Introduction
Coronavirus disease-19 (Covid-19) has challenged and transiently overwhelmed the health care system of all Western countries. Infection by severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2), the causative agent of Covid-19 is usually asymptomatic or pauci-symptomatic in healthy individuals [1, 2]. However, old age and age-related diseases are risk factors that predispose to a more severe disease course requiring hospitalization, respiratory assistance with oxygen (“moderate” Covid-19) or even intubation (“severe” Covid-19), often with long-term sequelae or a fatal outcome [3, 4]. Cancer is one of the established risk factors for severe Covid-19 [5], which may be related to the facts that (i) malignant disease mostly develops in aged individuals, often in the context of other comorbidities [6–8] (ii) cancer manifests preferentially in the context of failing immunosurveillance and mediates local and systemic immunosuppressive effects during tumor progression [9, 10]; and (iii) antineoplastic therapies, in particular combination chemotherapies, have debilitating, pro-inflammatory, immunosuppressive and senescence-accelerating side effects [11, 12].
Multiple studies have attempted to identify biomarkers that distinguish patients likely to develop mild versus severe Covid-19 [1, 2, 13]. Obviously, such biomarkers are linked to deficient anti-SARS-CoV-2 immune responses (such as severe lymphopenia or, more specifically, inefficient type-1 interferon responses due to inherent immune defects or the production of autoantibodies that neutralize type-1 interferons) [14–16] or excessive inflammatory responses (that often manifest at the levels of granulocytes, monocytes and their products) [17, 18]. The search of predictive biomarkers has been involving advanced technologies including high-dimensional cytometry [17], single-cell transcriptomics [19] and proteomics [20]. An additional unbiased strategy for defining circulating factors useful for risk stratification is mass spectrometric metabolomics, a technique that requires a minimum of initial sample preparation (snap freezing of serum or heparin serum and its storage at -80° C), is entirely automatable and yields accurate information on hundreds of known metabolites (i.e. a combination of chromatographic retention times and masses that allow for the >bona fide identification of the corresponding chemical compound) as well as thousands of unknown metabolites [21–32].
Here, we used mass spectrometric metabolomics to identify biomarkers of Covid-19 severity in cancer patients recovered at the Gustave Roussy Cancer Campus, which comprises the largest cancer-specific research hospital in Europe. We identified a series of metabolites that correlate with disease severity in cancer patients.
Results
Study design and metadata
We determined the metabolome of serum samples from 204 cancer patients enrolled in the ONCOVID trial (https://clinicaltrials.gov/ct2/show/NCT04341207). Patients were divided into “controls” (no diagnosis of Covid-19, no signs of respiratory infection) and two categories of PCR-confirmed Covid-19 patients with “mild disease” (with ambulatory treatment) or “moderate/severe disease” (requiring hospitalization and respiratory assistance). Indeed, the number of patients with severe Covid-19 (requiring intubation and mechanical ventilation) was too low to be analyzed as a separate group. Importantly, the duration of viral shedding (short-term shedding: <40 days, long-term shedding: >40 days) was variable among Covid-19 infected cancer patients, yet tended to be shorter in mild than in moderate/severe cases (Figure 1) as already observed by Goubet et al. [33]. The clinical characteristics of both cohorts are summarized in Table 1. Serum samples from both cohorts were subjected to mass spectrometric metabolomics, yielding high-quality information on 239 identified metabolites (Figure 1 and Supplementary Table 1). Moreover, the cohort yielded 4276 non-identified mass spectrometric peaks (Supplementary Figure 1 and Supplementary Table 2).
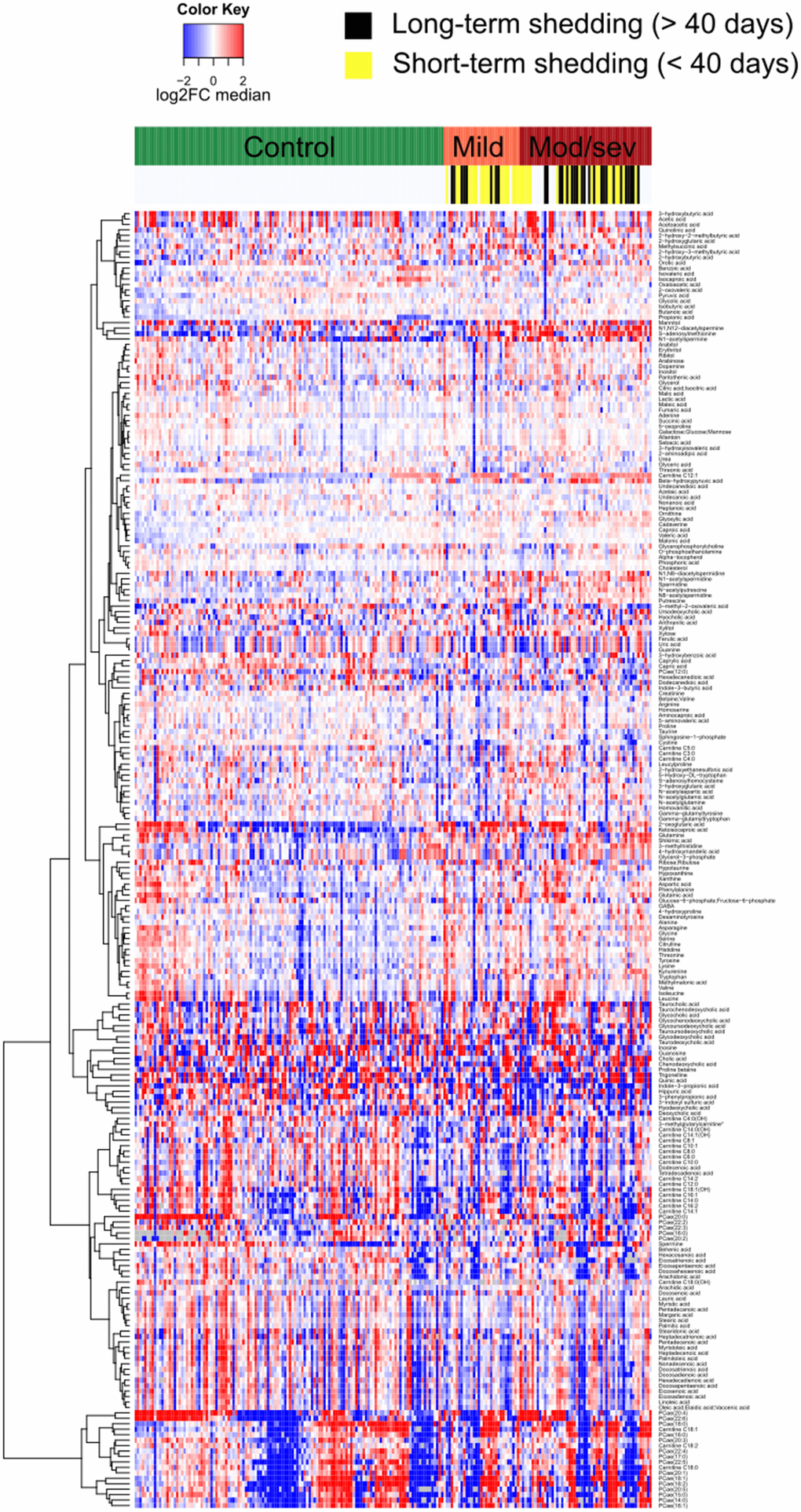
Figure 1. Heatmap representing the serum metabolome of each individual cancer patient clustered by clinical severity of Covid-19. Targeted metabolomic data on 211 serum samples from 204 patients were normalized areas of identified metabolites. Results are listed in Supplementary Table 1.
Table 1. Clinical characteristics of SARS-CoV-2 patients affected by cancer.
| Cancer patient characteristics | All N=204 | Control n=128 | COVID-19 | p | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Mild n=37 | Moderate or severe n=39 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Age (year) | Median (range) | 61 (19-89) | 60 (19-83) | 65 (19-89) | 64 (33-83) | - | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Gender – no (%) | Male | 84 (41) | 54 (42) | 12 (32) | 18 (46) | 0.44 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Female | 120 (59) | 74 (58) | 25 (68) | 21 (54) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Number of comorbidities – no (%) | 0 | 104 (51) | 71 (55) | 16 (43) | 17 (44) | 0.42 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 | 51 (25) | 27 (21) | 14 (38) | 10 (26) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 2 | 30 (15) | 18 (14) | 4 (11) | 8 (21) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| >3 | 19 (9) | 12 (9) | 3 (8) | 4 (10) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Comorbid conditions – no (%) | COPD | 15 (7) | 13 (10) | 1 (3) | 1 (3) | 0.51 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Obesity (BMI ≥ 30) | 18 (9) | 11 (9) | 2 (5) | 5 (13) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Hypertension | 66 (32) | 37 (29) | 15 (41) | 14 (36) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Congestive heart failure | 7 (3) | 4 (3) | 1 (3) | 2 (5) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Diabetes mellitus | 22 (11) | 11 (9) | 5 (14) | 6 (15) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Type of malignancy – no (%) | Solid tumors | 180 (88) | 121 (95) | 31 (84) | 28 (72) | 0.0004 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Hematological malignancies | 24 (12) | 7 (5) | 6 (16) | 11 (28) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cancer spread - no (%) | Localized | 89 (44) | 68 (53) | 11 (30) | 10 (26) | 0.002 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Locally advanced | 25 (12) | 11 (9) | 9 (24) | 5 (13) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metastatic | 90 (44) | 49 (38) | 17 (46) | 24 (61) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ECOG performance status – no (%) | 0 | 108 (53) | 81 (63) | 21 (57) | 6 (15) | <0.0001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 | 52 (25) | 33 (26) | 6 (16) | 13 (33) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 2 or more | 44 (22) | 14 (11) | 10 (27) | 20 (51) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Death – no (%) | Yes | 45 (22) | 27 (21) | 5 (14) | 13 (33) | 0.10 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SVS, Short Viral Shedding; LVS, Long Viral Shedding; no, number; COPD, Chronic Obstructive Pulmonary Disease; Ct, Cycle threshold. Statistic tests, Mann-Whitney or Chi-Square. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Depletion of two bacterial propionate derivatives in moderate/severe Covid-19
To identify metabolites the abundance of which increases or decreases with disease severity, we generated volcano plots that pinpoint variations in the metabolite concentration by at least 20% (up or down) with a p-value ≤0.05 (Figure 2A). Eleven metabolites fulfilled these criteria (Figure 2A) and were then subjected to random forest classification to identify which among them have the best predictive values (Figure 2B). Of note, two chemically related compounds, indole-3-proprionate and 3-phenylproprionate were reduced in cancer patients with moderate/severe Covid-19 compared to cancer patients with mild disease. A prior study has shown that critically ill patient with pneumonia have low serum levels of 3-phenylproprionate, a bacterial product that may be depleted as a result of antibiotic use [34]. Moreover, it is known that indole-3-proprionate is produced by Clostridium sporogenes in the human gastrointestinal tract [35, 36] and that low indole-3-proprionate serum levels reflect a diet poor in fibers [37] and a low microbiome diversity [38]. Thus, the depletion of indole-3-proprionate and 3-phenylproprionate (Figure 2C) that was found may indicate a state of intestinal dysbiosis.
