Introduction
Developmental dysplasia of the hip (DDH) (OMIM#142700) is a common congenital disease with both environmental and genetic components [1]. DDH patients present distinctively with hip joint laxity and skeletal dysplasia in the hip [2]. Subsequent dysfunction brings unacceptable life burden with skeletal development and physical growth [3]. DDH is a hereditary progression [4–6], though mechanical factors (e.g. breech delivery, high birth weight, primiparity and oligoamnios) are suggested [7, 8]. Several DDH susceptibility genes (e.g. GDF5, TBX4, ASPN, PAPPA2 and TGFB1) were discovered by association study in Chinese and Caucasian populations [9–13]. Gene mutations not only confer the sequence changes, but can also elicit certain structural change in the microenvironment [14, 15]. A variety of environmental factors have also been predicted to participate in DDH [16]. Yet the complex etiology of DDH remains to be clarified.
Various members (e.g. type II collagen, type XI collagen) in the collagen family are vital in skeletal development and connective tissue formation. Type XI collagen is composed of three subunits (α1, α2, α3) and expressed in bone, connective tissue and cartilage. It forms the major structure of articular cartilage and the core of collagen fibers together with type II collagen [17, 18]. COL11A2 (collagen, type XI, alpha 2) gene (OMIM#120290) codes for the α2 subunit of type collagen XI. Mutations in COL11A2 were reported to cause many inherited skeletal disorders like autosomal recessive otospondylomegaepiphyseal dysplasia (OSMED) (OMIM#215150) [19] and Marshall’s or Stickler syndrome (OMIM#184840) [20]. Several lines of evidence suggest that COL11A2 plays an important role in skeletal development. Nevertheless, there is no study of COL11A2 concerning developmental dysplasia of the hip. In the present study, we have explored the association between COL11A2 gene and DDH in the Han Chinese population.
Results
Genetic association of locus rs9277935 in COL11A2 gene with DDH
Genotype distribution in both groups were conformed to Hardy-Weinberg equilibrium (p>0.01). Genotype and allele frequency distribution of Chinese Han population DDH group and healthy controls are shown in Table 1. The statistical analysis results indicate there is a trend of significant difference in allele frequency for the two groups (p=0.099). A significant difference in genotype distribution in a recessive model (TT+GT vs. GG) between two groups (P=0.017) was demonstrated. Due to the high prevalence of female patients, the genotype in the female from both groups was analyzed. (Table 2) There was significantly greater frequency of minor allele G in the female DDH patients (0.49 vs. 0.43, p=0.024). The recessive model also revealed significantly higher distribution of GG genotype in female DDH patients (p=0.006).
Table 1. Association between rs9277935 and DDH in different genetic models.
| Genotype | Allele frequency | P value | |||||||
| T vs G | GG vs others | TT vs others | GG | GT | TT | No. | G | T | |
| Case | 76 | 183 | 91 | 350 | 0.48 | 0.52 | |||
| Control | 127 | 269 | 199 | 595 | 0.44 | 0.56 | 0.099 | 0.017 | 0.89 |
Table 2. Association between rs9277935 and DDH in the female population.
| Genotype | Allele frequency | P value | |||||||
| GG | GT | TT | No. | G | T | T vs G | GG vs others | TT vs others | |
| Female case | 71 | 163 | 78 | 312 | 0.49 | 0.51 | |||
| Female control | 91 | 202 | 153 | 446 | 0.43 | 0.57 | 0.024 | 0.006 | 0.43 |
Dysregulated joint tissue expression of the COL11A2 in DDH and the controls
Expression level of the COL11A2 was summarized in the patients and in the controls, respectively (Figure 1A). DDH patients had significantly lower expression of the COL11A2 in both joint cartilage and ligament compared with the control. (2.43 ± 1.65 vs 4.05 ± 1.89, p = 0.002 for articular cartilage; 2.46 ±1.68 vs. 3.85 ± 2.73, p = 0.007 for joint ligament). Moreover, immunofluorescent assay also demonstrated significantly lower COL11A2 expression in the tissue sections of cartilage and ligament tissues in DDH patients observed under confocal microscopy.
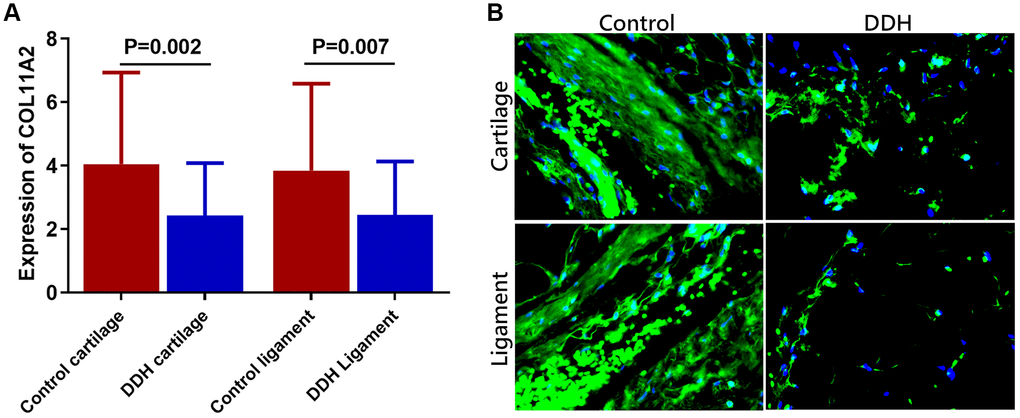
Figure 1. Tissue expression of COL11A2 in patients and controls. (A) DDH patients were found to have significantly lower expression of the COL11A2 in the articular cartilage and ligament as compared with the controls (2.43 ± 1.65 vs 4.05 ± 1.89, p = 0.002 for articular cartilage; 2.46 ±1.68 vs. 3.85 ± 2.73, p = 0.007 for joint ligament). (B) Immunofluorescent assay of COL11A2 (green) expression and nucleus (blue) in the cartilage and ligament tissues in different groups of patients observed under confocal microscopy.
