Introduction
Aging is a natural event, driven by multiple genetic and environmental factors and is also a gradual and irreversible process [1, 2]. Research on aging has made great progress. The reasons for aging are diverse, and their typical hallmarks are described as follows: genomic instability, telomere attrition, epigenetic alterations, loss of proteostasis, deregulated nutrient sensing, mitochondrial dysfunction, cell senescence, stem cell failure and altered intercellular communication [3, 4].
Proteostasis is maintained by a proteostasis network [5]. This network, which can sense and respond to misfolded proteins in all cells individually or together in different subnetworks, is composed of molecular chaperones, protein degradation mechanisms, and stress response pathways [6]. However, during aging, the ability of cells to maintain protein stability is weakened [7]. The molecular chaperone-mediated degradation of protein equilibrium mechanisms is disrupted, leading to increased protein oxidation, mis-location and aggregation in organisms [8]. The misfolded proteins cannot be degraded, and they accumulate in the body, eventually causing aging or aging-related diseases (such as Alzheimer's disease (AD) and Parkinson’s disease (PD)) in humans [9].
Protein is degraded via two primary pathways: the ubiquitin-proteasome pathway (UPS) and the autophagic lysosomal pathway [10]. According to the occurrence of autophagy, it includes three forms: macroautophagy, microautophagy, and molecular chaperone-mediated autophagy [11]. The main role of autophagy is to degrade many different substrates during metabolic stress to maintain the energy balance [12], and it is usually used to degrade cytoplasmic components, such as organelles and macromolecular compounds [13]. Lysosomes are an essential catalytic part of the autophagic lysosomal degradation system [14]. The lysosome contains a large number of hydrolases that are involved in the degradation of proteins, which are decomposed into small peptides and amino acids to obtain energy or synthesize new proteins [15]. The autophagic lysosomal pathway maintains protein homeostasis through this mechanism of degradation.
The ubiquitin-proteasome pathway is the major selective protein degradation pathway in eukaryotic cells [13]. The UPS can mediate the degradation of short-lived regulatory proteins, ensuring the orderly progression of related biological processes and promoting the degradation of defective proteins [10, 16]. The ubiquitin-proteasome pathway consists of a protein recruiting pathway and a protein degradation pathway. The first part consists of three enzymes: ubiquitin-activating enzyme (E1), ubiquitin-conjugation enzymes (E2), and ubiquitin ligase enzymes (E3). As a result of the cascade of three ubiquitinases, ubiquitin eventually binds to the substrate protein. Protein degradation is mainly accomplished by the proteasome, where the substrate protein is recognized and degraded by the proteasome [5].
Before the ubiquitin cascade forms a sufficiently large branched structure to activate the proteasome, the deubiquitinating enzymes (DUBs) can remove the ubiquitin molecule, reversing the ubiquitination process and maintaining protein stability; this process protects the protein from degradation [17].
The Drosophila ubiquitin-specific protease 7 (dUSP7) is one of the most abundant deubiquitinating proteases. Many reports had proved that dUSP7 is closely related to various biological processes. dUSP7 plays a very important role in DNA repair, Tip60-mediated apoptosis, various cancers, and organism development [18–21]. Besides, previous studies have revealed that Usp7 is essential for some organs development and size, such as wing development [14], brain size [22], eye development [23].
Only a few studies on the role of dUSP7 in aging have been reported. In C. elegans, the lack of math-33, a homologous protein of dUSP7, can reduce the lifespan of C. elegans [24]. To explore whether the regulatory mechanism of dUSP7 in aging is conserved, we examined the aging effects of dUSP7 in Drosophila. As the signaling pathways that regulate aging are evolutionarily conserved, our results could further elucidate the mechanism of dUSP7 in the regulation of aging.
Results
Knockdown of dusp7 significantly shortens the Drosophila lifespan
Using classical Gal4 driver and GeneSwitch system of Drosophila to design a gene which is able to be time and tissue-specific expression (Figure 1A). CG1490 RNAi is with usp7-RNAi gene strains of Drosophila. To explore the effect of ubiquitin-specific protease dUSP7 on the lifespan of Drosophila, we first tested the mRNA levels of dusp7 when dusp7 was knocked down by DaGS-gal4. The results showed that the expression of dusp7 was significantly decreased compared with the control group (Figure 1C). When dusp7 was knocked down in Drosophila, the lifespan was significantly decreased in comparison with the control group. The decrease was up to approximately 28.9%, with the mean lifespan decreasing from 69.1 to 52.1 days, p<0.0001 (Figure 1A). When we knocked down the expression of dusp7 only in Drosophila guts, the lifespan was also significantly decreased compared with the control group by up to approximately 13.3%, with the mean lifespan decreasing from 58.7 to 52.7 days, p<0.0001 (Figure 1B). These results indicate that dUSP7 is necessary to maintain the lifespan in Drosophila.
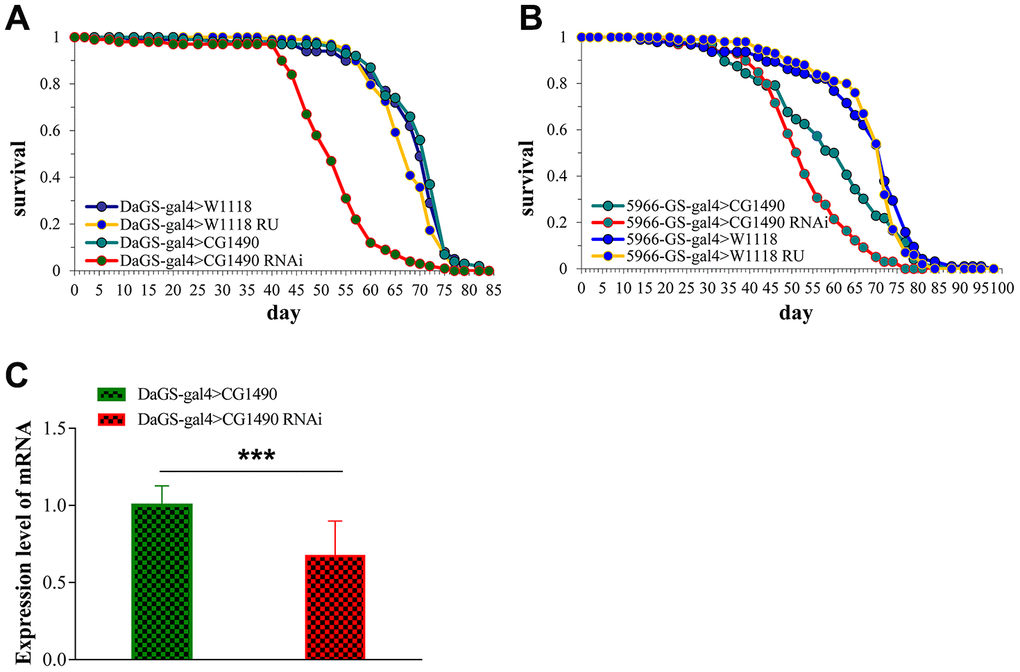
Figure 1. The effect of dusp7 knockdown on the lifespan of Drosophila. (A) The dusp7 expression level when RU486 was added to induce dusp7 knockdown (*** p<0.001; > means hybridization). (B) The lifespan curve when dusp7 was knocked down in Drosophila. (C) The lifespan curve under specific knockdown of dusp7 in the Drosophila intestinal track.