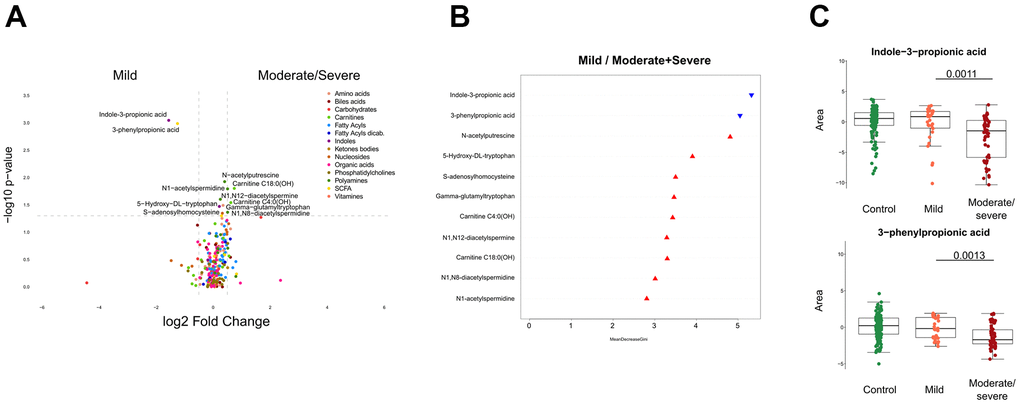
Figure 2. Identification of metabolites discriminating cancer patients by clinical severity of SARS-CoV-2. (A) Volcano plot comparing mild with moderate/severe Covid-19 patients, classified by chemical classes. X-axis: log2 fold change of metabolites; Y-axis: fold change of –log10 P value determined by the Mann–Whitney test. (B) Random forest classification model based on metabolites altered (p < 0.05) between mild and moderate/severe Covid-19 cases. The downregulated and upregulated metabolites in mild compared to moderate/severe patients are marked in blue and red, respectively. (C) Indole-3-propionic acid and 3-phenylproprionic levels in patients. All boxes indicate the interquartile range Q1 to Q3 with Q2 (median levels) in the center. The range of outliers is depicted by whiskers. The black bar of the figure indicates the p-value.
Identification of disease severity-associated acetylated polyamine derivatives
Several acetylated polyamine derivatives (N1-acetylputrescine, N1-acetylspermidine, N1,N8-diacetylspermidine and N1,N12-diacetylspermine) were overabundant in moderate/severe compared to mild Covid-19 (Figures 2A, 3A) when they were analyzed individually (Figure 3A). S-adenosylmethionine, a metabolite that is connected to polyamine synthesis [39], also correlated with Covid-19 severity (Figure 3B) echoing a prior report [40]. The random forest calculations based on all metabolites that changed significantly based on the volcano plot analyses (Figure 2B) yielded a classification model with an out-of-bag (OOB) error rate of 22.35% (Table 2). The boxplot representation illustrates that acetylated polyamine derivatives tend to be higher in Covid-19 infected patients than in controls and that they significantly increase with disease severity (Figure 3A, 3B). The ratio of N1-acetylspermidine over spermidine, as well as the ratio of N1,N8-diacetylspermine over spermidine, increased, but no such increase was found for the ratios of N1-acetylputrescine over putrescine and N1,N12-diacetylspermine over spermine (which was actually reduced) (Supplementary Figure 2). The sum of all acetylated polyamines (N1-acetylputrescine + N1-acetylspermidine + N1,N8-diacetylspermidine + N1,N12-diacetylspermine) yielded a lower discriminative p-value than each of them alone (Figure 4). Altogether, these results support the idea that acetylated polyamine derivatives correlate with Covid-19 severity. When the duration of PCR-detectable SARS-CoV-2 shedding was used to distinguish short-term carriers (<40 days) from long-term-carriers (>40 days), the serum levels of N1-acetylspermidine and N8-acetylspermidine were found to be slightly but significantly increased in long-term carriers (Figure 5). Thus, failure to eliminate SARS-CoV-2 is associated with an increase of selected acetylpolyamines.
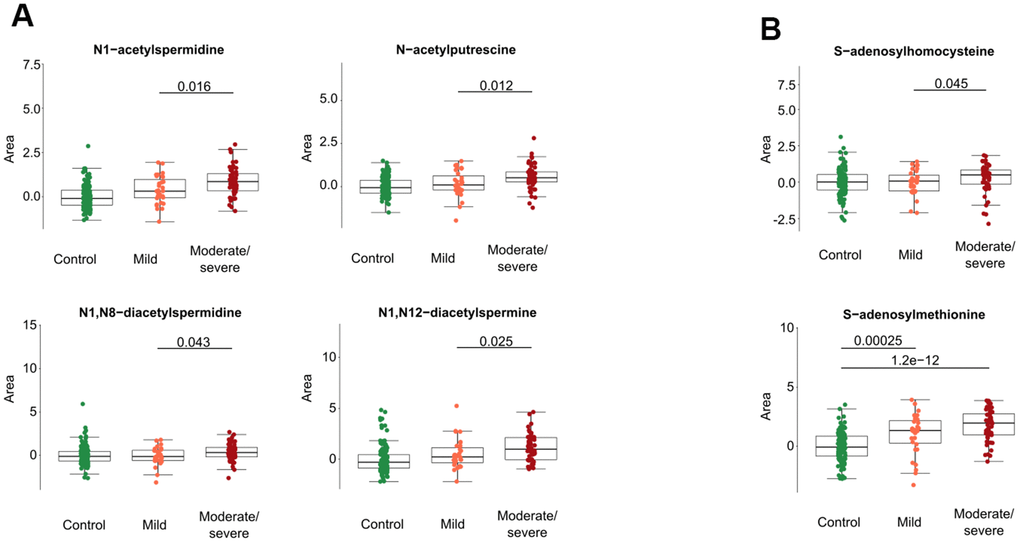
Figure 3. Acetylated polyamine derivatives and associated metabolites in cancer patients with different levels of Covid-19 severity. Acetylated polyamine derivatives were identified by targeted metabolomics data (A). S-adenosylhomocysteine and S-adenosylmethionine are shown (B). All data represent the normalized areas of mass spectrometric peaks and were analyzed by non-parametric unpaired Wilcoxon test (Mann–Whitney) for each two-group comparison. The black bar of the figure indicates the p-value.
Table 2. Error rate estimates for the Random Forest SARS-CoV-2.
| Prediction: OOB estimate of error rate: 22.35% | ||||
| Predicted label | ||||
| Actual label | Label | Mild | Moderate/severe | Class error |
| Mild | 16 | 15 | 48.39% | |
| Moderate/severe | 4 | 50 | 7.41% | |
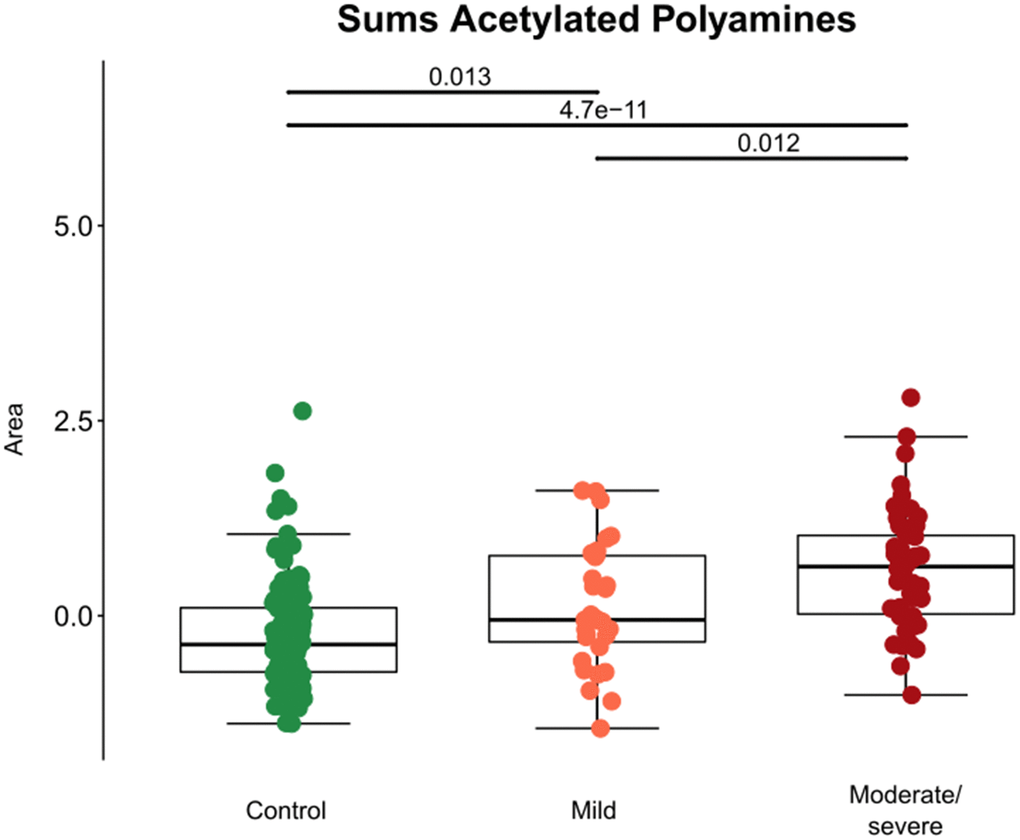
Figure 4. Aggregate analyses of acetylated polyamine derivatives in cancer patients with different levels of Covid-19 severity. For each patient, the sum of the normalized peak areas corresponding to N1-acetylputrescin, N1-acetylspermidine, N1,N12-diacetylspermine and N1,N8-diacetylspermidine were calculated and shown with black bars indicating p-values.
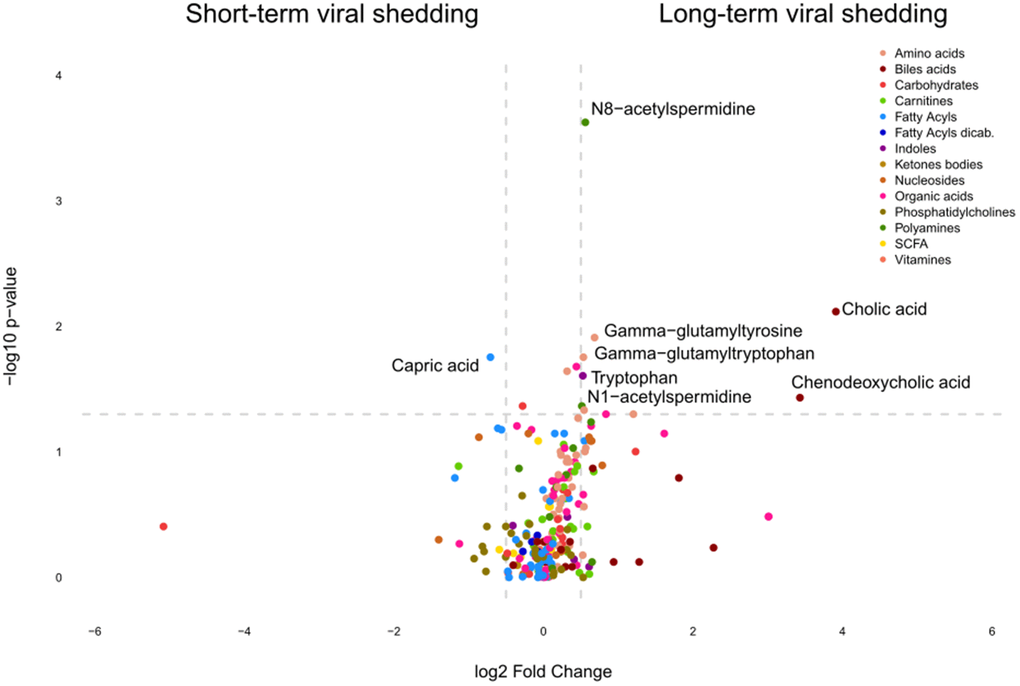
Figure 5. Identification of metabolites discriminating cancer patients according to the duration of SARS-CoV-2 shedding. Volcano plot comparing Covid-19 patients with short versus long viral shedding (determined by RT-PCT of nasopharyngeal PCRs, the threshold between short and long shedding being 40 days), classified by chemical classes. X-axis: log2 fold change of metabolites; Y-axis: fold change of –log10 P value determined by the Mann–Whitney test.