Relationship between the genotype of rs9277935 and the COL11A2 expression
Results of the comparison of the COL11A2 expression among patients with different genotypes are shown in Figure 2A. The mean value of COL11A2 expression in articular cartilage and joint ligament were respectively 3.07 ± 2.11 and 3.15 ± 2.33 for genotype TT, 2.59 ± 1.87 and 2.66 ± 1.95 for genotype TG, and 1.62 ± 1.13 and 1.52 ± 1.45 for genotype GG. Patients with genotype TT were found to have a remarkably higher COL11A2 expression than those with genotype GG in both articular cartilage and joint ligament. (p = 0.02 for articular cartilage; p = 0.036 for joint ligament) Different COL11A2 expression was also demonstrated in tissue sections of cartilage and ligament tissues in DDH patients with different genotypes for rs9277935 with immunofluorescent assay, showing significantly greater COL11A2 expression in patients with TT genotype compared to those with GG genotype. (Figure 2B).
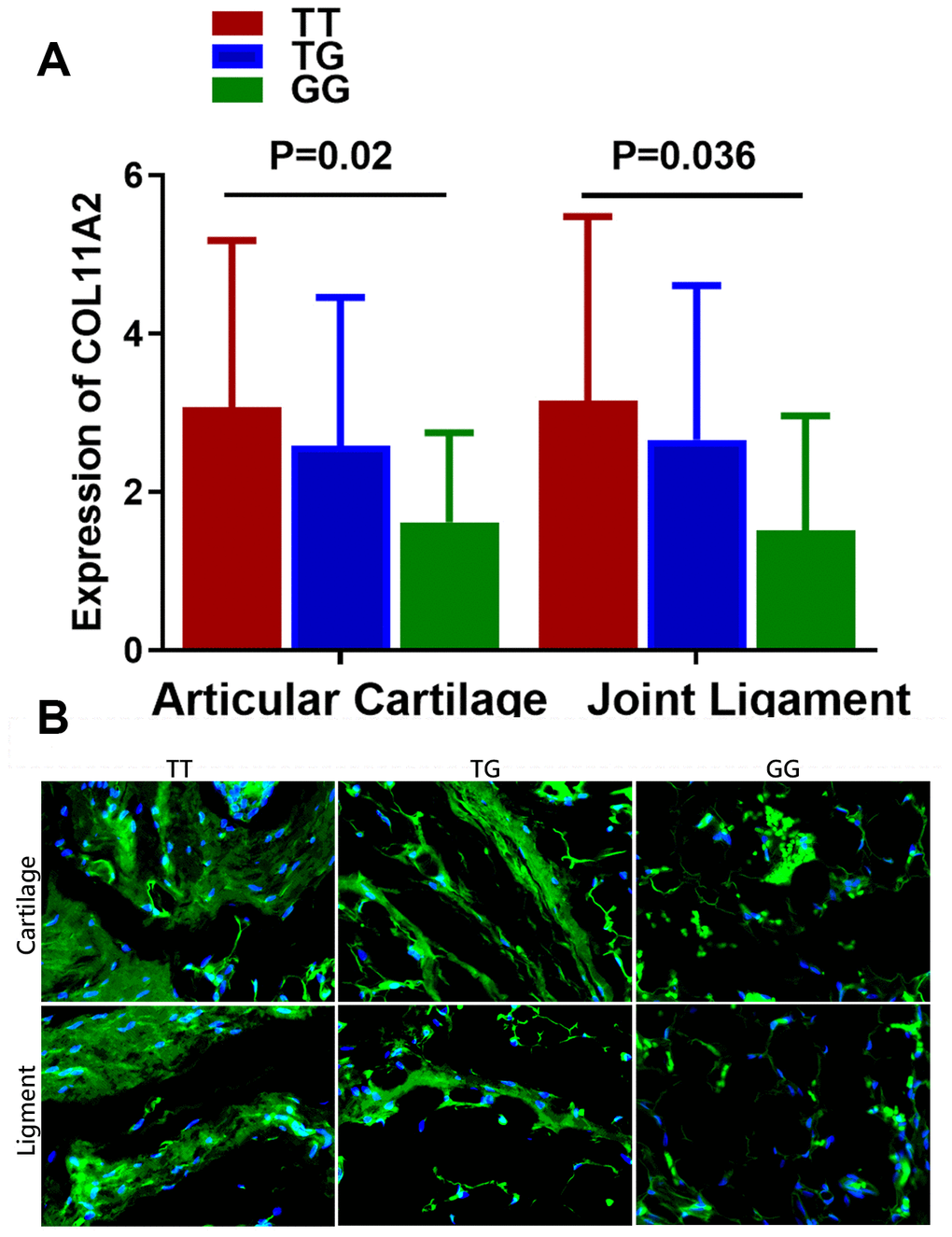
Figure 2. Relationship between the genotype of rs9277935 and COL11A2 expression in DDH joint tissues. (A) DDH Patients with genotype TT have a significantly increased expression of COL11A2 than those with GG (p = 0.02 for articular cartilage; p = 0.036 for joint ligament). (B) Immunofluorescent assay of COL11A2 (green) expression and nucleus (blue) in the cartilage and ligament tissues in DDH patients with different genotypes observed under confocal microscopy.