Knockdown of dusp7 significantly decreased the climbing ability of Drosophila
The climbing ability of Drosophila weakens as the Drosophila age. To explore the climbing ability of Drosophila upon dusp7 knockdown, we tested the climbing ability of 10-day-old and 30-day-old Drosophila. For the 10-day-old flies, the climbing ability did not change between the control and knockdown groups; however, for the 30-day-old flies, the climbing ability was significantly decreased compared with the control group. The climbing ability performance index (PI) decreased from 0.515 to 0.285, p<0.05 (Figure 2A). We also gut-specifically knocked down the dusp7 gene in Drosophila, and the climbing ability was significantly decreased compared with the control group, decreasing the PI from 0.29 to 0.17, p<0.05 (Figure 2B). These findings indicate that dusp7 can reduce the muscle mass and cause the flies to weaken, thereby decreasing the climbing ability.
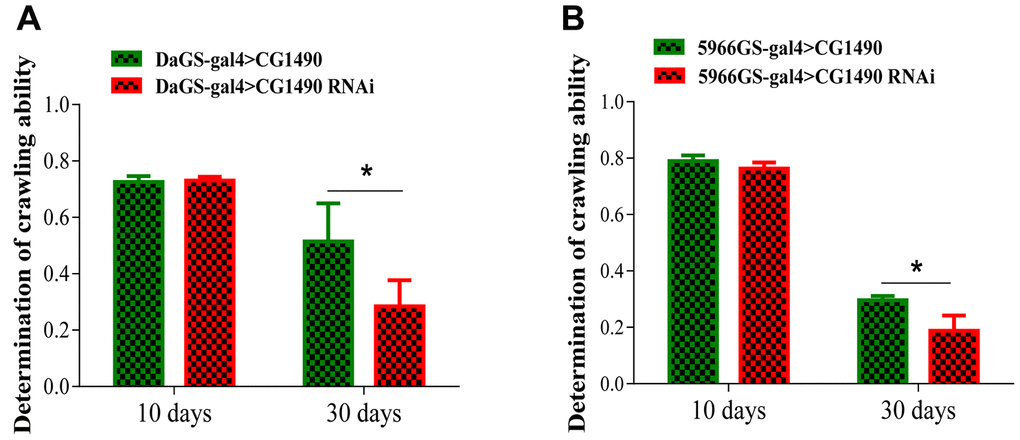
Figure 2. Knockdown of dusp7 decreases the climbing ability of Drosophila. (A) Effect on the climbing ability of Drosophila after dusp7 knockdown (* p<0.05). (B) Effect on the climbing ability of Drosophila after dusp7 knockdown in the gut (* p<0.05). (>: hybridization).
Knockdown of dusp7 promotes gut dysfunction in Drosophila
As gut function integrity is closely related to aging, it is important for the health of the organism. Therefore, we investigated whether dusp7 acts on the gut health of Drosophila. From the results of the “smurf” assay [25], we found that systemic knockdown and gut-specific knockdown of dusp7 significantly increased the proportion of “smurfs” in 40-day-old females compared with controls (p<0.05; Figure 3A, 3B), indicating that dusp7 knockdown reduces the gut barrier function.
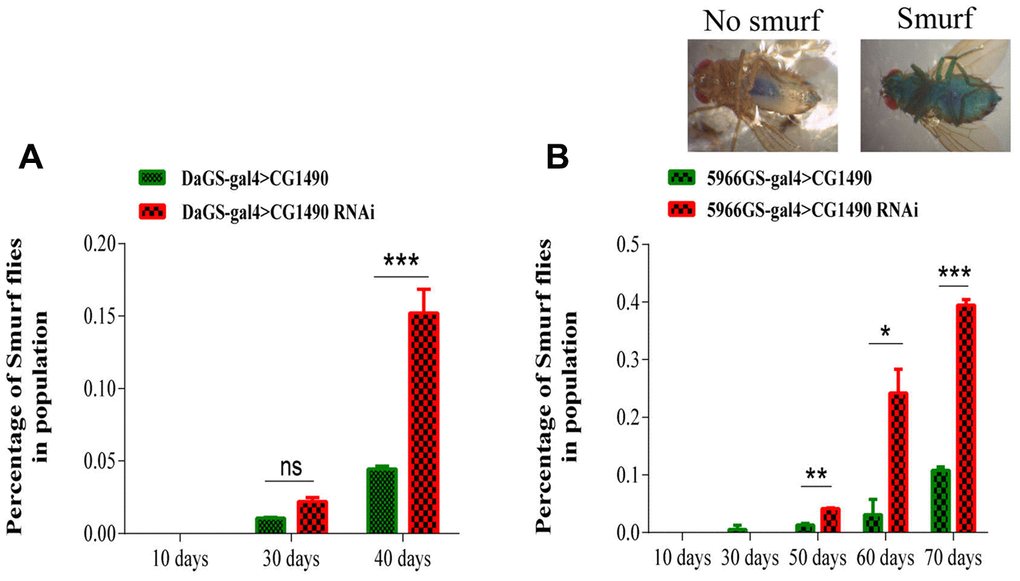
Figure 3. Effect of dusp7 knockdown on the intestinal function of Drosophila. (A) Effect on the intestinal integrity of Drosophila after systemic knockdown of the ubiquitin-specific protease dUSP7(*** p<0.001). (B) Effect on intestinal integrity of Drosophila after gut-specific knockdown of the ubiquitin-specific protease dUSP7 (* p<0.05; ** p<0.01; *** p<0.001). (>: hybridization).
dUSP7 is necessarily for the stress tolerance of Drosophila
Because stress tolerance and longevity are mechanistically and phenotypically linked [26], we investigated changes in the stress tolerance of Drosophila by knocking down dusp7. The 10-day-old flies were used to examine susceptibility to starvation, heat-shock, paraquat, and H2O2 stress. The results showed that flies with dusp7 knockdown (systemic or gut-specific) had significantly decreased tolerance to H2O2 compared with controls. The survival time for the systemic knockdown flies decreased by 23.08% (p<0.0001) and that of the gut-specific knockdown flies decreased by 10.5% (p<0.05) (Figure 4A, 4B). The tolerance to paraquat was similar to H2O2, where both systemic and gut-specific knockdown flies showed a significantly decreased the tolerance to paraquat compared with controls. The survival time for systemic knockdown decreased by up to 27.27% (p<0.0001), and gut-specific knockdown decreased by up to 10% (p<0.001) (Figure 4C, 4D). Moreover, we also validated the stress tolerance by using systemic overexpression (OE) dusp7 Drosophila (Supplementary Figure 1A)and the survival time showed a significantly increasing by up to 47.29%(p<0.0001) (Supplementary Figure 1B). Subsequently, we found that flies that were knocked down for dusp7 showed significantly decreased tolerance to starvation compared with controls (the survival time for systemic knockdown decreased by up to 25.37%, p<0.0001, and gut-specific knockdown decreased by up to 3.79%, p<0.01, Figure 4E, 4F). We also tested for heat-shock tolerance, and the results were the same as above. Flies knocked down for dusp7 (both systemic and gut-specific) showed significantly decreased tolerance to heat-shock compared with controls (the survival time for systemic knockdown decreased by up to 22.24%, p<0.001, and gut-specific knockdown decreased by up to 12.59%, p<0.01, Figure 4G, 4H). But there is no significant increasing in systemic overexpression dusp7 Drosophila (Supplementary Figure 1C). These results indicated that usp7 is necessary for the stress tolerance of Drosophila.