Correlations between circulating metabolites and cytokines
In the final step of data analysis, we established a correlation matrix to visualize positive or negative associations among significantly altered metabolites and cytokines that were measured in the serum from patients enrolled (Figure 6). Non-hierarchical clustering revealed positive associations among several acetylpolyamines, SAM, as well 5-hydroxy-tryptophan and 2 saturated acyl carnitines (butyryl-L-carnitine, arachidyl-L-carnitine with 4 and 20 carbon atoms in the acyl chain, respectively). Among the acetylated polyamines, N1-acetylputrescine correlated with several cytokines in particular, the interferons (IFN) IFNα2a and IFNγ, as well as the interleukins (IL) IL-2 and IL-10, but no such correlation was found for any of the other acetylpolyamines (N1-acetylspermidine, N1,N8-diacetylspermidine and N1,N12-diacetylspermine). All immune and inflammation-related parameters clustered together, including a positive association between indole-3-propionate and 3-phenylpropionate (Figure 6 and Supplementary Figures 3–6). In particular, the association between 3-phenylproprionate and circulating tumor necrosis factor-α (TNFα) was highly significant (Figure 7A). Moreover, this bacterial metabolite exhibited a positive correlation with indole−3−propionate (Figure 7B), which in turn exhibited a negative correlation with N1-acetylspermidine, N1,N8-diacetylspermidine and N1,N12-diacetylspermine (Figure 7C–7E). This anticorrelation supports the general conclusions of this paper.
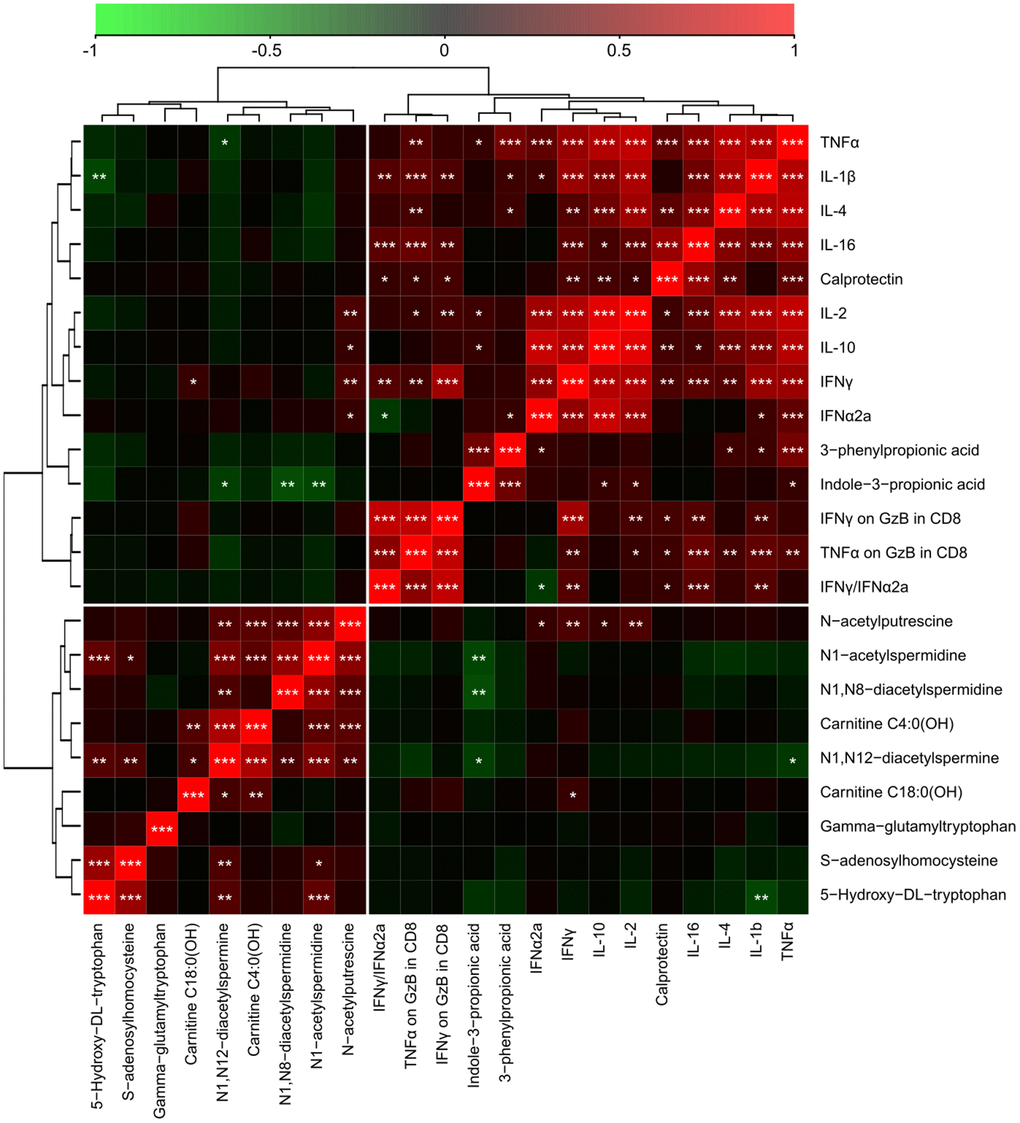
Figure 6. Integration of metabolic with inflammatory markers in serum samples from cancer patients with different levels of Covid-19 severity. The correlation heatmap of data was generated by means of Pearson’s method, and clustered using the ward. D2 method. The red color indicates positive correlations with a FDR<0.05, and the green color marks negative correlations with FDR<0.05. Black indicates non-significant (FDR>0.05) associations. *p < 0.05, **p < 0.01, ***p < 0.001. TNFα: tumor necrosis factor alpha. GzB: Granzyme B. TNFα on GzB in CD8 is the ratio between soluble TNFα and GzB in CD8+ T cells (by flow cytometry) and IFNγ on GzB in CD8 is the ratio between soluble IFNγ GzB in CD8+ T cells (by flow cytometry).
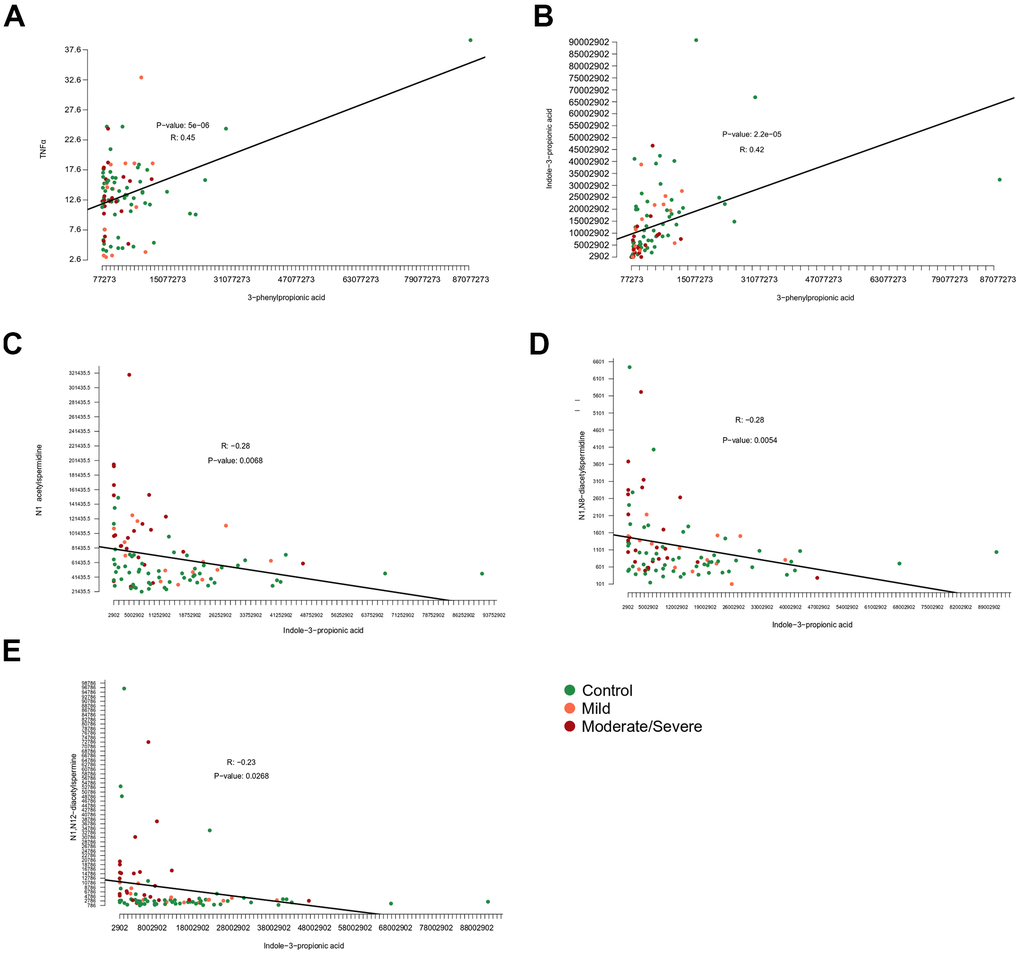
Figure 7. Correlation among metabolites and inflammatory markers. Correlations of (A) 3−phenylpropionic acid and tumor necrosis factor alpha (TNFα), (B) 3−phenylpropionic acid and indole-3-phenylpropionic acid, (C) indol-3-propionic acid and N1-acetylspermidine or (D) N1,N8-diacetylspermidine or (E) N1,N12-diacetylspermine.
Discussion
Here, we present an unbiased metabolomics-based approach to identify circulating metabolites that are elevated in severe Covid-19, in the context of cancer. Our findings indicate that some bacterial metabolites are depleted in severe Covid-19, suggesting the presence of intestinal dysbiosis. Whether this dysbiosis results from severe SARS-CoV-2 infection, reflects a preexisting condition or results from antibiotic treatments remains to be determined.
Prior studies performed on general (non-cancer-specific) patient cohort have revealed a disease severity-associated increase in the tryptophan metabolite kynurenine [24, 28, 30, 41], which is well known for its immunosuppressive properties. Indeed, addition of epacadostat, an inhibitor of the kynurenine-generating enzyme indole 2,3-dioxygenase 1 (IDO1) has been shown to suppress the SARS-CoV-2-induced proinflammatory cytokine release ex vivo [31].
Interestingly, high levels of kynurenine metabolites were identified in the murine serum metabolome after infection by H1N1 influenza virus infection, suggesting that distinct viruses may affect this inflammation-relevant metabolic pathway [42]. In our work, a general (non-cancer-specific) cohort, we found that another immunosuppressive tryptophan derivative, anthranilic acid [43, 44], had a poor prognostic value, correlating with the maintenance of high interleukin-10 and -18 levels [45]. In sharp contrast, we did not find anthranilic acid to be increased in Covid-19 patients with cancer, most likely because the levels of this immunosuppressive metabolite are already increased at baseline, in Covid-19-free cancer patients as compared to cancer-free control individuals (Figure 8). This suggests that malignant disease is associated with an augmentation of circulating anthranilic acid concentrations. Indeed, some cancers such as HER2-positive and triple-negative mammary carcinomas overexpress the enzymes kynurenine 3-monooxygenase (KMO) and kynureninase (KYNU), causing increased production of anthranilic acid [46]. Moreover, poor prognosis prostate cancer is associated with elevated circulating anthranilic acid levels [47]. However, at this point the causes of the elevation of anthranilic acid across different cancer types remain unclear.