Allelic difference in Col11a2 expression by rs9277935 and the chondrogenic effects of Col11a2 on BMSCs in vitro
To verify the allelic difference in Col11a2 expression by locus rs9277935 we tested the luciferase activity driven by different alleles. (Figure 3A) In comparison, it was apparent that T allele drove greater luciferase expression than G, with a 40% difference seen in ATDC5 (Figure 3A), indicating rs9277935 as a causative locus driving the luciferase expression change. These data clearly demonstrate that the rs9277935 variant associated with DDH were functional and mediated dysregulated Col11a2 expression, which partly explained the decreased Col11a2 expression in DDH. To test the involvement of dysregulated Col11a2 expression in chondrogenesis, potential chondrogenic effects of Col11a2 on chondrogenesis of bone marrow stem cells (BMSCs) were further analyzed in vitro. (Figure 3B to I) Addition of Col11a2 blocking antibody lowered production of glycosaminoglycans (GAGs) while application of exogenous recombinant Col11a2 (50ng/ml) for 2 weeks in culture induced greater synthesized glycosaminoglycans (GAGs) with strong Alcian blue staining, indicating differentiation of BMSCs into chondrocyte-like cells. The chondrogenic effects of exogenous Col11a2 was further evidenced by its neutralization in the group with Col11a2 blocking antibody. Chondrocytes generated in the several groups were next analyzed with immunostaining and RT-PCR (Figure 3C–3I) for expression of genes expressed by chondrocytes (ACAN, SOX9, COL1A1, COL2A1). Compared to the control, exogenous Col11a2 treatment led to significant chondrogenic effects evidenced by chondrogenic gene expression (Figure 3C–3F) and GAG deposition with higher DNA quantification of collagen I and II (Figure 3G–3I). The chondrogenic effects of Col11a2 on BMSCs were blocked with supplemented Col11a2 blocking antibody in the other two groups. These results indicated that Col11a2 as an important mediator in chondrogenesis and cartilage formation.
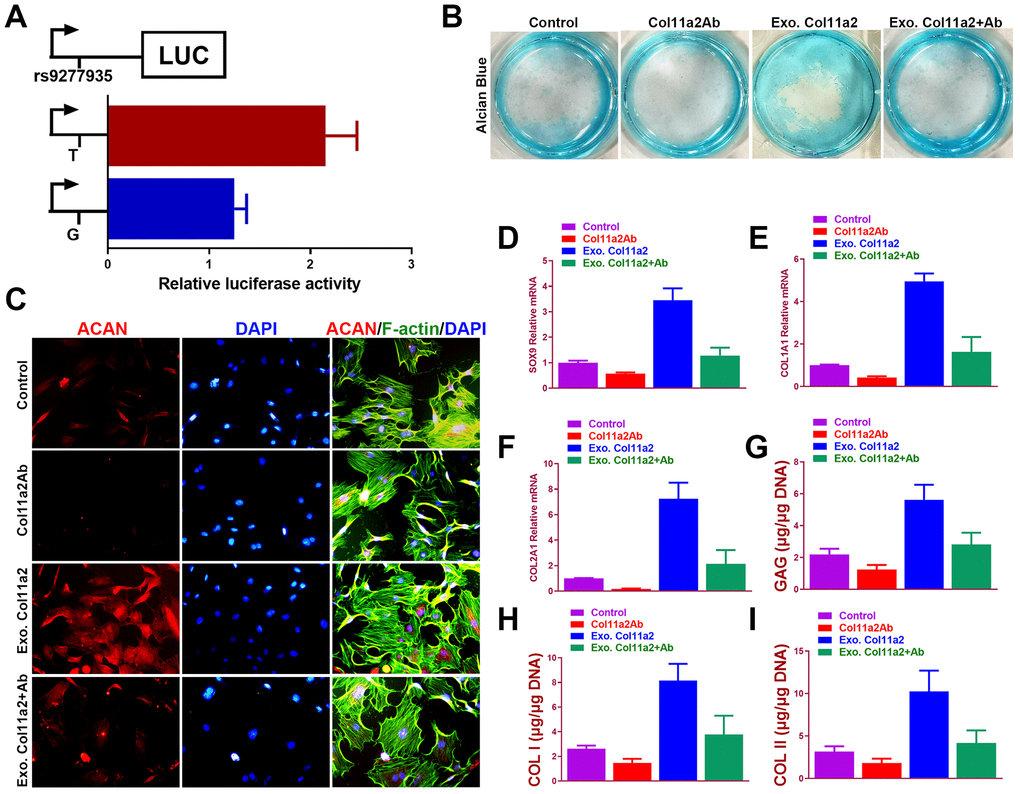
Figure 3. Chondrogenic effects of Col11a2 in vitro. (A) Luciferase activity to indicate the allelic difference in Col11a2 expression driven by rs9277935 (n=6 for each) in ATDC5 cells. (B) Alcian blue staining of BMSCs in different treatment groups at 2 weeks to indicate GAG production in the culture plate. (C) Immunofluorescent assay of Aggrecan (red) expression, cytoskeleton (green) nucleus (blue) in different treatment groups observed under confocal microscopy. Scale bar=200μm. (D–F) Expression level of chondrogenic markers for BMSCs in different treatment groups (n=6 for each) in (B) and (G–I) quantification of GAG production, COL I and II DNA in generated cartilaginous tissues (n=6 for each). *P < 0.05 between control group and other groups. Data are presented as averages ± SD. One-way analysis of variance (ANOVA) with post-hoc Tukey’s B test was applied. Ab, antibody; Exo, exogenous; ACAN, aggrecan; GAG, glycosaminoglycan.