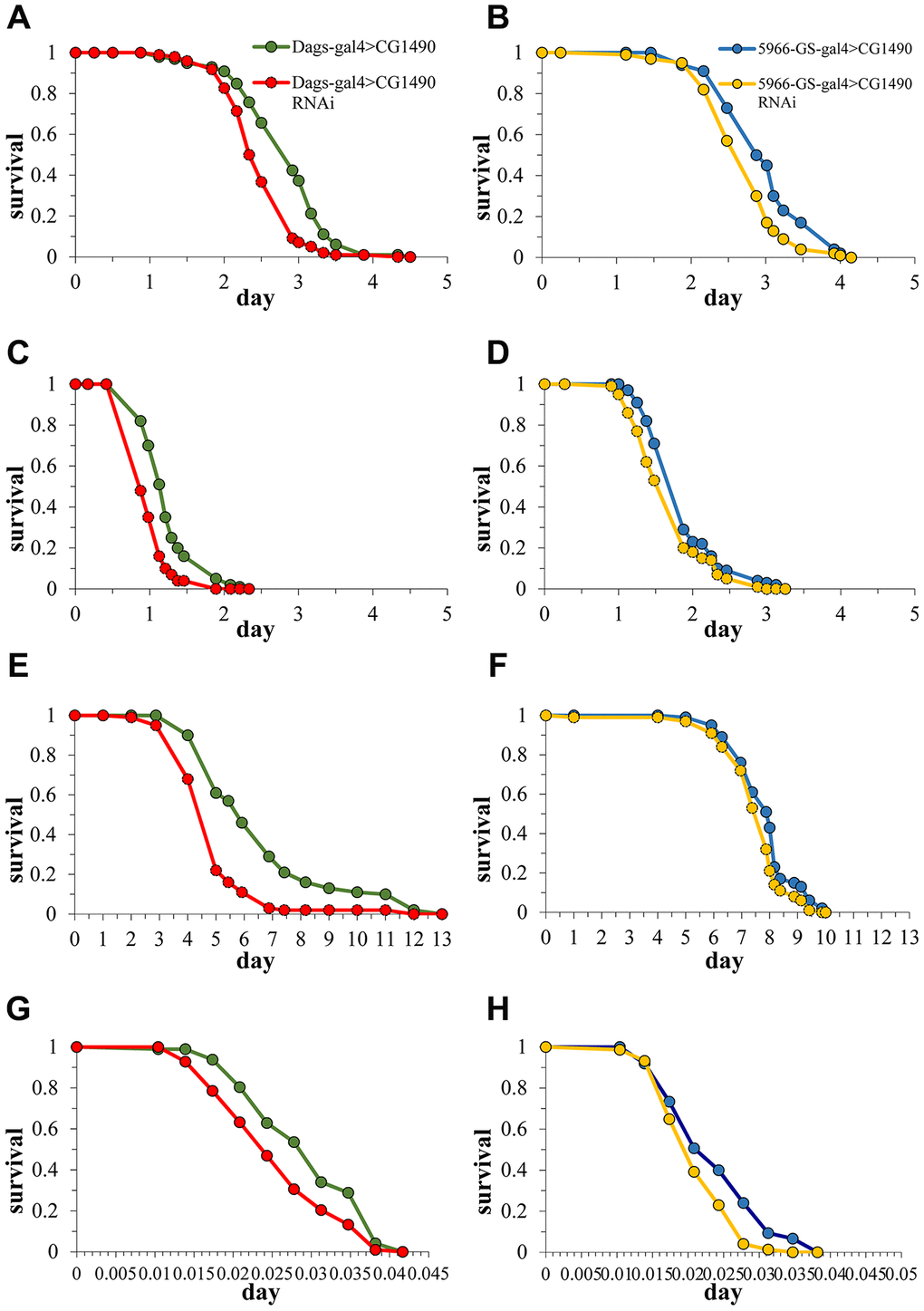
Figure 4. Effect of dusp7 knockdown on Drosophila stress tolerance. (A) Effect of dusp7 knockdown on Drosophila tolerance to H2O2. (B) Effect of gut-specific dusp7 knockdown on Drosophila tolerance to H2O2. (C) Effect of dusp7 knockdown on Drosophila tolerance to paraquat. (D) Effect of gut-specific dusp7 knockdown on Drosophila tolerance to paraquat. (E) Effect of dusp7 knockdown on Drosophila tolerance to starvation. (F) Effect of gut-specific dusp7 knockdown on Drosophila tolerance to starvation. (G) Effect of dusp7 knockdown on Drosophila tolerance to heat stimulation. (H) Effect of dusp7 gut-specific knockdown on Drosophila tolerance to heat stimulation. (>: hybridization).
dusp7 regulates protein degradation and autophagy
Loss of protein homeostasis is considered to be one of the "hallmarks of aging" [3, 27]. Protein homeostasis is maintained by the protein stabilization network (PN), which is composed of molecular chaperones, protein degradation mechanisms, and stress response pathways [6]. When dusp7 was knocked down, we found that knockdown flies significantly increased their ubiquitination of proteins and monoubiquitin (Figure 5A, 5B). The dusp7 knockdown flies also significantly decreased their protein and mRNA expression levels of autophagy-related 5 (atg5) (Figure 5C, 5D). Additionally, we detected a decrease of autolysosome in their midgut epithelial cells (Figure 5E). These results indicated that dusp7 may regulate protein degradation and autophagy to some extent.