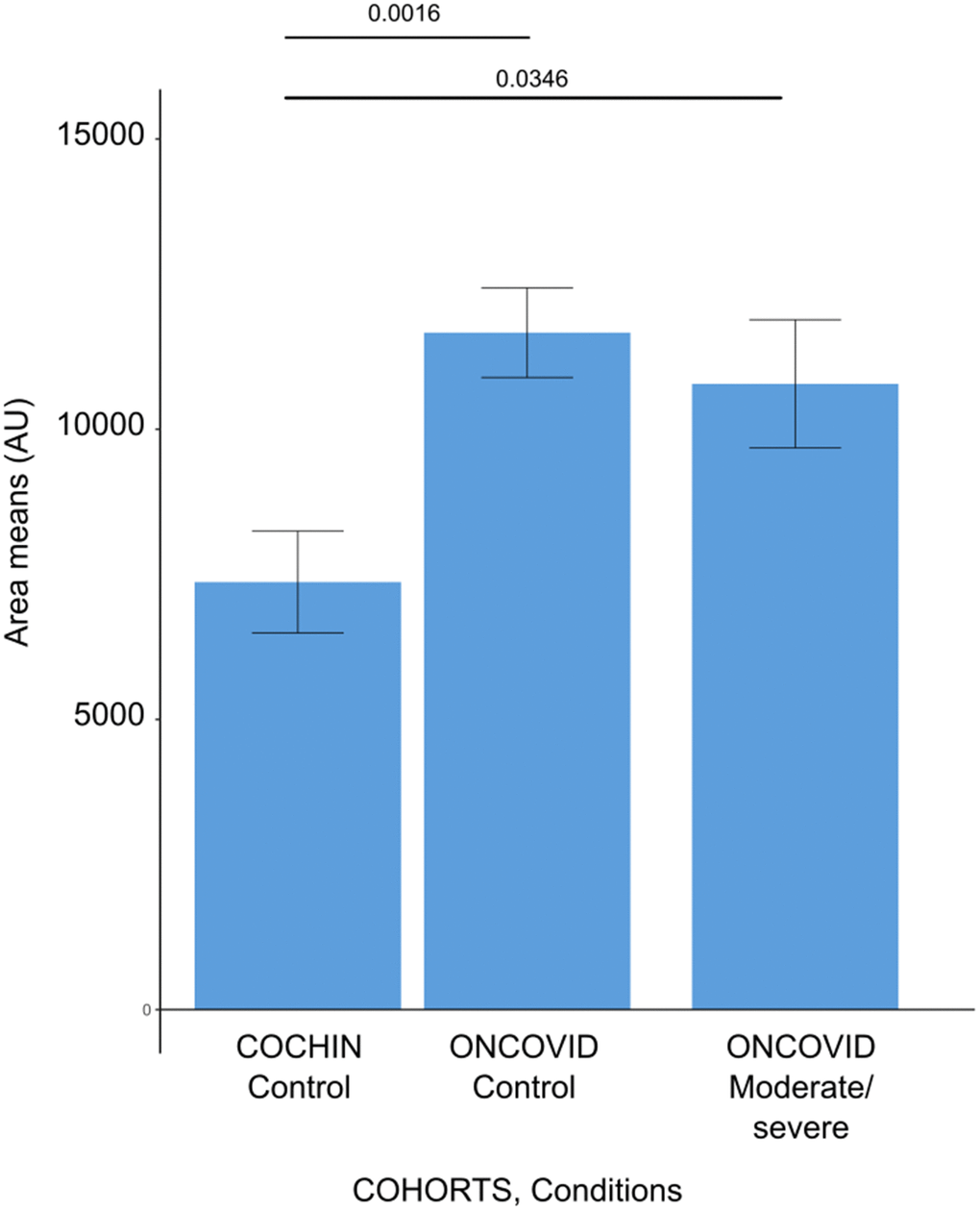
Figure 8. Anthranilic acid levels in different patient cohorts. Barplots demonstrating the anthranilic acid area levels for each subject issued from the cancer-free and Covid-19-free control cohort (COCHIN Control), previously described by Danlos et al., [45], and our dataset of cancer patients without Covid-19 (ONCOVID controls) and the cancer patients with moderate or severe Covid-19 (ONCOVID Moderate/Severe). Error bars show standard errors of the mean.
The most important finding of this study concerns the Covid-19 severity-associated surge in acetylated polyamines found in cancer patients. Increased acetylated polyamine levels, in particular mono- and diacetyl spermidine and spermine derivatives (such as N1-acetylspermidine, N1,N8-diacetylspermidine and N1,N12-diacetylspermine), have been found in cancer-free cohorts of Covid-19 patients to be associated with disease severity [40, 45]. Hence, this alteration appears to be a general feature of moderate or severe Covid-19 infection requiring hospitalization, irrespective of the presence or absence of neoplasia.
However, the functional implications of these findings are elusive. Spermidine inhibits SARS-CoV-2 replication in vitro [48]. Spermidine supplementation is known to exert immunostimulatory effects and to improve the effects of anticancer chemotherapy and immunochemotherapy [39, 49–52]. However, acetylated polyamine derivatives have not been investigated with respect to their potential immunomodulatory effects.
It is important to note that acetylated polyamine derivatives have previously been associated with the risk of developing hepatocellular carcinoma [53], poor prognosis triple-negative breast cancer [54], non-small cell lung cancer [55, 56], colorectal cancer [57], pancreas carcinoma [58], lethal cardiovascular disease [59, 60], Parkinson disease [61] and an elderly-type gut microbiota [62]. This contrasts with the observations that nutritional uptake of spermidine (and spermine but not putrescine) is epidemiologically linked to a decrease in the risk of lethal cancer, cardiovascular disease and cognitive decline [39, 63] and that experimental spermidine supplementation has wide oncopreventive, cardioprotective and neuroprotective effects in preclinical models [64–68].
It is possible that the elevated levels of mono- and di-acetyl spermidine reflect a higher spermidine catabolism, reducing the bioavailability of endogenous spermidine. Indeed, intracellular spermidine and spermine are acetylated by spermidine/spermine N1-acetyltransferase-1 (SAT1) and the resulting acetylated products can be either extruded from the cell or oxidized to putrescine [39, 63]. Thus, transgenic overexpression of SAT1 or its pharmacological activation depletes spermidine and spermine but increases the levels of mono- or diacetylated spermidine and spermine while limiting cellular proliferation and favoring the induction of apoptosis [69, 70]. Inhibition of SAT1 reduces acetyl-coenzyme A consumption in white adipose tissue or the liver, thus enhancing lipogenesis, while its activation stimulates beige adipocyte biogenesis as well as the expression of pro-inflammatory genes [71–73]. Transgene-enforced overexpression of SAT1 accelerates aging in mice [74] as it enhances carcinogenesis [75, 76], while its knockout protects against liver and kidney ischemia-reperfusion damage [77], CCL4-induced acute liver injury [78], and endotoxin- or cisplatin-induced acute kidney injury [79, 80]. Enhanced expression of SAT1 is a biomarker of kidney ischemia-reperfusion damage [81], poor prognosis prostate cancer [82], and radioresistance in brain tumors [83]. Amantadine, a clinically approved antiviral drug, is a substrate for SAT1 and hence can be used to indirectly measure SAT1 activity by assessing the quantity of urinary acetylamantadine. Using this test, patients with breast or lung cancer exhibit an enhanced SAT1 activity [84]. Of note, SAT1 is upregulated by interferons [85], perhaps explaining the shift in polyamine metabolism associated with severe Covid-19. On theoretical grounds, the depletion of intracellular spermidine resulting from SAT1 activation may favor SARS-CoV-2 replication [48] and enfeeble the antiviral immune response due to the depletion of bioavailable spermidine [49, 51, 52, 83, 86]. However, the effects of polyamine catabolism on infections by respiratory viruses have not yet been studied in suitable animal models.
In conclusion, it appears that the severity of Covid-19 affecting cancer cells is strongly related to the acetylation of polyamines, in particular that of spermidine and spermine. It will be important to understand the mechanisms as well as the functional consequences of this over-acetylation detectable in cancer patients.
Materials and Methods
Standards and reagents
Acetonitrile (Sigma Aldrich)
Isopropanol (Sigma Aldrich)
Methanol (Sigma Aldrich)
Chloroform (Sigma Aldrich)
Acetic acid (Sigma Aldrich)
Formic acid (Sigma Aldrich)
Methoxyamine hydrochloride (Sigma Aldrich)
MSTFA - N-Methyl-N-(trimethylsilyl) trifluoroacetamide (Sigma Aldrich)
Pyridine (Sigma Aldrich)
3 nitrophenylhydrazine (Sigma Aldrich)
N-(3-Dimethylaminopropyl)-N′-ethylcarbodiimide hydrochloride (EDC) (Sigma Aldrich)
Sulfosalicylic acid (Sigma Aldrich)
Study ONCOVID design and participants
Gustave Roussy Cancer Center mentored the “ONCOVID” trial and collaborated with the academic authors. All patients provided written informed consent. This clinical trial was conducted in accordance with the principles of the Declaration of Helsinki. Protocol approval was obtained from an independent ethics committee (ethics protocol number EudraCT No: 2020-001250-21). The protocol is available with the full text of this article at https://clinicaltrials.gov/ct2/show/NCT04341207.
Blood samples were drawn from patients enrolled in ONCOVID at Gustave Roussy Cancer Campus (Villejuif, France). Whole human peripheral blood was collected into sterile vacutainer tubes. Serum were collected after centrifugation at 600 × g for 10 min at room temperature and transferred to −80° C freezer to await analysis. Serum samples were used to perform metabolomic approaches and monitor the concentrations of soluble factors. Fixed whole blood were used for spectral flow cytometry. For details, please referred from previously described by Goubet et al. [33].
Evaluation of SARS-CoV-2 RNA shedding
The duration of viral shedding was defined as the number of days from the first positive to the first negative RT-qPCR, after longitudinal monitoring. In order to prevent an overvaluation of this duration, we considered in this analysis only patients with an interval below 40 days between the last positive RT-qPCR and the first negative RT-qPCR. Six patients had one negative RT-qPCR followed by positive RT-qPCR. We extend the duration to the second negative RT-qPCR for 3 patients with a cycle threshold below 35 for the gene coding replication-transcription complex and within 6 days after the first negative result [33].
Sample preparation serum (dry tubes)
A volume of 50 μL of serum were mixed with 500 μL a cold solvent mixture with ISTD (MeOH/Water, 9/1, -20° C), into 1.5 mL microtubes, vortexed and centrifuged (10 min at 15000 g, 4° C) to obtain protein precipitation. Then upper phase of supernatant was split in three parts: 150 μL were used for GC-MS experiment in injection vial, 40 μL were used for the SCFA (Short Chain Fatty Acids) UHPLC-MS method, and 150 μL were used for others UHPLC-MS experimentations as described [45]. Extracted biological samples were pooled in quality controls samples, used for data correction during the data treatment.
Widely-targeted analysis of intracellular metabolites gas chromatography (GC) coupled to a triple quadrupole (QQQ) mass spectrometer
GC-MS/MS method was performed on a 7890B gas chromatography (Agilent Technologies, Waldbronn, Germany) coupled to a triple quadrupole 7000C (Agilent Technologies, Waldbronn, Germany) equipped with a High sensitivity electronic impact source (EI) operating in positive mode [45]. The scan mode used was the MRM for biological samples. Peak detection and integration of analytes were performed using the Agilent Mass Hunter quantitative software (B.07.01), exported as tables and processed with R software (version 4.0.3) and the GRMeta package (Github/kroemerlab).
Targeted analysis of bile acids by ion pairing high performance liquid chromatography (HPLC) coupled to a QTRAP 6500+ mass spectrometer
Targeted analysis was performed on a RRLC 1260 system (Agilent Technologies, Waldbronn, Germany) coupled to a QTRAP 6500+ (Sciex) equipped with an electrospray source operating in negative mode. The source conditions were: ion spray source temperature at 450° C, curtain (CUR) gas pressure at 25 psi, gas 1 (GS1) pressure at 30 psi and gas 2 (GS2) pressure at 70 psi.
2.5 μL of sample were injected on a Column Poroshell 120 EC-C8 (100 mm x 2.1 mm particle size 2.7 μm) from Agilent technologies, protected by a guard column XDB-C18 (5 mm × 2.1 mm particle size 1.8 μm) and heated at 40° C in a Pelletier oven.