Discussion
In the present study, we first explored the association between COL11A2 gene and DDH. The minor allele frequency of rs9277935 in the control group in our study (0.44) is close to that reported in HapMap for Chinese Han population (0.47). A trend of difference of allele frequency between DDH and control group was demonstrated. Whereas a significant difference of) genotype distribution in a recessive model (TT+GT vs. GG between two groups was identified. Further analysis in female population showed significant difference in both genotypes and allele frequency distributions, suggesting association between rs9277935 in gene COL11A2 and DDH occurrence in female patients. DDH patients were found to have a significantly lower expression of COL11A2 in both the cartilage and ligament than the control groups. Besides, patients with genotype TT of rs9277935 were found to have a remarkably increased expression of COL11A2 than those with GG. Allele G is indicative of lower expression of COL11A2 as well as a higher risk of the development of DDH. Taken together, rs9277935 could be a functional variant regulating the expression of COL11A2.
DDH is a complex polygenic disease, whereby multiple susceptibility genes have been reported. Jia et al reported that rs726252 in pregnancy - associated plasma protein - A2 gene, with significantly greater distribution of TT genotype in DDH patients, was associated with DDH [10]. The research conducted by Shi et al showed an obvious association between the D repeat polymorphism of ASPN and DDH, which indicated that ASPN is an important regulator in the etiology of DDH [11]. In addition, Wang et al discovered that a single nucleotide polymorphism in Tbx4 was also associated with DDH [12]. Furthermore, Jin et al found that GDF5 is important in the etiology of congenital dysplasia of the hip [13]. A recent study showed that TGFB1 played a genetic role in the risk of DDH [9]. All of these indicated the complex genetic components in DDH occurrence. DDH patients usually present distinctively with laxity of joint capsule, skeletal dysplasia of the femoral head and acetabular dysplasia. However, the patients retrieved in the present study were mostly the elder DDH patients with secondary arthritis. In this case, the exact primary cause of DDH was difficult to determine in these patients. Further studies of DDH in infants or adolescents could help to address this issue.
COL11A2 is an indispensable gene involved in collagen formation and extracellular matrix building, which are fundamental in development of cartilage and articular joint, shaping of joint morphology and biomechanical balance maintaining [21–24], indicating a possible role of COL11A2 in DDH development. This gene codes for type collagen XI, which is expressed in cartilage and bone. Collagen type XI constitutes the major structure of articular cartilage with collagen type II. It is wildly expressed in cartilage as a target gene downstream of Sox9 [25]. While Sox9 gene haplo-insufficiency can lead to genetic skeletal disorders like campomelic dysplasia (CMD) (OMIM#114290) characterized by bowing long bones [26, 27]. COL11A2 mutations are also associated with a lot of inherited skeletal diseases, whereby Otospondylomegaepiphyseal dysplasia (OSMED) is autosomal recessive disease, featuring auditory dysfunction and short asymmetrical limbs [28]. Mutations of COL11A2 can also cause an skeletal dysplasia called fibrochondrogenesis (OMIM#614524) [29]. Moreover, Pihlajamaa et al reported a single nucleotide mutation in COL11A2 gene that converted glycine to glutamate (G955E) and caused Weissenbacher-Zweymuller syndrome (OMIM#277610) [30]. One C to T transition in exon 56, which led to premature translation termination codon in COL11A2 (R845X), was also reported in individuals with Weissenbacher-Zweymuller syndrome [31]. In addition, abnormal COL11A2 gene structure was also associated with Marshall’s or Stickler syndrome marked with skeletal dysplasia and joint pain [32].
Close relationship between COL11A2 gene and the aforementioned skeletal disorders is suggestive of its significant role in bone and joint development. This present study was the first report concerning COL11A2 in developmental dysplasia of the hip to date. The locus rs9277935 was formerly reported to associate with autoimmune disease (e.g. Wegener's granulomatosis) [33]. rs9277935 locates in the promoter region of gene COL11A2 and modifies the expression of COL11A2 as demonstrated by the allelic difference of COL11A2 expression driven by rs9277935 in the present study. It is possible that rs9277935 affected the binding of transcript factors or other motifs to the promoter of COL11A2 gene to regulate its final expression. In clinical practice, DDH patients usually present with laxity of joint capsule and chondrogenic dysplasia of the femoral head. Arthritic changes occur in most of DDH patients, afflicting mainly the chondrocytes of the femoral head. COL11A2 was an indispensable gene in chondrogenesis and rs9277935 modulated COL11A2 in the present study in DDH patients. Therefore, we further validated the chondrogenic role of COL11A2 in studies in vitro, suggesting that altered chondrogenesis with changed COL11A2 signaling by the variant rs9277935 could be the causative role in DDH. The association between rs9277935 and DDH was mediated by dysregulated chondrogenesis and joint cartilage formation caused by the allelic difference of COL11A2 gene expression. Experiments in vivo to further explore the function of COL11A2 in DDH occurrence is necessary.