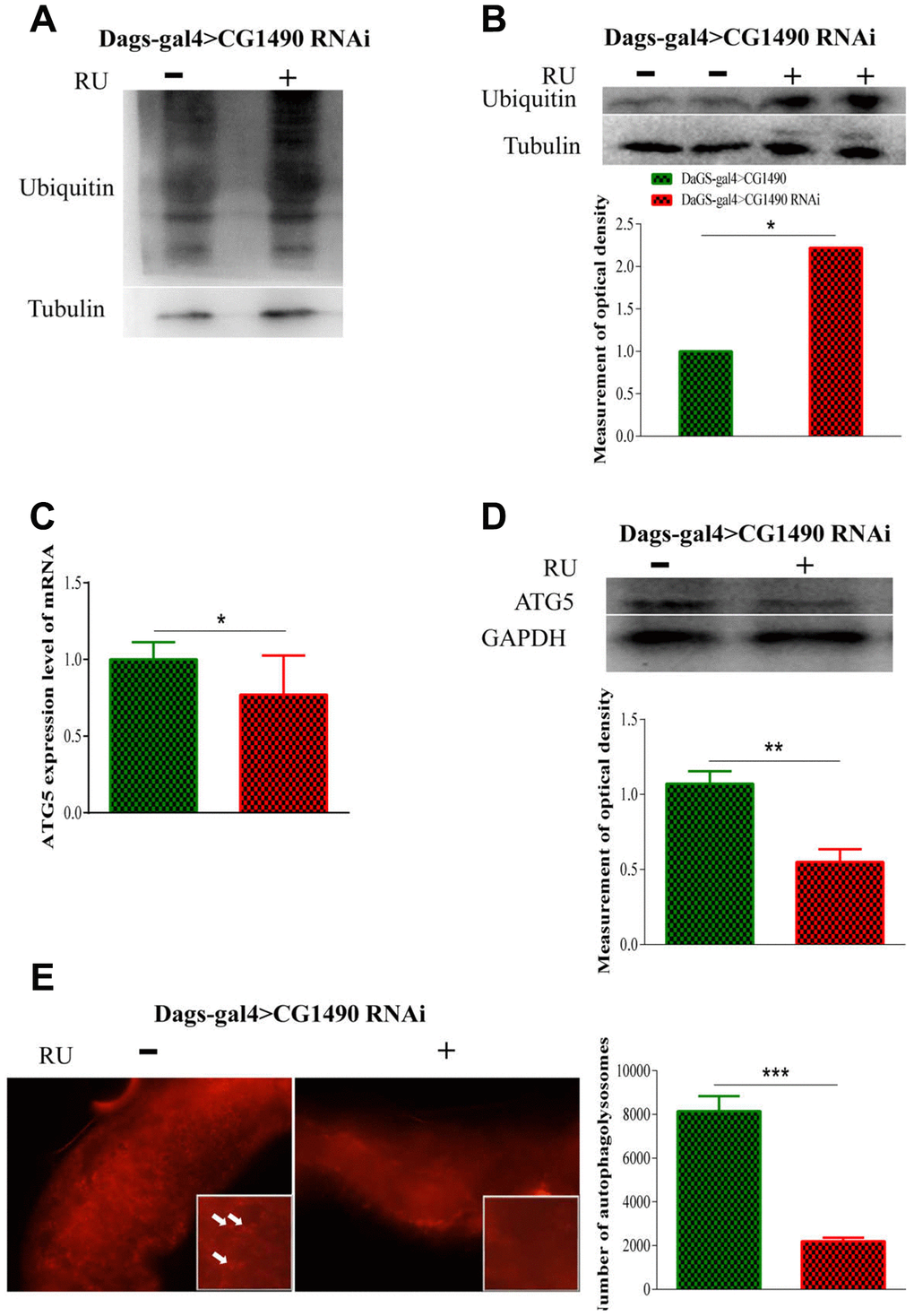
Figure 5. Effect of dusp7 in Drosophila on the control of protein in vivo. (A) The protein ubiquitination level after dusp7 knockdown in Drosophila. (B) The ubiquitination level after dusp7 knockdown in Drosophila (* p<0.05). (C) Autophagy-related gene expression after dusp7 knockdown in Drosophila (* p<0.05). (D) Autophagy-related protein expression after dusp7 knockdown in Drosophila (** p<0.01). (E) The effect of autolysosomes after dusp7 knockdown in Drosophila (*** p<0.001; white arrow points to autophagosome). (>: hybridization; + means adding Ru to induce expression; - means no adding RU).
DMC extends the lifespan of dusp7-knockdown drosophila
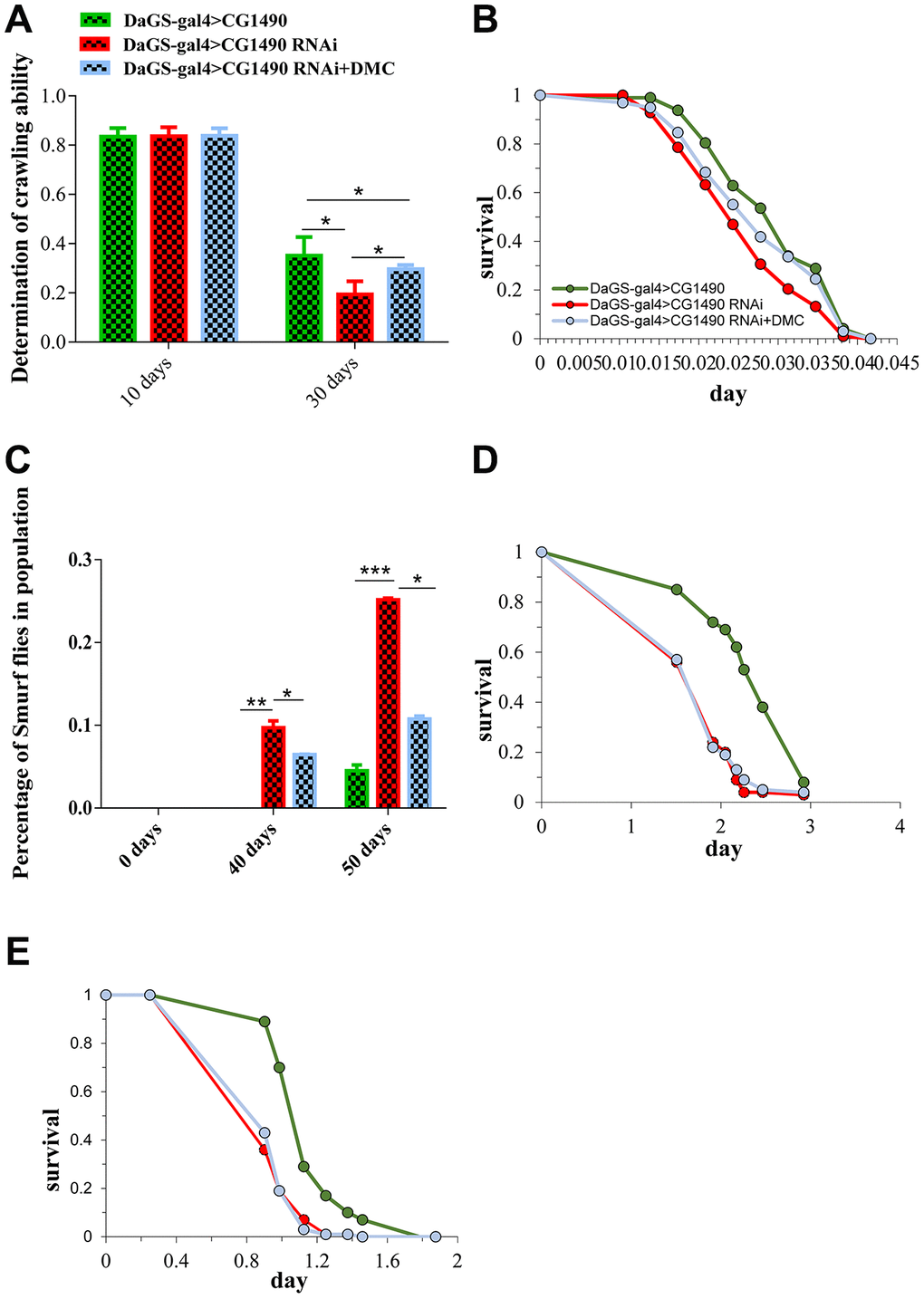
2,5-Dimethyl celecoxib (DMC) is a derivative of celecoxib, a non-steroidal anti-inflammatory drug that inhibits colon cancer and induces apoptosis [28, 29]. We previously found that DMC can extend the lifespan of Drosophila by increasing autophagy in the cell [30]. Therefore, we tested whether the addition of DMC could rescue the shortened lifespan of Drosophila caused by the knockdown of dusp7. The results showed that DMC was able to rescue the shortened lifespan caused by dusp7 knockdown (Figure 6A). Compared with the dusp7-knockdown group, the lifespan of the DMC-added group was significantly increased by approximately 6.36%, from 66 days to 70.2 days, p<0.0001.