Gradient mobile phase consisted of water with 0.2% of formic acid (A) and acetonitrile/isopropanol (1/1; v/v) (B) freshly made. Flow rate was set to 0.3 mL/min, and gradient as follow: initial condition was 70% phase A and 30% phase B, maintained during 1.5 min. Molecules were then eluted using a gradient from 30% to 60% phase B over 9 min. Column was washed using 98% mobile phase B for 2 minutes and equilibrated using 30% mobile phase B for 2 min. After each injection, needle was washed twice with isopropanol and thrice with water. The autosampler was kept at 4° C.
Collision gas was nitrogen. Scan mode used was the MRM for biological samples. Peak detection and integration of the analytes were performed using the Sciex MultiQuant quantitative software (Version 3.0.3), exported as tables and processed with R software (version 4.0.3) and the GRMeta package (Github/kroemerlab).
Targeted analysis of polyamines by ion pairing ultra-high performance liquid chromatography (UHPLC) coupled to a triple quadrupole (QQQ) mass spectrometer
Targeted analysis was performed on a UHPLC 1290 system (Agilent Technologies, Waldbronn, Germany) coupled to a Triple Quadrupole 6470 (Agilent Technologies) equipped with an electrospray source operating in positive mode. The gas temperature was set to 350° C with a gas flow of 12 l/min. The capillary voltage was set to 2.5 kV.
10 μL of sample were injected on a Column Kinetex C18 (150 mm x 2.1 mm particle size 2.6 μm) from Phenomenex, protected by a guard column C18 (5 mm × 2.1 mm) and heated at 40° C in a Pelletier oven.
The gradient mobile phase consisted of water with 0.1 % of Heptafluorobutyric acid (HFBA, Sigma-Aldrich) (A) and acetonitrile with 0.1 % of HFBA (B) freshly made. The flow rate was set to 0.2 ml/min, and gradient as follow: initial condition was 95% phase A and 5% phase B. Molecules were then eluted using a gradient from 5% to 40% phase B over 10 min. The column was washed using 90% mobile phase B for 2.5 minutes and equilibrated using 5% mobile phase B for 4 min. The autosampler was kept at 4° C.
The collision gas was nitrogen. The scan mode used was the MRM for biological samples. Peak detection and integration of analytes were performed using the Agilent Mass Hunter quantitative software (B.07.01), exported as tables and processed with R software (version 4.0.3) and the GRMeta package (Github/kroemerlab).
Targeted analysis of Short Chain Fatty Acid by ultra-high performance liquid chromatography (UHPLC) coupled to a Triple Quadrupole (QQQ) mass spectrometer
Targeted analysis was performed on a UHPLC 1290 system (Agilent Technologies, Waldbronn, Germany) coupled to a Triple Quadrupole 6470 (Agilent Technologies) equipped with an electrospray source operating in negative mode. The gas temperature was set to 300° C with a gas flow of 12 l/min. The capillary voltage was set to 5000 kV.
10 μL of sample were injected on a Column Zorbax Eclipse XBD C18 (100 mm x 2.1 mm particle size 1.8 μm) from Agilent technologies, protected by a guard column XDB-C18 (5 mm × 2.1 mm particle size 1.8 μm) and heated at 50° C by a Pelletier oven.
Gradient mobile phase consisted of water with 0.01% of formic acid (A) and acetonitrile with 0.01% of formic acid (B). Flow rate was set to 0.4 mL/min, and gradient as follow: initial condition was 80% phase A and 20% phase B, maintained during 6 min. Molecules were then eluted using a gradient from 20% to 45% phase B over 7 min. Column was washed using 95% mobile phase B for 5 minutes and equilibrated using 20% mobile phase B for 4 min. Autosampler was kept at 4° C.
The collision gas was nitrogen. The scan mode used was the MRM for biological samples. Peak detection and integration of analytes were performed using the Agilent Mass Hunter quantitative software (B.07.01), exported as tables and processed with R software (version 4.0.3) and the GRMeta package (Github/kroemerlab).
Pseudo-targeted analysis of metabolites by ultra-high performance liquid chromatography (UHPLC) coupled to a Q-Exactive mass spectrometer. Reversed phase acetonitrile method
The profiling experiment was performed with a Dionex Ultimate 3000 UHPLC system (Thermo Scientific) coupled to a Q-Exactive (Thermo Scientific) equipped with an electrospray source operating in both positive and negative mode and full scan mode from 100 to 1200 m/z. The Q-Exactive parameters were: sheath gas flow rate 55 au, auxiliary gas flow rate 15 au, spray voltage 3.3 kV, capillary temperature 300° C, S-Lens RF level 55 V. The mass spectrometer was calibrated with sodium acetate solution dedicated to low mass calibration.
10 μL of sample were injected on a SB-Aq column (100 mm × 2.1 mm particle size 1.8 μm) from Agilent Technologies, protected by a guard column XDB-C18 (5 mm × 2.1 mm particle size 1.8 μm) and heated at 40° C in a Pelletier oven. The gradient mobile phase consists of water with 0.2% of acetic acid (A) and acetonitrile (B). The flow rate was set to 0.3 mL/min. Initial condition is 98% phase A and 2% phase B. Molecules were then eluted using a gradient from 2% to 95% phase B in 22 min. The column was washed using 95% mobile phase B for 2 minutes and equilibrated using 2% mobile phase B for 4 min.
The autosampler was kept at 4° C.
Peak detection and integration were performed using the Thermo Xcalibur quantitative software (3.1.), exported as tables and processed with R software (version 4.0.3) and the GRMeta package (Github/kroemerlab).
In parallel, raw data files obtained by the pseudo-targeted analysis described above were used to perform unbiased profiling analyses using the Thermo Scientific™ Compound Discoverer™ small molecule identification software (version 3.1).
Data analysis using Compound Discoverer™
After sample injection and data acquisition, raw data files were processed with Compound Discoverer software following a customized node-based workflow for identifying unknown compounds in metabolomics.
First, spectra selection and retention time alignment were performed, followed by removal of background noise and baseline correction. Next, the processing workflow found chromatographic peaks for unknown compounds (molecular weight, MW, x retention time, RT) extracting all relevant spectral and chromatographic information, to predict the elemental composition of the unknowns. The possible identity of the unknown compounds was then searched against selected MS databases, such as ChemSpider (from MS1 scans by using MW or predicted composition when available), mZcloud (MS/MS spectral library), built-in databases (custom, local libraries), and Metabolika or KEGG databases (metabolic pathway search). Annotations are assigned to the detected compounds, to rank putative database results. Finally, the software performed statistical analysis using a multivariate method approach, e. g. PCA (unsupervised), and data visualization, e.g. volcano plots.
Compounds found to be statistically different between groups, were checked, and all data was exported to R software (version 4.0.3) for data representation.
Random forest calculation
Targeted data used for random forest calculation were areas corrected by means of quality controls, log2 transformed, and centered on the mean of the control samples (AreaCorrLog2Cen datasheet of the raw data, Supplementary Table 1).
We oriented this calculation on the severity levels of the patient, and focused on the differences of mild versus moderated + severe patients. Data were submitted to a Kruskal-Wallis test, and metabolites were selected according to fold changes and p-values. Mild versus moderate/severe. The two groups had a similar weight in the random forest calculation.
Cytokine measurements
Serum samples and cytokines measurements were prepared according the Goubet et al., method [33]. In this study, only these cytokines considered are, IFN-γ, IL-10, IL-16, IL-1β, IL-2, IL-4, IL-6, TNF-α and Calprotectin and IFN-α2a.
Author Contributions
MB and SD performed the metabolomics, analyzed the mass spectrometric data and generated all Figures, as well as Table 2. LD, C A-C-S. AD, FXD, LC, AG and AP collected and analyzed the clinical data for Table 1. A-GG collected PCR and cytokine data. CG-I, FM and NN contributed to the optimization and realization of mass spectrometry experiments. FM provided conceptual input. LZ and GK supervised the study. GK and SD wrote the paper with input by all other authors.
Conflicts of Interest
L.Z. and G.K. are scientific cofounders of everImmune, a company that develops bacteria for the treatment of cancer. GK is a scientific cofounder of Samsara Therapeutics and Therafast Bio. F.M. has equity interest in and is advisor of TLL The Longevity Labs GmbH and Samsara Therapeutics.
AG, being part of the Drug Development Department (DITEP) at Gustave Roussy, has the following DITEP-relevant disclosures:
Principal/sub-Investigator of Clinical Trials for Abbvie, Adaptimmune, Adlai Nortye USA Inc, Aduro Biotech, Agios Pharmaceuticals, Amgen, Argen-X Bvba, Astex Pharmaceuticals, Astra Zeneca Ab, Aveo, Basilea Pharmaceutica International Ltd, Bayer Healthcare Ag, Bbb Technologies Bv, Beigene, BicycleTx Ltd, Blueprint Medicines, Boehringer Ingelheim, Boston Pharmaceuticals, Bristol Myers Squibb, Ca, Celgene Corporation, Chugai Pharmaceutical Co, Clovis Oncology, Cullinan-Apollo, Curevac, Daiichi Sankyo, Debiopharm, Eisai, Eisai Limited, Eli Lilly, Exelixis, Faron Pharmaceuticals Ltd, Forma Tharapeutics, Gamamabs, Genentech, Glaxosmithkline, H3 Biomedicine, Hoffmann La Roche Ag, Imcheck Therapeutics, Innate Pharma, Institut De Recherche Pierre Fabre, Iris Servier, Iteos Belgium SA, Janssen Cilag, Janssen Research Foundation, Kura Oncology, Kyowa Kirin Pharm. Dev, Lilly France, Loxo Oncology, Lytix Biopharma As, Medimmune, Menarini Ricerche, Merck Sharp and Dohme Chibret, Merrimack Pharmaceuticals, Merus, Millennium Pharmaceuticals, Molecular Partners Ag, Nanobiotix, Nektar Therapeutics, Novartis Pharma, Octimet Oncology Nv, Oncoethix, Oncopeptides, Orion Pharma, Ose Pharma, Pfizer, Pharma Mar, Pierre Fabre, Medicament, Roche, Sanofi Aventis, Seattle Genetics, Sotio A.S, Syros Pharmaceuticals, Taiho Pharma, Tesaro, Turning Point Therapeutics, Xencor.
Research Grants from Astrazeneca, BMS, Boehringer Ingelheim, GSK, INCA, Janssen Cilag, Merck, Novartis, Pfizer, Roche, Sanofi.
Non-financial support (drug supplied) from Astrazeneca, Bayer, BMS, Boringher Ingelheim, GSK, Medimmune, Merck, NH TherAGuiX, Pfizer, Roche.
The authors declare that none of these activities are directly related to the present study.
Funding
This project was specifically supported by Association pour la Recherche sur le Cancer. The Gustave Roussy sponsored clinical study on Covid-19 (ONCOVID; NCT NCT04341207 has been supported by the Fondation Gustave Roussy, the Dassault family, Malakoff Humanis, Agnès b., Izipizi, Ralph Lauren and Sanofi). L.Z. and G.K. were supported by RHU Torino Lumière (ANR-16-RHUS-0008), ONCOBIOME H2020 network, the Seerave Foundation, the Ligue contre le Cancer (équipe labelisée). In addition, GK was supported by the Ligue contre le Cancer (équipe labellisée); Agence National de la Recherche (ANR) – Projets blancs; AMMICa US23/CNRS UMS3655; Association “Ruban Rose”; Cancéropôle Ile-de-France; Fondation pour la Recherche Médicale (FRM); a donation by Elior; European Joint Programme on Rare Diseases (EJPRD); Gustave Roussy Odyssea, the European Union Horizon 2020 Project Oncobiome (grant number 825410); Fondation Carrefour; Institut National du Cancer (INCa); Inserm (HTE); Institut Universitaire de France; LeDucq Foundation; the LabEx Immuno-Oncology (ANR-18-IDEX-0001); the RHU Torino Lumière; the Seerave Foundation; the SIRIC Stratified Oncology Cell DNA Repair and Tumor Immune Elimination (SOCRATE); and the SIRIC Cancer Research and Personalized Medicine (CARPEM). This study contributes to the IdEx Université de Paris ANR-18-IDEX-0001. LZ is supported by the French-German ANR grant “Ileobiome” 19-CE15-0029-01. A-GG was supported by Fondation pour la Recherche Médicale (FRM). F.M. is grateful to the Austrian Science Fund FWF (SFB LIPOTOX F3007 and F3012, DK-MCD W1226, as well as grants P29203, P29262, P27893, P31727) and the Austrian Federal Ministry of Education, Science and Research as well as the University of Graz for grants “Unkonventionelle Forschung-InterFast and Fast4Health” as well as “flysleep” (BMWFW-80.109/0001-WF/V/3b/2015). We acknowledge the support of the field of excellence BioHealth, of NAWI Graz and the BioTechMed-Graz flagship project “EPIAge”. L.D. received support by the Philanthropia Fondation Gustave Roussy. (ONCOVID; NCT NCT04341207 has been supported by the Fondation Gustave Roussy, the Dassault family, Malakoff Humanis, Agnès b., Izipizi, Ralph Lauren and Sanofi).