Conclusions
In conclusion, we demonstrated the locus rs9277935 of the chondrogenic gene COL11A2 was functionally associated with DDH with dysregulated COL11A2 expression in hip joint tissues in Chinese Han population. Genotype TT of rs9277935 is associated with higher expression of COL11A2 in DDH. Genetic variants of COL11A2 could be a potential biomarker for early diagnosis of DDH. Replication work in other ethnic populations is needed.
Materials and Methods
Patients
We enrolled 350 radiologically confirmed DDH patients and 595 healthy controls to conduct a case-control candidate gene association study. DDH patients were consecutively recruited from the department of orthopedics, the first affiliated hospital of Soochow University. Controls were recruited from physical examination center in the first affiliated hospital of Soochow University. All the subjects were Han Chinese living in or around Suzhou. The study was approved by the ethical committee of the Soochow University, and informed consents were obtained from all patients and controls.
Sample collection
After informed consent was obtained from the families, blood samples were collected for genomic DNA extraction using the commercial kit (QIAGEN) according to the standard protocol. Hip joint articular cartilage and ligament were obtained in 65 DDH patients during hip arthroplasty surgery. 20 age-matched trauma patients undergoing amputation surgery were recruited as control. Tissue samples were placed in separate sterile tubes and immediately stored in liquid nitrogen. Frozen samples were stored at -80 °C. Total RNA was extracted using Trizol reagent (QIAGEN) according to the manufacturer’s protocol. To avoid genomic DNA contamination in RNA, samples were treated with DNase (QIAGEN). Total RNA was then reverse-transcribed from 2ug of RNA using the PrimeScript RT Master Mix kit (TaKaRa).
Genotyping of targeted locus
According to the manufacture’s protocol, the DNA of all the subjects was extracted either from the buccal swabs using the DNA IQ System (Promega, Madison, WI) or peripheral blood using the NucleoSpin Blood QuickPure Kit (Macherey-Nagel GmbH & Co. KG, Düren, German). All the samples were genotyped with Taqman assay. The sample was genotyped by uninformed laboratory personnel. Genotyping, data input and statistical results were examined by two authors independently. Five percent of samples were randomly selected to repeat, and 100% consistency was obtained.
Tissue expression of the COL11A2 gene in DDH patients and controls
The tissue expression of COL11A2 was measured with real-time PCR using gene-specific primers as follows: forward 5’-GCCTCAGCCTAGCAGAT -3’, reverse 5’- ATCACTCCATGGGTGTCCAATA -3’ for the COL11A2 gene, and forward 5’-GAGTC AACGGATTTGGTCGT-3’, reverse 5’-TTGATTTTGGAGGGATCTCG-3’ for the endogenous control gene Glyceraldehyde-3-phosphate dehydrogenase (GAPDH). For real-time PCR, 1 μL of cDNA was amplified for 40 cycles by SYBR.
Premix Ex TaqTM II (TaKaRa) in ABI 7900HT mentioned above. Melting curve analysis was done at the end of the reaction to assess the quality of the final PCR products. All samples were analyzed in triplicate using the 2-ΔΔCt method.
Immunofluorescent staining of histological sections
Frozen sections of hip joint tissue were fixed with 4% paraformaldehyde for 5 min. After washing with TBS-T, the sections were incubated with 10 mg/mL hyaluronidase (Sigma) at 37 °C for 30 min. After blocking with 10% goat normal serum, the sections were incubated with primary antibody for 2 hat room temperature. Paraffin-embedded sections were deparaffinized and incubated in 1mM EDTA (pH 8.0) at 80 °C for 15 mins to retrieve the antigen. Then sections were treated with 10 mg/mL hyaluronidase at 37 °C for 30 min. The primary antibodies anti-COL11A2 were used (diluted 1:200, Abcam). Immune complexes were detected using secondary antibody. DAPI was also used to detect the nucleus.
Construction of COL11A2-pGL3-basic luciferase reporter plasmids
COL11A2-pGL3-basic luciferase reporter plasmids for rs9277935 were constructed as previously reported [2]. Briefly, ATDC5 cells were transfected using 500 ng of pGL3 plasmid DNA and 15 ng of pTK-RL Renilla plasmid in combination with Exgen 500 Transfection reagent (Fermentas, Sankt Leon-Rot, Germany) [34–36]. Twenty-four hours after transfection ATDC5 were lysed, using passive lysis buffer (Promega), and the protein extracted and stored at -20 °C. Lysate sample was mixed with luciferase activating reagent II (Promega) and a 1-s luciferase activity reading was measured using a luminometer (EG&G Berthold, Bad Wildbad, Germany). Stop and Glo (Promega) was added to each sample to measure renilla activity. An empty vector transfection was performed as a control. Each experiment contained six replicates, and was repeated three times, producing a total of 18 data points. A Student’s t-test was performed to assess any significant differences in luciferase activity between the different alleles.