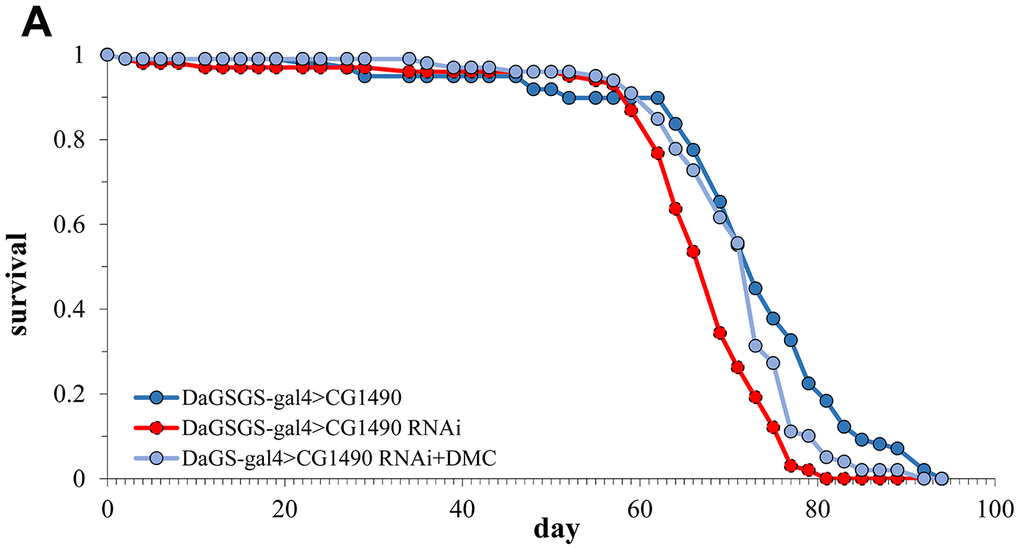
Figure 6. Lifespan of dusp7-knockdown group after adding DMC (A) Effect on the lifespan of adding DMC to dusp7-knockdown Drosophila (>: hybridization).
DMC improves autophagy and ubiquitination in dusp7-knockdown Drosophila
To test the effect of DMC on the protein degradation pathway, we investigated changes in the levels of protein ubiquitination, ubiquitin, autophagy, and atg5 mRNA in Drosophila. We found that dusp7-knockdown flies had significantly increased levels of protein ubiquitination and ubiquitin protein, but the addition of DMC decreased the level of ubiquitin protein (Figure 8A). Knockdown of dusp7 significantly decreased the protein and mRNA expression levels of atg5, but the addition of DMC increased these levels (Figure 8B, 8C). Additionally, DMC could increase autolysosomes in the midgut epithelial cells (Figure 8D). The results suggest that DMC can enhance autophagy and ubiquitination.
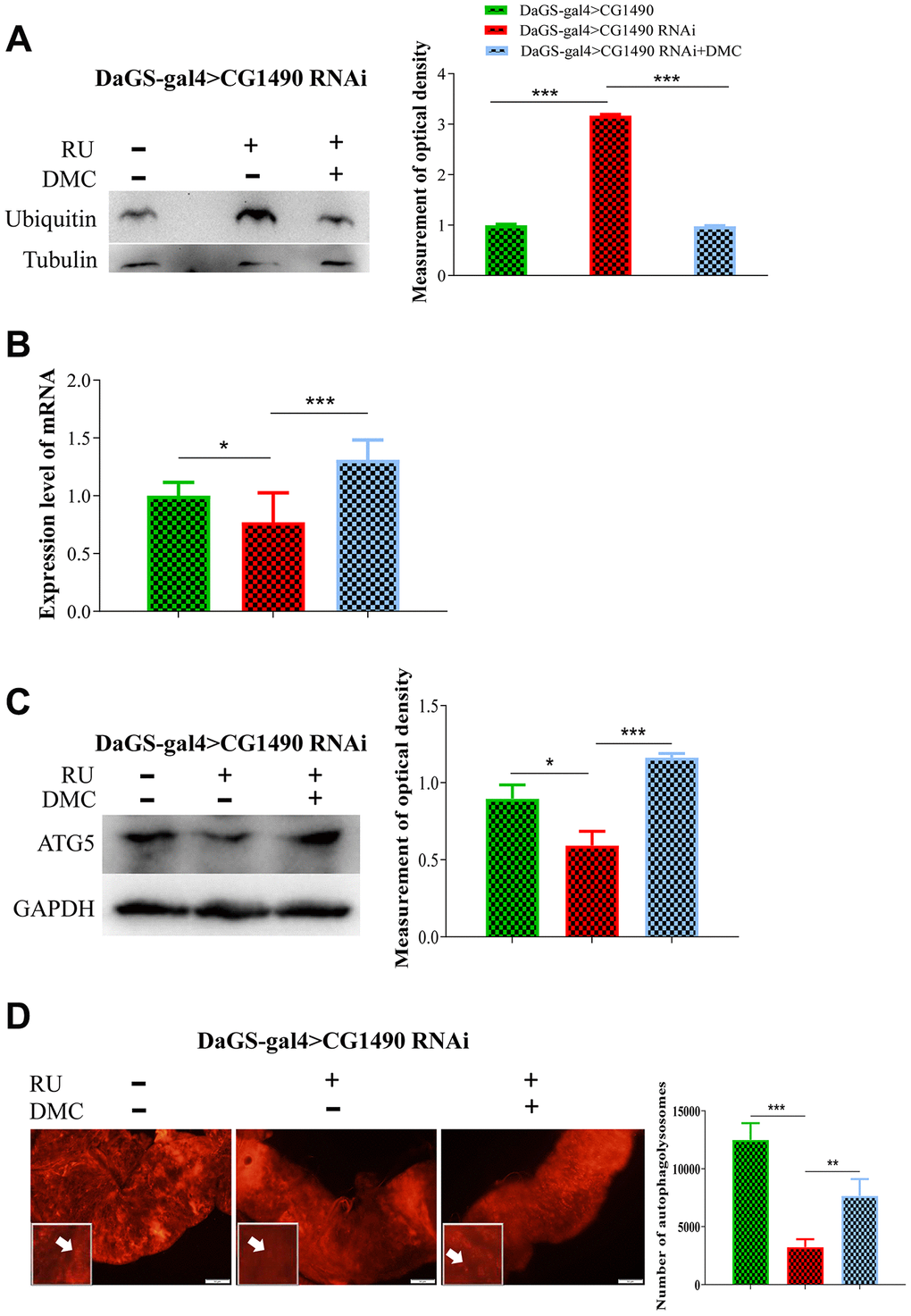
Figure 8. Effect of DMC addition on protein degradation in dusp7-knockdown Drosophila. (A) Expression level of ubiquitin protein (*** p<0.001). (B) Expression level of autophagy-associated gene (* p<0.05; ***p<0.001). (C) Expression level of autophagy-associated protein (* p<0.05; *** p<0.001). (D) Number of autolysosomes (** p<0.01; *** p<0.001; white arrow points to autophagosome). (>: hybridization; + means adding corresponding ingredients; - means no adding corresponding ingredients).