References
- 1. Raoult D, Zumla A, Locatelli F, Ippolito G, Kroemer G. Coronavirus infections: Epidemiological, clinical and immunological features and hypotheses. Cell Stress. 2020; 4:66–75. https://doi.org/10.15698/cst2020.04.216 [PubMed]
- 2. Melenotte C, Silvin A, Goubet AG, Lahmar I, Dubuisson A, Zumla A, Raoult D, Merad M, Gachot B, Hénon C, Solary E, Fontenay M, André F, et al. Immune responses during COVID-19 infection. Oncoimmunology. 2020; 9:1807836. https://doi.org/10.1080/2162402X.2020.1807836 [PubMed]
- 3. Gandhi RT, Lynch JB, Del Rio C. Mild or Moderate Covid-19. N Engl J Med. 2020; 383:1757–66. https://doi.org/10.1056/NEJMcp2009249 [PubMed]
- 4. Berlin DA, Gulick RM, Martinez FJ. Severe Covid-19. N Engl J Med. 2020; 383:2451–60. https://doi.org/10.1056/NEJMcp2009575 [PubMed]
- 5. Derosa L, Melenotte C, Griscelli F, Gachot B, Marabelle A, Kroemer G, Zitvogel L. The immuno-oncological challenge of COVID-19. Nat Cancer. 2020; 1:946–64. https://doi.org/10.1038/s43018-020-00122-3
- 6. Hanahan D, Weinberg RA. Hallmarks of cancer: The next generation. Cell. 2011; 144:646–74. https://doi.org/10.1016/j.cell.2011.02.013 [PubMed]
- 7. López-Otín C, Blasco MA, Partridge L, Serrano M, Kroemer G. The hallmarks of aging. Cell. 2013; 153:1194–217. https://doi.org/10.1016/j.cell.2013.05.039 [PubMed]
- 8. López-Otín C, Kroemer G. Hallmarks of Health. Cell. 2021; 184:33–63. https://doi.org/10.1016/j.cell.2020.11.034 [PubMed]
- 9. Zitvogel L, Tesniere A, Kroemer G. Cancer despite immunosurveillance: immunoselection and immunosubversion. Nat Rev Immunol. 2006; 6:715–27. https://doi.org/10.1038/nri1936 [PubMed]
- 10. Palucka AK, Coussens LM. The Basis of Oncoimmunology. Cell. 2016; 164:1233–47. https://doi.org/10.1016/j.cell.2016.01.049 [PubMed]
- 11. Petroni G, Buqué A, Zitvogel L, Kroemer G, Galluzzi L. Immunomodulation by targeted anticancer agents. Cancer Cell. 2021; 39:310–45. https://doi.org/10.1016/j.ccell.2020.11.009 [PubMed]
- 12. Prasanna PG, Citrin DE, Hildesheim J, Ahmed MM, Venkatachalam S, Riscuta G, Xi D, Zheng G, van Deursen J, Goronzy J, Kron SJ, Anscher MS, Sharpless NE, et al. Therapy-Induced Senescence: Opportunities to Improve Anti-Cancer Therapy. J Natl Cancer Inst. 2021. [Epub ahead of print]. https://doi.org/10.1093/jnci/djab064 [PubMed]
- 13. Routy B, Derosa L, Zitvogel L, Kroemer G. COVID-19: a challenge for oncology services. Oncoimmunology. 2020; 9:1760686. https://doi.org/10.1080/2162402X.2020.1760686 [PubMed]
- 14. Chen H, Guo J, Wang C, Luo F, Yu X, Zhang W, Li J, Zhao D, Xu D, Gong Q, Liao J, Yang H, Hou W, Zhang Y. Clinical characteristics and intrauterine vertical transmission potential of COVID-19 infection in nine pregnant women: a retrospective review of medical records. Lancet. 2020; 395:809–15. https://doi.org/10.1016/S0140-6736(20)30360-3 [PubMed]
- 15. Zhang Q, Bastard P, Liu Z, Le Pen J, Moncada-Velez M, Chen J, Ogishi M, Sabli IK, Hodeib S, Korol C, Rosain J, Bilguvar K, Ye J, et al, and COVID-STORM Clinicians, and COVID Clinicians, and Imagine COVID Group, and French COVID Cohort Study Group, and CoV-Contact Cohort, and Amsterdam UMC Covid-19 Biobank, and COVID Human Genetic Effort, and NIAID-USUHS/TAGC COVID Immunity Group. Inborn errors of type I IFN immunity in patients with life-threatening COVID-19. Science. 2020; 370:eabd4570. https://doi.org/10.1126/science.abd4570 [PubMed]
- 16. Bastard P, Rosen LB, Zhang Q, Michailidis E, Hoffmann HH, Zhang Y, Dorgham K, Philippot Q, Rosain J, Béziat V, Manry J, Shaw E, Haljasmägi L, et al, and HGID Lab, and NIAID-USUHS Immune Response to COVID Group, and COVID Clinicians, and COVID-STORM Clinicians, and Imagine COVID Group, and French COVID Cohort Study Group, and Milieu Intérieur Consortium, and CoV-Contact Cohort, and Amsterdam UMC Covid-19 Biobank, and COVID Human Genetic Effort. Autoantibodies against type I IFNs in patients with life-threatening COVID-19. Science. 2020; 370:eabd4585. https://doi.org/10.1126/science.abd4585 [PubMed]
- 17. Silvin A, Chapuis N, Dunsmore G, Goubet AG, Dubuisson A, Derosa L, Almire C, Hénon C, Kosmider O, Droin N, Rameau P, Catelain C, Alfaro A, et al. Elevated Calprotectin and Abnormal Myeloid Cell Subsets Discriminate Severe from Mild COVID-19. Cell. 2020; 182:1401–18.e18. https://doi.org/10.1016/j.cell.2020.08.002 [PubMed]
- 18. Del Valle DM, Kim-Schulze S, Huang HH, Beckmann ND, Nirenberg S, Wang B, Lavin Y, Swartz TH, Madduri D, Stock A, Marron TU, Xie H, Patel M, et al. An inflammatory cytokine signature predicts COVID-19 severity and survival. Nat Med. 2020; 26:1636–43. https://doi.org/10.1038/s41591-020-1051-9 [PubMed]
- 19. Zheng H, Rao AM, Dermadi D, Toh J, Murphy Jones L, Donato M, Liu Y, Su Y, Dai CL, Kornilov SA, Karagiannis M, Marantos T, Hasin-Brumshtein Y, et al. Multi-cohort analysis of host immune response identifies conserved protective and detrimental modules associated with severity across viruses. Immunity. 2021; 54:753–68.e5. https://doi.org/10.1016/j.immuni.2021.03.002 [PubMed]
- 20. Stukalov A, Girault V, Grass V, Karayel O, Bergant V, Urban C, Haas DA, Huang Y, Oubraham L, Wang A, Hamad MS, Piras A, Hansen FM, et al. Multilevel proteomics reveals host perturbations by SARS-CoV-2 and SARS-CoV. Nature. 2021; 594:246–52. https://doi.org/10.1038/s41586-021-03493-4 [PubMed]
- 21. Viltard M, Durand S, Pérez-Lanzón M, Aprahamian F, Lefevre D, Leroy C, Madeo F, Kroemer G, Friedlander G. The metabolomic signature of extreme longevity: naked mole rats versus mice. Aging (Albany NY). 2019; 11:4783–800. https://doi.org/10.18632/aging.102116 [PubMed]
- 22. Blasco H, Bessy C, Plantier L, Lefevre A, Piver E, Bernard L, Marlet J, Stefic K, Benz-de Bretagne I, Cannet P, Lumbu H, Morel T, Boulard P, et al. The specific metabolome profiling of patients infected by SARS-COV-2 supports the key role of tryptophan-nicotinamide pathway and cytosine metabolism. Sci Rep. 2020; 10:16824. https://doi.org/10.1038/s41598-020-73966-5 [PubMed]
- 23. Bruzzone C, Bizkarguenaga M, Gil-Redondo R, Diercks T, Arana E, García de Vicuña A, Seco M, Bosch A, Palazón A, San Juan I, Laín A, Gil-Martínez J, Bernardo-Seisdedos G, et al. SARS-CoV-2 Infection Dysregulates the Metabolomic and Lipidomic Profiles of Serum. iScience. 2020; 23:101645. https://doi.org/10.1016/j.isci.2020.101645 [PubMed]
- 24. Fraser DD, Slessarev M, Martin CM, Daley M, Patel MA, Miller MR, Patterson EK, O’Gorman DB, Gill SE, Wishart DS, Mandal R, Cepinskas G. Metabolomics Profiling of Critically Ill Coronavirus Disease 2019 Patients: Identification of Diagnostic and Prognostic Biomarkers. Crit Care Explor. 2020; 2:e0272. https://doi.org/10.1097/CCE.0000000000000272 [PubMed]
- 25. Shi D, Yan R, Lv L, Jiang H, Lu Y, Sheng J, Xie J, Wu W, Xia J, Xu K, Gu S, Chen Y, Huang C, et al. The serum metabolome of COVID-19 patients is distinctive and predictive. Metabolism. 2021; 118:154739. https://doi.org/10.1016/j.metabol.2021.154739 [PubMed]
- 26. Su Y, Chen D, Yuan D, Lausted C, Choi J, Dai CL, Voillet V, Duvvuri VR, Scherler K, Troisch P, Baloni P, Qin G, Smith B, et al, and ISB-Swedish COVID19 Biobanking Unit. Multi-Omics Resolves a Sharp Disease-State Shift between Mild and Moderate COVID-19. Cell. 2020; 183:1479–95.e20. https://doi.org/10.1016/j.cell.2020.10.037 [PubMed]
- 27. Grajeda-Iglesias C, Durand S, Daillère R, Iribarren K, Lemaitre F, Derosa L, Aprahamian F, Bossut N, Nirmalathasan N, Madeo F, Zitvogel L, Kroemer G. Oral administration of Akkermansia muciniphila elevates systemic antiaging and anticancer metabolites. Aging (Albany NY). 2021; 13:6375–405. https://doi.org/10.18632/aging.202739 [PubMed]
- 28. Marín-Corral J, Rodríguez-Morató J, Gomez-Gomez A, Pascual-Guardia S, Muñoz-Bermúdez R, Salazar-Degracia A, Pérez-Terán P, Restrepo MI, Khymenets O, Haro N, Masclans JR, Pozo OJ. Metabolic Signatures Associated with Severity in Hospitalized COVID-19 Patients. Int J Mol Sci. 2021; 22:4794. https://doi.org/10.3390/ijms22094794 [PubMed]
- 29. Páez-Franco JC, Torres-Ruiz J, Sosa-Hernández VA, Cervantes-Díaz R, Romero-Ramírez S, Pérez-Fragoso A, Meza-Sánchez DE, Germán-Acacio JM, Maravillas-Montero JL, Mejía-Domínguez NR, Ponce-de-León A, Ulloa-Aguirre A, Gómez-Martín D, Llorente L. Metabolomics analysis reveals a modified amino acid metabolism that correlates with altered oxygen homeostasis in COVID-19 patients. Sci Rep. 2021; 11:6350. https://doi.org/10.1038/s41598-021-85788-0 [PubMed]
- 30. Sindelar M, Stancliffe E, Schwaiger-Haber M, Anbukumar DS, Albrecht RA, Liu WC, Travis KA, García-Sastre A, Shriver LP, Patti GJ. Longitudinal Metabolomics of Human Plasma Reveals Robust Prognostic Markers of COVID-19 Disease Severity. medRxiv. 2021. [Epub ahead of print]. https://doi.org/10.1101/2021.02.05.21251173 [PubMed]