Isolation and treatment of BMSC in vitro
Mouse BMSCs were isolated from rabbit bone marrow aspirates [37]. Briefly, marrow aspirates (20 mL volume) were harvested and immediately transferred into plastic tubes. Isolated BMSCs were expanded in α-MEM containing 10% FBS, 4.5 mg/mL D-glucose,0.1 mM nonessential amino acids, 1 mM sodium pyruvate, 100 mM Hepes buffer, 100 Ul/mL penicillin, 100 μg/mL streptomycin, and 0.29 mg/mL L-glutamate. Medium was changed twice a week and BMSCs were used at P2 for the following experiments [34, 38]. In the exogenous COL11A2 group, COL11A2 (50 ng/ml) was added in the medium for 2 weeks. COL11A2 neutralizing peptide was added in some of the cultures according to the protocol. Medium was also changed twice a week.
Immunofluorescence staining of chondrogenic marker ACAN was conducted to compare the phenotypes of generated chondrocytes in different groups and observed under confocal microscopy (Leica, Japan). The expression of chondrogenesis marker (SOX9, Col1A1 and Col2A1) after 2-week incubation was analyzed by real-time polymerase chain reaction (RT-PCR) using an ABI 7300 RT-PCR system (Applied Biosystems, USA). Two-week-old tissue generated under different conditions was observed and stained with alcian blue for proteoglycan production. The stained images were taken using a light microscope (Leica Microsystems, Germany).
GAGs and types I and II collagen were quantitatively assayed (6 vs 6) and normalized to DNA content. GAG production, Collagen I and II expression were compared among different treatment groups.
Statistics
Hardy-Weinberg equilibrium was calculated by chi-squared test in both control and case groups. The association between the DDH patients and the control subjects in the stages was tested by SAS software (version 9.2 - SAS Institute, Cary, NC, USA). Bilateral chi square tests were conducted to determine the significance of differences in allelic frequencies and P < 0.05 was considered to be statistically significant. The Student t test was used to compare the difference of COL11A2 expression between the patients and the controls. DDH patients were classified into three groups according to the genotypes of each SNP, and One-way ANOVA test was used to compare the COL11A2 expression among different genotypes.
Author Contributions
Renjie Xu and Yuxing Zhang conceived the study. Junlan Lu and Kexin Wang completed the collection and assembly of data. Xin Jiang undertook the statistical analysis. Renjie Xu and Ye Sun produced the initial draft manuscript. Ye Sun and Yuxin Zhang critically revised the article for important intellectual content and gave final approval of the article. All authors read the final manuscript.
Conflicts of Interest
The authors declare that they have no conflicts of interest.
Funding
This study was funded by China National Natural Science Funds (No. 81802122), the Project funded by China Postdoctoral Science Foundation (No. 2019M661559), the Funds from Shanghai jiao tong university for the Clinical and Translational Research Center for 3D Printing Technology and the Fundamental research program funding of Ninth People's Hospital affiliated to Shanghai Jiao Tong university School of Medicine (JYZZ078).
References
- 1. Grzegorzewski A, Jóźwiak M, Pawlak M, Modrzewski T, Buchcic P, Masłoń A. Hip joint pain in children with cerebral palsy and developmental dysplasia of the hip: why are the differences so huge? BMC Musculoskelet Disord. 2014; 15:96. https://doi.org/10.1186/1471-2474-15-96 [PubMed]
- 2. Sun Y, You Y, Jiang W, Zhai Z, Dai K. 3D-bioprinting a genetically inspired cartilage scaffold with GDF5-conjugated BMSC-laden hydrogel and polymer for cartilage repair. Theranostics. 2019; 9:6949–61. https://doi.org/10.7150/thno.38061 [PubMed]
- 3. Judd J, Clarke NM. Treatment and prevention of hip dysplasia in infants and young children. Early Hum Dev. 2014; 90:731–34. https://doi.org/10.1016/j.earlhumdev.2014.08.011 [PubMed]
- 4. Wynne-Davies R. A family study of neonatal and late-diagnosis congenital dislocation of the hip. J Med Genet. 1970; 7:315–33. https://doi.org/10.1136/jmg.7.4.315 [PubMed]
- 5. Stevenson DA, Mineau G, Kerber RA, Viskochil DH, Schaefer C, Roach JW. Familial predisposition to developmental dysplasia of the hip. J Pediatr Orthop. 2009; 29:463–66. https://doi.org/10.1097/BPO.0b013e3181aa586b [PubMed]