Discussion
USP7 is an evolutionarily conserved deubiquitinase with high homology between different species [31]. In our study, we found that dUSP7 is necessary to maintain the normal fly lifespan, and flies knocked down for dusp7 had significantly shortened lifespans and a reduced ability to respond to starvation, oxidative stress and heat stress. In addition, we found that USP7 regulates aging in relation to the autophagy and ubiquitin signaling pathways. Moreover, DMC can partially restore the shortened lifespan and relative aberrant phenotypes caused by dusp7 knockdown.
The dUSP7 homolog MATH-33 has been shown to regulate lifespan in nematodes by deubiquitinating DAF16 [24]. Our results showed that dUSP7 can also regulate the lifespan of Drosophila. When systemic or gut-specific dusp7 expression in Drosophila was knocked down, the lifespan was significantly decreased compared with control groups. This result is consistent with the phenotype of mutant math-33 in nematodes. With aging, the body's stress tolerance, intestinal function and muscle capacity decline [30, 32]. We found that the systemic knockdown of dusp7 gave rise to more accumulated aging characteristics than gut-specific dusp7 knockdown. This suggests that dusp7 is expressed systemically in Drosophila. When knockdown occurred only in the intestine, dusp7 could also work in other parts of the body, and it could continue to play a compensatory role. Thus, the aging symptoms of Drosophila were not as serious under gut-specific dusp7 knockdown compared to systemic knockdown.
Two pathways have been shown to regulate proteostasis that have been proven to be associated with aging. The first is the ubiquitin-proteasome pathway, which is mainly involved in the regulation of short-lived proteins and regulatory proteins [33]. The second is the autophagic lysosomal pathway, which is mainly involved in the regulation of long-lived proteins [34]. The two pathways work together to keep proteins relatively stable in the body [13]. In our study, when dusp7 was knocked down, the level of ubiquitination was significantly increased compared with the control. These results indicate that the protein homeostasis is disrupted and the ubiquitinated proteins are increased due to dusp7 knockdown. Thomas et al. found that MATH-33 removes ubiquitin moieties from DAF-16 to promote DAF-16 stability in response to decreased IIS in C. elegans [24]. The human USP7 stabilizes GATA1 by removing K48-linked polyubiquitin [35]. In response to DNA damage, BRE/BRCC45 recruits USP7 to regulate CDC25A stability [36]. These reports demonstrate the significance of USP7 in maintaining protein homeostasis. We also found that the level of autophagy was significantly reduced compared with control. We observed that knockdown of dusp7 reduces the ability of flies to cope with stress such as starvation and oxidative damage. Therefore, knockdown of dusp7 could lead to loss of proteostasis in Drosophila, possibly leading to a reduced lifespan.
DMC, a celecoxib derivative that has lost the ability to inhibit cyclooxygenase-2 (COX-2), does not cause the side effects of celecoxib; hence, it is used as a celecoxib replacement [28, 29, 37]. It has been reported that DMC can extend the lifespan of wild-type Drosophila by activating foxo, which is a key factor of the insulin-signaling pathway [30]. However, no reports have yet shown that DMC can restore the lifespan reduction caused by losing protein homeostasis. In our study, we found that DMC could rescue the shortened lifespan of Drosophila caused by dusp7 knockdown. The protein and mRNA expression levels of atg5 were significantly increased compared with the knockdown group that did not receive DMC, suggesting that the addition of DMC could enhance autophagy. Furthermore, we found that the addition of DMC not only enhances autophagy but also reduces the levels of ubiquitination and ubiquitin protein that are abnormally elevated in dusp7-knockdown Drosophila. Therefore, we suggest that DMC possibly regulates proteostasis to extend the lifespan of dusp7-knockdown Drosophila. DMC can rescue parts of the senescence traits of Drosophila, but it has no effect on the antioxidation ability. This indicates that DMC may extend the lifespan independent from the increase of antioxidant capacity.
In conclusion, dUSP7 is an essential component for maintaining the normal lifespan of Drosophila. Knockdown of dusp7 could significantly reduce the lifespan, lessen the climbing ability, reduce the stress tolerance and accelerate the loss of intestinal barrier integrity. At the same time, the level of ubiquitin increased and the level of autophagy decreased in Drosophila. Finally, the protein homeostasis balance in the body was upset. The insulin signaling pathway inhibitor DMC could partially rescue the most aberrant phenotypes, suggesting that dusp7 maintains the normal lifespan and partially depends on the insulin-signaling pathway in Drosophila. Due to the high conservation of the deubiquitinase dUSP7 across different species, it is expected that the regulatory pathway in aging also could be evolutionarily conserved. These results could provide insights regarding the role of human USP7 in aging and the potential development of targeted drugs.
Materials and Methods
Fly stocks and feeding
DaGS-gal4, 5966GS-gal4, usp7-RNAi (CG1490, BL34708), and W1118 were obtained from the Bloomington Drosophila Stock Center (http://flystocks.bio.indiana.edu), usp7-overexpression line (kindly gifted by Professor Zhou). The Drosophila were fed with either yeast medium or amino acid medium [38, 39]. The resulting hybrid, DaGS-CG1490 OE (Over Expression), DaGS-CG1490 RNAi, 5966GS-W1118, and 5966GS-CG1490 RNAi, was used in experiment.
Lifespan curve analysis
The survival time was recorded for the selected Drosophila. During the experiment, the culture medium of Drosophila was replaced every Monday, Wednesday and Friday until the experimental Drosophila had all died, and a survival curve was constructed according to the Drosophila survival data. Statistical significance analysis was performed using the log-rank test. More than two independent experiments were carried out for most of the lifespan assays.
Intestinal integrity test of Drosophila
Blue dye No. 1 was added to the culture medium for the intestinal integrity tests. When the Drosophila were maintained on the dyed medium for 9 hours, if the intestinal function was damaged, the dye entered the body through the intestine, making the whole abdomen and even the whole body blue. At this point, the Drosophila are termed "smurfs". In contrast, if the intestinal tract function was intact, the Drosophila showed no change.
Drosophila stress index
Each group of experiments required 200 flies that had been fed for 10 days. For the H2O2 and paraquat assays, Drosophila were fed 20 mM paraquat or 5% H2O2 diluted in a 5% glucose solution supplied on filter paper. In the starvation test, the Drosophila were placed in a tube containing 1.5% agarose to provide only moisture. The number of deaths was recorded every four hours in the above experiments. For the heat assay, Drosophila were placed in empty tubes at 39° C, and deaths were recorded every hour.
Drosophila climbing ability
A Drosophila was placed in a tube, which was gently shaken so that the fruit fly went into the bottom of the tube. The Drosophila climbed freely for 45 s and was then photographed. The assay was repeated three times with independent groups of Drosophila. The speed was calculated by the formula: PI=(ntot+ntop-nbot)/2/ntot (ntot indicates all Drosophila, ntop indicates the Drosophila at the top of the tube, nbot indicates the Drosophila at the bottom of the tube, and PI stands for climbing ability).