- 31. Xiao N, Nie M, Pang H, Wang B, Hu J, Meng X, Li K, Ran X, Long Q, Deng H, Chen N, Li S, Tang N, et al. Integrated cytokine and metabolite analysis reveals immunometabolic reprogramming in COVID-19 patients with therapeutic implications. Nat Commun. 2021; 12:1618. https://doi.org/10.1038/s41467-021-21907-9 [PubMed]
- 32. Zheng H, Jin S, Li T, Ying W, Ying B, Chen D, Ning J, Zheng C, Li Y, Li C, Chen C, Li X, Gao H. Metabolomics reveals sex-specific metabolic shifts and predicts the duration from positive to negative in non-severe COVID-19 patients during recovery process. Comput Struct Biotechnol J. 2021; 19:1863–73. https://doi.org/10.1016/j.csbj.2021.03.039 [PubMed]
- 33. Goubet AG, Dubuisson A, Geraud A, Danlos FX, Terrisse S, Silva CA, Drubay D, Touri L, Picard M, Mazzenga M, Silvin A, Dunsmore G, Haddad Y, et al. Prolonged SARS-CoV-2 RNA virus shedding and lymphopenia are hallmarks of COVID-19 in cancer patients with poor prognosis. Cell Death Differ. 2021. [Epub ahead of print]. https://doi.org/10.1038/s41418-021-00817-9 [PubMed]
- 34. Chernevskaya E, Beloborodova N, Klimenko N, Pautova A, Shilkin D, Gusarov V, Tyakht A. Serum and fecal profiles of aromatic microbial metabolites reflect gut microbiota disruption in critically ill patients: a prospective observational pilot study. Crit Care. 2020; 24:312. https://doi.org/10.1186/s13054-020-03031-0 [PubMed]
- 35. Wikoff WR, Anfora AT, Liu J, Schultz PG, Lesley SA, Peters EC, Siuzdak G. Metabolomics analysis reveals large effects of gut microflora on mammalian blood metabolites. Proc Natl Acad Sci USA. 2009; 106:3698–703. https://doi.org/10.1073/pnas.0812874106 [PubMed]
- 36. Zhang LS, Davies SS. Microbial metabolism of dietary components to bioactive metabolites: opportunities for new therapeutic interventions. Genome Med. 2016; 8:46. https://doi.org/10.1186/s13073-016-0296-x [PubMed]
- 37. de Mello VD, Paananen J, Lindström J, Lankinen MA, Shi L, Kuusisto J, Pihlajamäki J, Auriola S, Lehtonen M, Rolandsson O, Bergdahl IA, Nordin E, Ilanne-Parikka P, et al. Indolepropionic acid and novel lipid metabolites are associated with a lower risk of type 2 diabetes in the Finnish Diabetes Prevention Study. Sci Rep. 2017; 7:46337. https://doi.org/10.1038/srep46337 [PubMed]
- 38. Menni C, Hernandez MM, Vital M, Mohney RP, Spector TD, Valdes AM. Circulating levels of the anti-oxidant indoleproprionic acid are associated with higher gut microbiome diversity. Gut Microbes. 2019; 10:688–95. https://doi.org/10.1080/19490976.2019.1586038 [PubMed]
- 39. Madeo F, Hofer SJ, Pendl T, Bauer MA, Eisenberg T, Carmona-Gutierrez D, Kroemer G. Nutritional Aspects of Spermidine. Annu Rev Nutr. 2020; 40:135–59. https://doi.org/10.1146/annurev-nutr-120419-015419 [PubMed]
- 40. Thomas T, Stefanoni D, Reisz JA, Nemkov T, Bertolone L, Francis RO, Hudson KE, Zimring JC, Hansen KC, Hod EA, Spitalnik SL, D’Alessandro A. COVID-19 infection alters kynurenine and fatty acid metabolism, correlating with IL-6 levels and renal status. JCI Insight. 2020; 5:e140327. https://doi.org/10.1172/jci.insight.140327 [PubMed]
- 41. Lawler NG, Gray N, Kimhofer T, Boughton B, Gay M, Yang R, Morillon AC, Chin ST, Ryan M, Begum S, Bong SH, Coudert JD, Edgar D, et al. Systemic Perturbations in Amine and Kynurenine Metabolism Associated with Acute SARS-CoV-2 Infection and Inflammatory Cytokine Responses. J Proteome Res. 2021; 20:2796–811. https://doi.org/10.1021/acs.jproteome.1c00052 [PubMed]
- 42. Chandler JD, Hu X, Ko EJ, Park S, Lee YT, Orr M, Fernandes J, Uppal K, Kang SM, Jones DP, Go YM. Metabolic pathways of lung inflammation revealed by high-resolution metabolomics (HRM) of H1N1 influenza virus infection in mice. Am J Physiol Regul Integr Comp Physiol. 2016; 311:R906–16. https://doi.org/10.1152/ajpregu.00298.2016 [PubMed]
- 43. Platten M, Ho PP, Youssef S, Fontoura P, Garren H, Hur EM, Gupta R, Lee LY, Kidd BA, Robinson WH, Sobel RA, Selley ML, Steinman L. Treatment of autoimmune neuroinflammation with a synthetic tryptophan metabolite. Science. 2005; 310:850–55. https://doi.org/10.1126/science.1117634 [PubMed]
- 44. Yong-Gang X, Ming-Zhe W, Jin-Yan Z, Zhi-Hai P, Jun-Ming X. Combination of N-(3'4'-dimethoxycinnamoyl) anthranilic acid with cyclosporin A treatment preserves immunosuppressive effect and reduces the side effect of cyclosporin A in rat. Eur J Pharmacol. 2014; 728:16–23. https://doi.org/10.1016/j.ejphar.2014.01.055 [PubMed]
- 45. Danlos FX, Grajeda-Iglesias C, Durand S, Sauvat A, Roumier M, Cantin D, Colomba E, Rohmer J, Pommeret F, Baciarello G, Willekens C, Vasse M, Griscelli F, et al. Metabolomic analyses of COVID-19 patients unravel stage-dependent and prognostic biomarkers. Cell Death Dis. 2021; 12:258. https://doi.org/10.1038/s41419-021-03540-y [PubMed]
- 46. Heng B, Bilgin AA, Lovejoy DB, Tan VX, Milioli HH, Gluch L, Bustamante S, Sabaretnam T, Moscato P, Lim CK, Guillemin GJ. Differential kynurenine pathway metabolism in highly metastatic aggressive breast cancer subtypes: beyond IDO1-induced immunosuppression. Breast Cancer Res. 2020; 22:113. https://doi.org/10.1186/s13058-020-01351-1 [PubMed]
- 47. Khan A, Choi SA, Na J, Pamungkas AD, Jung KJ, Jee SH, Park YH. Noninvasive Serum Metabolomic Profiling Reveals Elevated Kynurenine Pathway’s Metabolites in Humans with Prostate Cancer. J Proteome Res. 2019; 18:1532–41. https://doi.org/10.1021/acs.jproteome.8b00803 [PubMed]
- 48. Gassen NC, Papies J, Bajaj T, Emanuel J, Dethloff F, Chua RL, Trimpert J, Heinemann N, Niemeyer C, Weege F, Hönzke K, Aschman T, Heinz DE, et al. SARS-CoV-2-mediated dysregulation of metabolism and autophagy uncovers host-targeting antivirals. Nat Commun. 2021; 12:3818. https://doi.org/10.1038/s41467-021-24007-w [PubMed]
- 49. Pietrocola F, Pol J, Vacchelli E, Rao S, Enot DP, Baracco EE, Levesque S, Castoldi F, Jacquelot N, Yamazaki T, Senovilla L, Marino G, Aranda F, et al. Caloric Restriction Mimetics Enhance Anticancer Immunosurveillance. Cancer Cell. 2016; 30:147–60. https://doi.org/10.1016/j.ccell.2016.05.016 [PubMed]
- 50. Lévesque S, Le Naour J, Pietrocola F, Paillet J, Kremer M, Castoldi F, Baracco EE, Wang Y, Vacchelli E, Stoll G, Jolly A, De La Grange P, Zitvogel L, et al. A synergistic triad of chemotherapy, immune checkpoint inhibitors, and caloric restriction mimetics eradicates tumors in mice. Oncoimmunology. 2019; 8:e1657375. https://doi.org/10.1080/2162402X.2019.1657375 [PubMed]
- 51. Zhang H, Alsaleh G, Feltham J, Sun Y, Napolitano G, Riffelmacher T, Charles P, Frau L, Hublitz P, Yu Z, Mohammed S, Ballabio A, Balabanov S, et al. Polyamines Control eIF5A Hypusination, TFEB Translation, and Autophagy to Reverse B Cell Senescence. Mol Cell. 2019; 76:110–25.e9. https://doi.org/10.1016/j.molcel.2019.08.005 [PubMed]
- 52. Alsaleh G, Panse I, Swadling L, Zhang H, Richter FC, Meyer A, Lord J, Barnes E, Klenerman P, Green C, Simon AK. Autophagy in T cells from aged donors is maintained by spermidine and correlates with function and vaccine responses. Elife. 2020; 9:e57950. https://doi.org/10.7554/eLife.57950 [PubMed]
- 53. Stepien M, Keski-Rahkonen P, Kiss A, Robinot N, Duarte-Salles T, Murphy N, Perlemuter G, Viallon V, Tjønneland A, Rostgaard-Hansen AL, Dahm CC, Overvad K, Boutron-Ruault MC, et al. Metabolic perturbations prior to hepatocellular carcinoma diagnosis: Findings from a prospective observational cohort study. Int J Cancer. 2021; 148:609–25. https://doi.org/10.1002/ijc.33236 [PubMed]
- 54. Fahrmann JF, Vykoukal J, Fleury A, Tripathi S, Dennison JB, Murage E, Wang P, Yu CY, Capello M, Creighton CJ, Do KA, Long JP, Irajizad E, et al. Association Between Plasma Diacetylspermine and Tumor Spermine Synthase With Outcome in Triple-Negative Breast Cancer. J Natl Cancer Inst. 2020; 112:607–16. https://doi.org/10.1093/jnci/djz182 [PubMed]
- 55. Kato M, Onishi H, Matsumoto K, Motoshita J, Tsuruta N, Higuchi K, Katano M. Prognostic significance of urine N1, N12-diacetylspermine in patients with non-small cell lung cancer. Anticancer Res. 2014; 34:3053–59. [PubMed]
- 56. Wikoff WR, Hanash S, DeFelice B, Miyamoto S, Barnett M, Zhao Y, Goodman G, Feng Z, Gandara D, Fiehn O, Taguchi A. Diacetylspermine Is a Novel Prediagnostic Serum Biomarker for Non-Small-Cell Lung Cancer and Has Additive Performance With Pro-Surfactant Protein B. J Clin Oncol. 2015; 33:3880–86. https://doi.org/10.1200/JCO.2015.61.7779 [PubMed]
- 57. Venäläinen MK, Roine AN, Häkkinen MR, Vepsäläinen JJ, Kumpulainen PS, Kiviniemi MS, Lehtimäki T, Oksala NK, Rantanen TK. Altered Polyamine Profiles in Colorectal Cancer. Anticancer Res. 2018; 38:3601–07. https://doi.org/10.21873/anticanres.12634 [PubMed]
- 58. Fahrmann JF, Bantis LE, Capello M, Scelo G, Dennison JB, Patel N, Murage E, Vykoukal J, Kundnani DL, Foretova L, Fabianova E, Holcatova I, Janout V, et al. A Plasma-Derived Protein-Metabolite Multiplexed Panel for Early-Stage Pancreatic Cancer. J Natl Cancer Inst. 2019; 111:372–79. https://doi.org/10.1093/jnci/djy126 [PubMed]
