S, Sakakibara Y, Fujisaki N, Suzuki E, Ando K, Iijima KM. EDEM Function in ERAD Protects against Chronic ER Proteinopathy and Age-Related Physiological Decline in Drosophila. Dev Cell. 2017; 41:652–664.e5. https://doi.org/10.1016/j.devcel.2017.05.019 [PubMed]
- 16. Favrin G, Bean DM, Bilsland E, Boyer H, Fischer BE, Russell S, Crowther DC, Baylis HA, Oliver SG, Giannakou ME. Identification of novel modifiers of Aβ toxicity by transcriptomic analysis in the fruitfly. Sci Rep. 2013; 3:3512. https://doi.org/10.1038/srep03512 [PubMed]
- 17. Liu N, Landreh M, Cao K, Abe M, Hendriks GJ, Kennerdell JR, Zhu Y, Wang LS, Bonini NM. The microRNA miR-34 modulates ageing and neurodegeneration in Drosophila. Nature. 2012; 482:519–23. https://doi.org/10.1038/nature10810 [PubMed]
- 18. Napoletano F, Occhi S, Calamita P, Volpi V, Blanc E, Charroux B, Royet J, Fanto M. Polyglutamine Atrophin provokes neurodegeneration in Drosophila by repressing fat. EMBO J. 2011; 30:945–58. https://doi.org/10.1038/emboj.2011.1 [PubMed]
- 19. Bauer J, Antosh M, Chang C, Schorl C, Kolli S, Neretti N, Helfand SL. Comparative transcriptional profiling identifies takeout as a gene that regulates life span. Aging (Albany NY). 2010; 2:298–310. https://doi.org/10.18632/aging.100146 [PubMed]
- 20. Lee KS, Lee BS, Semnani S, Avanesian A, Um CY, Jeon HJ, Seong KM, Yu K, Min KJ, Jafari M. Curcumin extends life span, improves health span, and modulates the expression of age-associated aging genes in Drosophila melanogaster. Rejuvenation Res. 2010; 13:561–70. https://doi.org/10.1089/rej.2010.1031 [PubMed]
- 21. Zhang W, Edwards A, Fan W, Zhu D, Zhang K. svdPPCS: an effective singular value decomposition-based method for conserved and divergent co-expression gene module identification. BMC Bioinformatics. 2010; 11:338. https://doi.org/10.1186/1471-2105-11-338 [PubMed]
- 22. McCarroll SA, Murphy CT, Zou S, Pletcher SD, Chin CS, Jan YN, Kenyon C, Bargmann CI, Li H. Comparing genomic expression patterns across species identifies shared transcriptional profile in aging. Nat Genet. 2004; 36:197–204. https://doi.org/10.1038/ng1291 [PubMed]
- 23. Kumimoto EL, Fore TR, Zhang B. Transcriptome Profiling Following Neuronal and Glial Expression of ALS-Linked SOD1 in Drosophila. G3 (Bethesda). 2013; 3:695–708. https://doi.org/10.1534/g3.113.005850 [PubMed]
- 24. DeVeale B, Brummel T, Seroude L. Immunity and aging: the enemy within? Aging Cell. 2004; 3:195–208. https://doi.org/10.1111/j.1474-9728.2004.00106.x [PubMed]
- 25. Myllymäki H, Valanne S, Rämet M. The Drosophila imd signaling pathway. J Immunol. 2014; 192:3455–62. https://doi.org/10.4049/jimmunol.1303309 [PubMed]
- 26. Buchon N, Silverman N, Cherry S. Immunity in Drosophila melanogaster—from microbial recognition to whole-organism physiology. Nat Rev Immunol. 2014; 14:796–810. https://doi.org/10.1038/nri3763 [PubMed]
- 27. Ponnappan S, Ponnappan U. Aging and immune function: molecular mechanisms to interventions. Antioxid Redox Signal. 2011; 14:1551–85. https://doi.org/10.1089/ars.2010.3228 [PubMed]
- 28. Badinloo M, Nguyen E, Suh W, Alzahrani F, Castellanos J, Klichko VI, Orr WC, Radyuk SN. Overexpression of antimicrobial peptides contributes to aging through cytotoxic effects in Drosophila tissues. Arch Insect Biochem Physiol. 2018; 98:e21464. https://doi.org/10.1002/arch.21464 [PubMed]
- 29. Lin YR, Parikh H, Park Y. Stress resistance and lifespan enhanced by downregulation of antimicrobial peptide genes in the Imd pathway. Aging (Albany NY). 2018; 10:622–31. https://doi.org/10.18632/aging.101417 [PubMed]