Western blots
Fruit fly total protein was extracted with a Cell Total Protein Extraction Kit (Sangon Biotech C510003). Aliquots of 40-100 μg protein extract were separated by gel electrophoresis. The proteins were then transferred to polyvinylidene fluoride membranes. The primary antibodies used were as follows: ATG-5 antibody (NOVUS NB110-53818SS), UB antibody (Cell Signaling Technology 3936S), α-tubulin antibody (Sigma T9026), and GAPDH antibody (Servicebio GB11002). All western blot experiments were repeated at least twice.
qRT-PCR
Drosophila cultured for 10 days were frozen on ice, and total RNA was extracted by the TRIzol method and converted into cDNA. The real-time PCR program was performed by the SYBR Green method using the following primers: rp49-F: AGATCGTGAAGAAGCGCACCAAG; rp49-R: CACCAGGAACTTCTTGAATCCGG. atg5-F: TAGGCATATGCTTCCAGGCG; atg5-R: CACAGCTCCATCCTGGTGTT. dusp7-F: GTGAGAATACCTTGGCGGACTACG; dusp7-R: GCGGCAACTGGAGACCACATC. The relative expression levels of the target genes were normalized to the expression level of the housekeep gene rp49, and all of the experiments were repeated three times.
Autolysosome staining analysis
The experimental Drosophila were cultured for 20 days, and the guts were extracted and stained with 1 μM LysoTracker DS Red DND-99 (Invitrogen, Molecular Probes) for 3 minutes and then washed 3 times with PBS before mounting and performing microscopy.
Quantification and statistical analysis
The measurements represent the mean of at least three biological replicates in all graphs, and error bars represent the standard deviation. As appropriate, two-tailed Student’s t-test or one-way ANOVA with post hoc Dunnett’s test was used to calculate significance. All statistical analyses were carried out using GraphPad Prism 6.00 software. For all tests, p<0.05 was considered to be statistically significant. Asterisks reflecting the calculated p-values are shown above each measurement, and ns indicates that differences between measurements were not statistically significant. No collected data were excluded from any experimental or statistical analysis. Tests for normality or outliers were not performed.
Supplementary Materials
Author Contributions
L.C, W.S, X.F. and M.Y. designed and conducted the experiment with fly lifespan, “SMURF”, and analyzed data; Y.Z. and Q.W. conducted the experiments with Western Blots, analyzed data; W.S. T.L.and Y.L conducted the experiments With qRT-PCR and analyzed data; B.Z. and T.W conducted the experiments stress assay and analyzed data. X.M., D.Y., M.Z., Q.N. and D.L. participated in discussions. L.C, X.F.and M.Y. contributed to writing the manuscript and participated in discussions.
Acknowledgments
We thank all lab members for their valuable comments. We thank the Bloomington Drosophila Stock Center for the fly strains.
Conflicts of Interest
The authors declare no conflicts of interest.
Funding
This work was supported by the National Natural Science Foundation of China (81701392, 31771338).
References
- 1. Barbosa MC, Grosso RA, Fader CM. Hallmarks of aging: an autophagic perspective. Front Endocrinol (Lausanne). 2019; 9:790. https://doi.org/10.3389/fendo.2018.00790 [PubMed]
- 2. Sopjani M, Rinnerthaler M, Kruja J, Dermaku-Sopjani M. Intracellular signaling of the aging suppressor protein Klotho. Curr Mol Med. 2015; 15:27–37. https://doi.org/10.2174/1566524015666150114111258 [PubMed]
- 3. López-Otín C, Blasco MA, Partridge L, Serrano M, Kroemer G. The hallmarks of aging. Cell. 2013; 153:1194–217. https://doi.org/10.1016/j.cell.2013.05.039 [PubMed]
- 4. Singh PP, Demmitt BA, Nath RD, Brunet A. The genetics of aging: a vertebrate perspective. Cell. 2019; 177:200–20. https://doi.org/10.1016/j.cell.2019.02.038 [PubMed]
- 5. Shang F, Taylor A. Ubiquitin-proteasome pathway and cellular responses to oxidative stress. Free Radic Biol Med. 2011; 51:5–16. https://doi.org/10.1016/j.freeradbiomed.2011.03.031 [PubMed]
- 6. Balch WE, Morimoto RI, Dillin A, Kelly JW. Adapting proteostasis for disease intervention. Science. 2008; 319:916–19. https://doi.org/10.1126/science.1141448 [PubMed]
- 7. Kumsta C, Chang JT, Lee R, Tan EP, Yang Y, Loureiro R, Choy EH, Lim SH, Saez I, Springhorn A, Hoppe T, Vilchez D, Hansen M. The autophagy receptor p62/SQST-1 promotes proteostasis and longevity in C. Elegans by inducing autophagy. Nat Commun. 2019; 10:5648. https://doi.org/10.1038/s41467-019-13540-4 [PubMed]
- 8. Vila M, Laguna A, Carballo-Carbajal I. Intracellular crowding by age-dependent neuromelanin accumulation disrupts neuronal proteostasis and triggers Parkinson disease pathology. Autophagy. 2019; 15:2028–30. https://doi.org/10.1080/15548627.2019.1659621 [PubMed]
- 9. Koga H, Kaushik S, Cuervo AM. Protein homeostasis and aging: the importance of exquisite quality control. Ageing Res Rev. 2011; 10:205–15. https://doi.org/10.1016/j.arr.2010.02.001 [PubMed]
- 10. Löw P. The role of ubiquitin-proteasome system in ageing. Gen Comp Endocrinol. 2011; 172:39–43. https://doi.org/10.1016/j.ygcen.2011.02.005 [PubMed]
- 11. Rabinowitz JD, White E. Autophagy and metabolism. Science. 2010; 330:1344–48. https://doi.org/10.1126/science.1193497 [PubMed]
- 12. Rubinsztein DC, Mariño G, Kroemer G. Autophagy and aging. Cell. 2011; 146:682–95. https://doi.org/10.1016/j.cell.2011.07.030 [PubMed]
- 13. Vilchez D, Saez I, Dillin A. The role of protein clearance mechanisms in organismal ageing and age-related diseases. Nat Commun. 2014; 5:5659. https://doi.org/10.1038/ncomms6659 [PubMed]
- 14. Zhou Z, Yao X, Li S, Xiong Y, Dong X, Zhao Y, Jiang J, Zhang Q. Deubiquitination of ci/gli by Usp7/HAUSP regulates hedgehog signaling. Dev Cell. 2015; 34:58–72. https://doi.org/10.1016/j.devcel.2015.05.016 [PubMed]
- 15. Mizushima N, Komatsu M. Autophagy: renovation of cells and tissues. Cell. 2011; 147:728–41. https://doi.org/10.1016/j.cell.2011.10.026 [PubMed]
- 16. Schmidt M, Finley D. Regulation of proteasome activity in health and disease. Biochim Biophys Acta. 2014; 1843:13–25. https://doi.org/10.1016/j.bbamcr.2013.08.012 [PubMed]