- 6. Li L, Sun K, Zhang L, Zhao Q, Cheng X, Dang Y. Heritability and sibling recurrent risk of developmental dysplasia of the hip in Chinese population. Eur J Clin Invest. 2013; 43:589–94. https://doi.org/10.1111/eci.12084 [PubMed]
- 7. Stein-Zamir C, Volovik I, Rishpon S, Sabi R. Developmental dysplasia of the hip: risk markers, clinical screening and outcome. Pediatr Int. 2008; 50:341–45. https://doi.org/10.1111/j.1442-200X.2008.02575.x [PubMed]
- 8. Chan A, McCaul KA, Cundy PJ, Haan EA, Byron-Scott R. Perinatal risk factors for developmental dysplasia of the hip. Arch Dis Child Fetal Neonatal Ed. 1997; 76:F94–100. https://doi.org/10.1136/fn.76.2.f94 [PubMed]
- 9. Ma W, Zha Z, Chen K, Chen H, Wu Y, Ma J, Zeng S, Zhi L, Yao S. Genetic association study of common variants in TGFB1 and IL-6 with developmental dysplasia of the hip in Han Chinese population. Sci Rep. 2017; 7:10287. https://doi.org/10.1038/s41598-017-11185-1 [PubMed]
- 10. Jia J, Li L, Zhao Q, Zhang L, Ru J, Liu X, Li Q, Shi L. Association of a single nucleotide polymorphism in pregnancy-associated plasma protein-A2 with developmental dysplasia of the hip: a case-control study. Osteoarthritis Cartilage. 2012; 20:60–63. https://doi.org/10.1016/j.joca.2011.10.004 [PubMed]
- 11. Shi D, Dai J, Zhu P, Qin J, Zhu L, Zhu H, Zhao B, Qiu X, Xu Z, Chen D, Yi L, Ikegawa S, Jiang Q. Association of the D repeat polymorphism in the ASPN gene with developmental dysplasia of the hip: a case-control study in Han Chinese. Arthritis Res Ther. 2011; 13:R27. https://doi.org/10.1186/ar3252 [PubMed]
- 12. Hao Z, Dai J, Shi D, Xu Z, Chen D, Zhao B, Teng H, Jiang Q, Jiang Q. Association of a single nucleotide polymorphism in HOXB9 with developmental dysplasia of the hip: a case-control study. J Orthop Res. 2014; 32:179–82. https://doi.org/10.1002/jor.22507 [PubMed]
- 13. Dai J, Shi D, Zhu P, Qin J, Ni H, Xu Y, Yao C, Zhu L, Zhu H, Zhao B, Wei J, Liu B, Ikegawa S, et al. Association of a single nucleotide polymorphism in growth differentiate factor 5 with congenital dysplasia of the hip: a case-control study. Arthritis Res Ther. 2008; 10:R126. https://doi.org/10.1186/ar2540 [PubMed]
- 14. Kou Y, Koag MC, Lee S. N7 methylation alters hydrogen-bonding patterns of guanine in duplex DNA. J Am Chem Soc. 2015; 137:14067–70. https://doi.org/10.1021/jacs.5b10172 [PubMed]
- 15. Kou Y, Koag MC, Lee S. Structural and Kinetic Studies of the Effect of Guanine N7 Alkylation and Metal Cofactors on DNA Replication. Biochemistry. 2018; 57:5105–16. https://doi.org/10.1021/acs.biochem.8b00331 [PubMed]
- 16. Rhodes AM, Clarke NM. A review of environmental factors implicated in human developmental dysplasia of the hip. J Child Orthop. 2014; 8:375–79. https://doi.org/10.1007/s11832-014-0615-y [PubMed]
- 17. Mendler M, Eich-Bender SG, Vaughan L, Winterhalter KH, Bruckner P. Cartilage contains mixed fibrils of collagen types II, IX, and XI. J Cell Biol. 1989; 108:191–97. https://doi.org/10.1083/jcb.108.1.191 [PubMed]
- 18. Tsumaki N, Kimura T, Matsui Y, Nakata K, Ochi T. Separable cis-regulatory elements that contribute to tissue- and site-specific alpha 2(XI) collagen gene expression in the embryonic mouse cartilage. J Cell Biol. 1996; 134:1573–82. https://doi.org/10.1083/jcb.134.6.1573 [PubMed]
- 19. Kayserili H, Wollnik B, Güven G, Emiroğlu MU, Başerer N, Uyguner ZO. A novel homozygous COL11A2 deletion causes a C-terminal protein truncation with incomplete mRNA decay in a Turkish patient. Am J Med Genet A. 2011; 155A:180–85. https://doi.org/10.1002/ajmg.a.33780 [PubMed]
- 20. van Beelen E, Leijendeckers JM, Huygen PL, Admiraal RJ, Hoefsloot LH, Lichtenbelt KD, Stöbe L, Pennings RJ, Leuwer R, Snik AF, Kunst HP. Audiometric characteristics of two Dutch families with non-ocular Stickler syndrome (COL11A2). Hear Res. 2012; 291:15–23. https://doi.org/10.1016/j.heares.2012.07.001 [PubMed]
- 21. Baas D, Malbouyres M, Haftek-Terreau Z, Le Guellec D, Ruggiero F. Craniofacial cartilage morphogenesis requires zebrafish col11a1 activity. Matrix Biol. 2009; 28:490–502. https://doi.org/10.1016/j.matbio.2009.07.004 [PubMed]
- 22. Carter EM, Raggio CL. Genetic and orthopedic aspects of collagen disorders. Curr Opin Pediatr. 2009; 21:46–54. https://doi.org/10.1097/MOP.0b013e32832185c5 [PubMed]
- 23. Avcin T, Makitie O, Susic M, Miller S, Thorne C, Tenenbaum J, Laxer RM, Cole WG. Early-onset osteoarthritis due to otospondylomegaepiphyseal dysplasia in a family with a novel splicing mutation of the COL11A2 gene. J Rheumatol. 2008; 35:920–26. [PubMed]