- 30. Landis GN, Abdueva D, Skvortsov D, Yang J, Rabin BE, Carrick J, Tavaré S, Tower J. Similar gene expression patterns characterize aging and oxidative stress in Drosophila melanogaster. Proc Natl Acad Sci USA. 2004; 101:7663–68. https://doi.org/10.1073/pnas.0307605101 [PubMed]
- 31. Seroude L, Brummel T, Kapahi P, Benzer S. Spatio-temporal analysis of gene expression during aging in Drosophila melanogaster. Aging Cell. 2002; 1:47–56. https://doi.org/10.1046/j.1474-9728.2002.00007.x [PubMed]
- 32. Petersen AJ, Rimkus SA, Wassarman DA. ATM kinase inhibition in glial cells activates the innate immune response and causes neurodegeneration in Drosophila. Proc Natl Acad Sci USA. 2012; 109:E656–64. https://doi.org/10.1073/pnas.1110470109 [PubMed]
- 33. Cao Y, Chtarbanova S, Petersen AJ, Ganetzky B. Dnr1 mutations cause neurodegeneration in Drosophila by activating the innate immune response in the brain. Proc Natl Acad Sci USA. 2013; 110:E1752–60. https://doi.org/10.1073/pnas.1306220110 [PubMed]
- 34. Helmfors L, Boman A, Civitelli L, Nath S, Sandin L, Janefjord C, McCann H, Zetterberg H, Blennow K, Halliday G, Brorsson AC, Kågedal K. Protective properties of lysozyme on β-amyloid pathology: implications for Alzheimer disease. Neurobiol Dis. 2015; 83:122–33. https://doi.org/10.1016/j.nbd.2015.08.024 [PubMed]
- 35. Stephano F, Nolte S, Hoffmann J, El-Kholy S, von Frieling J, Bruchhaus I, Fink C, Roeder T. Impaired Wnt signaling in dopamine containing neurons is associated with pathogenesis in a rotenone triggered Drosophila Parkinson’s disease model. Sci Rep. 2018; 8:2372. https://doi.org/10.1038/s41598-018-20836-w [PubMed]
- 36. Myllymäki H, Rämet M. JAK/STAT pathway in Drosophila immunity. Scand J Immunol. 2014; 79:377–85. https://doi.org/10.1111/sji.12170 [PubMed]
- 37. Tanji T, Hu X, Weber AN, Ip YT. Toll and IMD pathways synergistically activate an innate immune response in Drosophila melanogaster. Mol Cell Biol. 2007; 27:4578–88. https://doi.org/10.1128/MCB.01814-06 [PubMed]
- 38. Kumar DK, Choi SH, Washicosky KJ, Eimer WA, Tucker S, Ghofrani J, Lefkowitz A, McColl G, Goldstein LE, Tanzi RE, Moir RD. Amyloid-β peptide protects against microbial infection in mouse and worm models of Alzheimer’s disease. Sci Transl Med. 2016; 8:340ra72. https://doi.org/10.1126/scitranslmed.aaf1059 [PubMed]
- 39. Iijima K, Chiang HC, Hearn SA, Hakker I, Gatt A, Shenton C, Granger L, Leung A, Iijima-Ando K, Zhong Y. Abeta42 mutants with different aggregation profiles induce distinct pathologies in Drosophila. PLoS One. 2008; 3:e1703. https://doi.org/10.1371/journal.pone.0001703 [PubMed]
- 40. Chen Y, Chen Y, Shi C, Huang Z, Zhang Y, Li S, Li Y, Ye J, Yu C, Li Z, Zhang X, Wang J, Yang H, et al. SOAPnuke: a MapReduce acceleration-supported software for integrated quality control and preprocessing of high-throughput sequencing data. Gigascience. 2018; 7:1–6. https://doi.org/10.1093/gigascience/gix120 [PubMed]
- 41. Thurmond J, Goodman JL, Strelets VB, Attrill H, Gramates LS, Marygold SJ, Matthews BB, Millburn G, Antonazzo G, Trovisco V, Kaufman TC, Calvi BR, Perrimon N, et al, and FlyBase Consortium. FlyBase 2.0: the next generation. Nucleic Acids Res. 2019; 47:D759–65. https://doi.org/10.1093/nar/gky1003 [PubMed]
- 42. Kim D, Langmead B, Salzberg SL. HISAT: a fast spliced aligner with low memory requirements. Nat Methods. 2015; 12:357–60. https://doi.org/10.1038/nmeth.3317 [PubMed]
- 43. Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014; 15:550. https://doi.org/10.1186/s13059-014-0550-8 [PubMed]