- 17. Haq S, Ramakrishna S. Deubiquitylation of deubiquitylases. Open Biol. 2017; 7:170016. https://doi.org/10.1098/rsob.170016 [PubMed]
- 18. Zlatanou A, Sabbioneda S, Miller ES, Greenwalt A, Aggathanggelou A, Maurice MM, Lehmann AR, Stankovic T, Reverdy C, Colland F, Vaziri C, Stewart GS. USP7 is essential for maintaining Rad18 stability and DNA damage tolerance. Oncogene. 2016; 35:965–76. https://doi.org/10.1038/onc.2015.149 [PubMed]
- 19. Masuya D, Huang C, Liu D, Nakashima T, Yokomise H, Ueno M, Nakashima N, Sumitomo S. The HAUSP gene plays an important role in non-small cell lung carcinogenesis through p53-dependent pathways. J Pathol. 2006; 208:724–32. https://doi.org/10.1002/path.1931 [PubMed]
- 20. Zhao GY, Lin ZW, Lu CL, Gu J, Yuan YF, Xu FK, Liu RH, Ge D, Ding JY. USP7 overexpression predicts a poor prognosis in lung squamous cell carcinoma and large cell carcinoma. Tumour Biol. 2015; 36:1721–29. https://doi.org/10.1007/s13277-014-2773-4 [PubMed]
- 21. Kon N, Zhong J, Kobayashi Y, Li M, Szabolcs M, Ludwig T, Canoll PD, Gu W. Roles of HAUSP-mediated p53 regulation in central nervous system development. Cell Death Differ. 2011; 18:1366–75. https://doi.org/10.1038/cdd.2011.12 [PubMed]
- 22. van der Knaap JA, Kumar BR, Moshkin YM, Langenberg K, Krijgsveld J, Heck AJ, Karch F, Verrijzer CP. GMP synthetase stimulates histone H2B deubiquitylation by the epigenetic silencer USP7. Mol Cell. 2005; 17:695–707. https://doi.org/10.1016/j.molcel.2005.02.013 [PubMed]
- 23. Sun X, Ding Y, Zhan M, Li Y, Gao D, Wang G, Gao Y, Li Y, Wu S, Lu L, Liu Q, Zhou Z. Usp7 regulates hippo pathway through deubiquitinating the transcriptional coactivator yorkie. Nat Commun. 2019; 10:411. https://doi.org/10.1038/s41467-019-08334-7 [PubMed]
- 24. Heimbucher T, Liu Z, Bossard C, McCloskey R, Carrano AC, Riedel CG, Tanasa B, Klammt C, Fonslow BR, Riera CE, Lillemeier BF, Kemphues K, Yates JR
3rd , et al. The deubiquitylase MATH-33 controls DAF-16 stability and function in metabolism and longevity. Cell Metab. 2015; 22:151–63. https://doi.org/10.1016/j.cmet.2015.06.002 [PubMed] - 25. Fan X, Liang Q, Lian T, Wu Q, Gaur U, Li D, Yang D, Mao X, Jin Z, Li Y, Yang M. Rapamycin preserves gut homeostasis during Drosophila aging. Oncotarget. 2015; 6:35274–83. https://doi.org/10.18632/oncotarget.5895 [PubMed]
- 26. Foley HB, Sun PY, Ramirez R, So BK, Venkataraman YR, Nixon EN, Davies KJ, Edmands S. Sex-specific stress tolerance, proteolysis, and lifespan in the invertebrate tigriopus californicus. Exp Gerontol. 2019; 119:146–56. https://doi.org/10.1016/j.exger.2019.02.006 [PubMed]
- 27. Labbadia J, Morimoto RI. Proteostasis and longevity: when does aging really begin? F1000Prime Rep. 2014; 6:7. https://doi.org/10.12703/P6-7 [PubMed]
- 28. Egashira I, Takahashi-Yanaga F, Nishida R, Arioka M, Igawa K, Tomooka K, Nakatsu Y, Tsuzuki T, Nakabeppu Y, Kitazono T, Sasaguri T. Celecoxib and 2,5-dimethylcelecoxib inhibit intestinal cancer growth by suppressing the Wnt/β-catenin signaling pathway. Cancer Sci. 2017; 108:108–15. https://doi.org/10.1111/cas.13106 [PubMed]
- 29. Atari-Hajipirloo S, Nikanfar S, Heydari A, Kheradmand F. Imatinib and its combination with 2,5-dimethyl-celecoxibinduces apoptosis of human HT-29 colorectal cancer cells. Res Pharm Sci. 2017; 12:67–73. https://doi.org/10.4103/1735-5362.199049 [PubMed]
- 30. Wu Q, Lian T, Fan X, Song C, Gaur U, Mao X, Yang D, Piper MD, Yang M. 2,5-dimethyl-celecoxib extends drosophila life span via a mechanism that requires insulin and target of rapamycin signaling. J Gerontol A Biol Sci Med Sci. 2017; 72:1334–41. https://doi.org/10.1093/gerona/glw244 [PubMed]
- 31. van der Knaap JA, Kozhevnikova E, Langenberg K, Moshkin YM, Verrijzer CP. Biosynthetic enzyme GMP synthetase cooperates with ubiquitin-specific protease 7 in transcriptional regulation of ecdysteroid target genes. Mol Cell Biol. 2010; 30:736–44. https://doi.org/10.1128/MCB.01121-09 [PubMed]
- 32. Bjedov I, Toivonen JM, Kerr F, Slack C, Jacobson J, Foley A, Partridge L. Mechanisms of life span extension by rapamycin in the fruit fly Drosophila melanogaster. Cell Metab. 2010; 11:35–46. https://doi.org/10.1016/j.cmet.2009.11.010 [PubMed]
- 33. Sha Z, Zhao J, Goldberg AL. Measuring the Overall Rate of Protein Breakdown in Cells and the Contributions of the Ubiquitin-Proteasome and Autophagy-Lysosomal Pathways. Methods Mol Biol. 2018; 1844:261–276. https://doi.org/10.1007/978-1-4939-8706-1_17 [PubMed]
- 34. Yu L, Chen Y, Tooze SA. Autophagy pathway: cellular and molecular mechanisms. Autophagy. 2018; 14:207–15. https://doi.org/10.1080/15548627.2017.1378838 [PubMed]
- 35. Liang L, Peng Y, Zhang J, Zhang Y, Roy M, Han X, Xiao X, Sun S, Liu H, Nie L, Kuang Y, Zhu Z, Deng J, et al. Deubiquitylase USP7 regulates human terminal erythroid differentiation by stabilizing GATA1. Haematologica. 2019; 104:2178–87. https://doi.org/10.3324/haematol.2018.206227 [PubMed]
- 36. Biswas K, Philip S, Yadav A, Martin BK, Burkett S, Singh V, Babbar A, North SL, Chang S, Sharan SK. BRE/BRCC45 regulates CDC25A stability by recruiting USP7 in response to DNA damage. Nat Commun. 2018; 9:537. https://doi.org/10.1038/s41467-018-03020-6 [PubMed]
- 37. Sobolewski C, Rhim J, Legrand N, Muller F, Cerella C, Mack F, Chateauvieux S, Kim JG, Yoon AY, Kim KW, Dicato M, Diederich M. 2,5-dimethyl-celecoxib inhibits cell cycle progression and induces apoptosis in human leukemia cells. J Pharmacol Exp Ther. 2015; 355:308–28. https://doi.org/10.1124/jpet.115.225011 [PubMed]
- 38. Bass TM, Grandison RC, Wong R, Martinez P, Partridge L, Piper MD. Optimization of dietary restriction protocols in Drosophila. J Gerontol A Biol Sci Med Sci. 2007; 62:1071–81. https://doi.org/10.1093/gerona/62.10.1071 [PubMed]
- 39. Piper MD, Blanc E, Leitão-Gonçalves R, Yang M, He X, Linford NJ, Hoddinott MP, Hopfen C, Soultoukis GA, Niemeyer C, Kerr F, Pletcher SD, Ribeiro C, Partridge L. A holidic medium for Drosophila melanogaster. Nat Methods. 2014; 11:100–05. https://doi.org/10.1038/nmeth.2731 [PubMed]