- 24. Noponen-Hietala N, Kyllönen E, Männikkö M, Ilkko E, Karppinen J, Ott J, Ala-Kokko L. Sequence variations in the collagen IX and XI genes are associated with degenerative lumbar spinal stenosis. Ann Rheum Dis. 2003; 62:1208–14. https://doi.org/10.1136/ard.2003.008334 [PubMed]
- 25. Park JS, Yi SW, Kim HJ, Oh HJ, Lee JS, Go M, Shim SH, Park KH. Verification of Long-Term Genetic Stability of hMSCs during Subculture after Internalization of Sunflower-Type Nanoparticles (SF-NPs). Theranostics. 2018; 8:5548–61. https://doi.org/10.7150/thno.29214 [PubMed]
- 26. Yokoi H, Yan YL, Miller MR, BreMiller RA, Catchen JM, Johnson EA, Postlethwait JH. Expression profiling of zebrafish sox9 mutants reveals that Sox9 is required for retinal differentiation. Dev Biol. 2009; 329:1–15. https://doi.org/10.1016/j.ydbio.2009.01.002 [PubMed]
- 27. Mayo JL, Holden DN, Barrow JR, Bridgewater LC. The transcription factor Lc-Maf participates in Col27a1 regulation during chondrocyte maturation. Exp Cell Res. 2009; 315:2293–300. https://doi.org/10.1016/j.yexcr.2009.04.020 [PubMed]
- 28. Tokgöz-Yılmaz S, Sahlı S, Fitoz S, Sennaroğlu G, Tekin M. Audiological findings in otospondylomegaepiphyseal dysplasia (OSMED) associated with a novel mutation in COL11A2. Int J Pediatr Otorhinolaryngol. 2011; 75:433–37. https://doi.org/10.1016/j.ijporl.2010.12.004 [PubMed]
- 29. Tompson SW, Faqeih EA, Ala-Kokko L, Hecht JT, Miki R, Funari T, Funari VA, Nevarez L, Krakow D, Cohn DH. Dominant and recessive forms of fibrochondrogenesis resulting from mutations at a second locus, COL11A2. Am J Med Genet A. 2012; 158A:309–14. https://doi.org/10.1002/ajmg.a.34406 [PubMed]
- 30. Pihlajamaa T, Prockop DJ, Faber J, Winterpacht A, Zabel B, Giedion A, Wiesbauer P, Spranger J, Ala-Kokko L. Heterozygous glycine substitution in the COL11A2 gene in the original patient with the Weissenbacher-Zweymüller syndrome demonstrates its identity with heterozygous OSMED (nonocular Stickler syndrome). Am J Med Genet. 1998; 80:115–20. https://doi.org/10.1002/(SICI)1096-8628(19981102)80:2<115::AID-AJMG5>3.0.CO;2-O [PubMed]
- 31. Harel T, Rabinowitz R, Hendler N, Galil A, Flusser H, Chemke J, Gradstein L, Lifshitz T, Ofir R, Elbedour K, Birk OS. COL11A2 mutation associated with autosomal recessive Weissenbacher-Zweymuller syndrome: molecular and clinical overlap with otospondylomegaepiphyseal dysplasia (OSMED). Am J Med Genet A. 2005; 132A:33–35. https://doi.org/10.1002/ajmg.a.30371 [PubMed]
- 32. Acke FR, Malfait F, Vanakker OM, Steyaert W, De Leeneer K, Mortier G, Dhooge I, De Paepe A, De Leenheer EM, Coucke PJ. Novel pathogenic COL11A1/COL11A2 variants in Stickler syndrome detected by targeted NGS and exome sequencing. Mol Genet Metab. 2014; 113:230–35. https://doi.org/10.1016/j.ymgme.2014.09.001 [PubMed]
- 33. Wieczorek S, Knaup S, Gross WL, Epplen JT. Genetic variability of RXRB, PPARA, and PPARG in Wegener’s granulomatosis. PPAR Res. 2009; 2009:786781. https://doi.org/10.1155/2009/786781 [PubMed]
- 34. Yi SW, Park JS, Kim HJ, Lee JS, Woo DG, Park KH. Multiply clustered gold-based nanoparticles complexed with exogenous pDNA achieve prolonged gene expression in stem cells. Theranostics. 2019; 9:5009–19. https://doi.org/10.7150/thno.34487 [PubMed]
- 35. Vonk LA, van Dooremalen SF, Liv N, Klumperman J, Coffer PJ, Saris DB, Lorenowicz MJ. Mesenchymal Stromal/stem Cell-derived Extracellular Vesicles Promote Human Cartilage Regeneration In Vitro.. Theranostics. 2018; 8:906–20. https://doi.org/10.7150/thno.20746 [PubMed]
- 36. Meng F, Li Z, Zhang Z, Yang Z, Kang Y, Zhao X, Long D, Hu S, Gu M, He S, Wu P, Chang Z, He A, Liao W. MicroRNA-193b-3p regulates chondrogenesis and chondrocyte metabolism by targeting HDAC3. Theranostics. 2018; 8:2862–83. https://doi.org/10.7150/thno.23547 [PubMed]
- 37. Sun Y, You Y, Jiang W, Wu Q, Wang B, Dai K. Generating ready-to-implant anisotropic menisci by 3D-bioprinting protein-releasing cell-laden hydrogel-polymer composite scaffold. Applied Materials Today. 2020; 18:100469. https://doi.org/10.1016/j.apmt.2019.100469
- 38. Lu J, Shen X, Sun X, Yin H, Yang S, Lu C, Wang Y, Liu Y, Huang Y, Yang Z, Dong X, Wang C, Guo Q, et al. Increased recruitment of endogenous stem cells and chondrogenic differentiation by a composite scaffold containing bone marrow homing peptide for cartilage regeneration. Theranostics. 2018; 8:5039–58. https://doi.org/10.7150/thno.26981 [PubMed]