- 59. Nayak A, Liu C, Mehta A, Ko YA, Tahhan AS, Dhindsa DS, Uppal K, Jones DP, Butler J, Morris AA, Quyyumi AA. N8-Acetylspermidine: A Polyamine Biomarker in Ischemic Cardiomyopathy With Reduced Ejection Fraction. J Am Heart Assoc. 2020; 9:e016055. https://doi.org/10.1161/JAHA.120.016055 [PubMed]
- 60. Tahir UA, Katz DH, Zhao T, Ngo D, Cruz DE, Robbins JM, Chen ZZ, Peterson B, Benson MD, Shi X, Dailey L, Andersson C, Vasan RS, et al. Metabolomic Profiles and Heart Failure Risk in Black Adults: Insights From the Jackson Heart Study. Circ Heart Fail. 2021; 14:e007275. https://doi.org/10.1161/CIRCHEARTFAILURE.120.007275 [PubMed]
- 61. Saiki S, Sasazawa Y, Fujimaki M, Kamagata K, Kaga N, Taka H, Li Y, Souma S, Hatano T, Imamichi Y, Furuya N, Mori A, Oji Y, et al. A metabolic profile of polyamines in Parkinson disease: A promising biomarker. Ann Neurol. 2019; 86:251–63. https://doi.org/10.1002/ana.25516 [PubMed]
- 62. Yoshimoto S, Mitsuyama E, Yoshida K, Odamaki T, Xiao JZ. Enriched metabolites that potentially promote age-associated diseases in subjects with an elderly-type gut microbiota. Gut Microbes. 2021; 13:1–11. https://doi.org/10.1080/19490976.2020.1865705 [PubMed]
- 63. Madeo F, Eisenberg T, Pietrocola F, Kroemer G. Spermidine in health and disease. Science. 2018; 359:eaan2788. https://doi.org/10.1126/science.aan2788 [PubMed]
- 64. Eisenberg T, Abdellatif M, Schroeder S, Primessnig U, Stekovic S, Pendl T, Harger A, Schipke J, Zimmermann A, Schmidt A, Tong M, Ruckenstuhl C, Dammbrueck C, et al. Cardioprotection and lifespan extension by the natural polyamine spermidine. Nat Med. 2016; 22:1428–38. https://doi.org/10.1038/nm.4222 [PubMed]
- 65. Kiechl S, Pechlaner R, Willeit P, Notdurfter M, Paulweber B, Willeit K, Werner P, Ruckenstuhl C, Iglseder B, Weger S, Mairhofer B, Gartner M, Kedenko L, et al. Higher spermidine intake is linked to lower mortality: a prospective population-based study. Am J Clin Nutr. 2018; 108:371–80. https://doi.org/10.1093/ajcn/nqy102 [PubMed]
- 66. Pietrocola F, Castoldi F, Kepp O, Carmona-Gutierrez D, Madeo F, Kroemer G. Spermidine reduces cancer-related mortality in humans. Autophagy. 2019; 15:362–65. https://doi.org/10.1080/15548627.2018.1539592 [PubMed]
- 67. Madeo F, Carmona-Gutierrez D, Hofer SJ, Kroemer G. Caloric Restriction Mimetics against Age-Associated Disease: Targets, Mechanisms, and Therapeutic Potential. Cell Metab. 2019; 29:592–610. https://doi.org/10.1016/j.cmet.2019.01.018 [PubMed]
- 68. Schroeder S, Hofer SJ, Zimmermann A, Pechlaner R, Dammbrueck C, Pendl T, Marcello GM, Pogatschnigg V, Bergmann M, Müller M, Gschiel V, Ristic S, Tadic J, et al. Dietary spermidine improves cognitive function. Cell Rep. 2021; 35:108985. https://doi.org/10.1016/j.celrep.2021.108985 [PubMed]
- 69. Mandal S, Mandal A, Johansson HE, Orjalo AV, Park MH. Depletion of cellular polyamines, spermidine and spermine, causes a total arrest in translation and growth in mammalian cells. Proc Natl Acad Sci USA. 2013; 110:2169–74. https://doi.org/10.1073/pnas.1219002110 [PubMed]
- 70. Pietilä M, Alhonen L, Halmekytö M, Kanter P, Jänne J, Porter CW. Activation of polyamine catabolism profoundly alters tissue polyamine pools and affects hair growth and female fertility in transgenic mice overexpressing spermidine/spermine N1-acetyltransferase. J Biol Chem. 1997; 272:18746–51. https://doi.org/10.1074/jbc.272.30.18746 [PubMed]
- 71. Castoldi F, Hyvönen MT, Durand S, Aprahamian F, Sauvat A, Malik SA, Baracco EE, Vacchelli E, Opolon P, Signolle N, Lefevre D, Bossut N, Eisenberg T, et al. Chemical activation of SAT1 corrects diet-induced metabolic syndrome. Cell Death Differ. 2020; 27:2904–20. https://doi.org/10.1038/s41418-020-0550-z [PubMed]
- 72. Liu C, Perez-Leal O, Barrero C, Zahedi K, Soleimani M, Porter C, Merali S. Modulation of polyamine metabolic flux in adipose tissue alters the accumulation of body fat by affecting glucose homeostasis. Amino Acids. 2014; 46:701–15. https://doi.org/10.1007/s00726-013-1548-3 [PubMed]
- 73. Yuan F, Zhang L, Cao Y, Gao W, Zhao C, Fang Y, Zahedi K, Soleimani M, Lu X, Fang Z, Yang Q. Spermidine/spermine N1-acetyltransferase-mediated polyamine catabolism regulates beige adipocyte biogenesis. Metabolism. 2018; 85:298–304. https://doi.org/10.1016/j.metabol.2018.04.007 [PubMed]
- 74. Cerrada-Gimenez M, Pietilä M, Loimas S, Pirinen E, Hyvönen MT, Keinänen TA, Jänne J, Alhonen L. Continuous oxidative stress due to activation of polyamine catabolism accelerates aging and protects against hepatotoxic insults. Transgenic Res. 2011; 20:387–96. https://doi.org/10.1007/s11248-010-9422-5 [PubMed]
- 75. Tucker JM, Murphy JT, Kisiel N, Diegelman P, Barbour KW, Davis C, Medda M, Alhonen L, Jänne J, Kramer DL, Porter CW, Berger FG. Potent modulation of intestinal tumorigenesis in Apcmin/+ mice by the polyamine catabolic enzyme spermidine/spermine N1-acetyltransferase. Cancer Res. 2005; 65:5390–98. https://doi.org/10.1158/0008-5472.CAN-05-0229 [PubMed]
- 76. Wang X, Feith DJ, Welsh P, Coleman CS, Lopez C, Woster PM, O’Brien TG, Pegg AE. Studies of the mechanism by which increased spermidine/spermine N1-acetyltransferase activity increases susceptibility to skin carcinogenesis. Carcinogenesis. 2007; 28:2404–11. https://doi.org/10.1093/carcin/bgm162 [PubMed]
- 77. Zahedi K, Lentsch AB, Okaya T, Barone S, Sakai N, Witte DP, Arend LJ, Alhonen L, Jell J, Jänne J, Porter CW, Soleimani M. Spermidine/spermine-N1-acetyltransferase ablation protects against liver and kidney ischemia-reperfusion injury in mice. Am J Physiol Gastrointest Liver Physiol. 2009; 296:G899–909. https://doi.org/10.1152/ajpgi.90507.2008 [PubMed]
- 78. Zahedi K, Barone SL, Xu J, Steinbergs N, Schuster R, Lentsch AB, Amlal H, Wang J, Casero RA
Jr , Soleimani M. Hepatocyte-specific ablation of spermine/spermidine-N1-acetyltransferase gene reduces the severity of CCl4-induced acute liver injury. Am J Physiol Gastrointest Liver Physiol. 2012; 303:G546–60. https://doi.org/10.1152/ajpgi.00431.2011 [PubMed] - 79. Zahedi K, Barone S, Kramer DL, Amlal H, Alhonen L, Jänne J, Porter CW, Soleimani M. The role of spermidine/spermine N1-acetyltransferase in endotoxin-induced acute kidney injury. Am J Physiol Cell Physiol. 2010; 299:C164–74. https://doi.org/10.1152/ajpcell.00512.2009 [PubMed]
- 80. Zahedi K, Barone S, Destefano-Shields C, Brooks M, Murray-Stewart T, Dunworth M, Li W, Doherty JR, Hall MA, Smith RD, Cleveland JL, Casero RA
Jr , Soleimani M. Activation of endoplasmic reticulum stress response by enhanced polyamine catabolism is important in the mediation of cisplatin-induced acute kidney injury. PLoS One. 2017; 12:e0184570. https://doi.org/10.1371/journal.pone.0184570 [PubMed] - 81. Zahedi K, Wang Z, Barone S, Prada AE, Kelly CN, Casero RA, Yokota N, Porter CW, Rabb H, Soleimani M. Expression of SSAT, a novel biomarker of tubular cell damage, increases in kidney ischemia-reperfusion injury. Am J Physiol Renal Physiol. 2003; 284:F1046–55. https://doi.org/10.1152/ajprenal.00318.2002 [PubMed]
- 82. Huang W, Eickhoff JC, Mehraein-Ghomi F, Church DR, Wilding G, Basu HS. Expression of spermidine/spermine N(1) -acetyl transferase (SSAT) in human prostate tissues is related to prostate cancer progression and metastasis. Prostate. 2015; 75:1150–59. https://doi.org/10.1002/pros.22996 [PubMed]
- 83. Brett-Morris A, Wright BM, Seo Y, Pasupuleti V, Zhang J, Lu J, Spina R, Bar EE, Gujrati M, Schur R, Lu ZR, Welford SM. The polyamine catabolic enzyme SAT1 modulates tumorigenesis and radiation response in GBM. Cancer Res. 2014; 74:6925–34. https://doi.org/10.1158/0008-5472.CAN-14-1249 [PubMed]
- 84. Maksymiuk AW, Tappia PS, Sitar DS, Akhtar PS, Khatun N, Parveen R, Ahmed R, Ahmed RB, Cheng B, Huang G, Bach H, Hiebert B, Ramjiawan B. Use of amantadine as substrate for SSAT-1 activity as a reliable clinical diagnostic assay for breast and lung cancer. Future Sci OA. 2018; 5:FSO365. https://doi.org/10.4155/fsoa-2018-0106 [PubMed]
- 85. Mounce BC, Poirier EZ, Passoni G, Simon-Loriere E, Cesaro T, Prot M, Stapleford KA, Moratorio G, Sakuntabhai A, Levraud JP, Vignuzzi M. Interferon-Induced Spermidine-Spermine Acetyltransferase and Polyamine Depletion Restrict Zika and Chikungunya Viruses. Cell Host Microbe. 2016; 20:167–77. https://doi.org/10.1016/j.chom.2016.06.011 [PubMed]
- 86. Roine A, Veskimäe E, Tuokko A, Kumpulainen P, Koskimäki J, Keinänen TA, Häkkinen MR, Vepsäläinen J, Paavonen T, Lekkala J, Lehtimäki T, Tammela TL, Oksala NK. Detection of prostate cancer by an electronic nose: a proof of principle study. J Urol. 2014; 192:230–34. https://doi.org/10.1016/j.juro.2014.01.113 [PubMed]


