Introduction
The Hedgehog (Hh) pathway is an evolutionarily conserved signaling axis essential for the regulation of diverse fundamental biological processes, including embryogenesis and tissue homeostasis [1]. Four major components, including Hh ligands, the Patched (PTCH) receptor, the Smoothened (SMO) intermediator and the zinc finger-containing Glioblastoma (GLI) transcription factor, are crucial for mediating signal transduction from the cell membrane to the nucleus. In the absence of Hh ligands, SMO function is inactivated by the PTCH receptor. GLI is converted to the repressor form, which blocks gene transcription. Upon the binding of Hh ligands to the PTCH receptor, SMO inhibition is relieved, leading to the nuclear accumulation of GLI and subsequent activation of Hh target genes [2].
The Hh pathway is crucial for mammalian embryonic development. Activation of the Hh pathway depends on the presence of a specialized cellular organelle known as the primary cilium, where the active SMO protein resides to promote the nuclear translocation of GLI proteins [3]. Normal ciliogenesis relies on a group of planar cell polarity (PCP) effector proteins, including the fuzzy planar cell polarity protein (FUZ), the inturned planar cell polarity protein (INTU) and the WD repeat containing planar cell polarity effector (WDPCP) [4, 5]. The intraflagellar transport (IFT) machinery governs the designated distribution of cargo proteins alongside the ciliary axoneme in support of ciliogenesis and activation of Hh signaling [6]. IFT-A and IFT-B are two subsets of protein complexes necessary for controlling retrograde and anterograde trafficking of cargo proteins [7]. The PCP effectors are indispensable for the initial ciliary recruitment and subsequent transport of IFT-A proteins. In mammalian embryos lacking these essential PCP effector genes, both IFT-A and IFT-B trafficking are impaired. In turn, failure of Hh signaling occurs due to ciliogenesis defects, and this leads to severe developmental retardation and early embryonic mortality [4, 5, 8].
Recently, emerging evidence has emphasized the involvement of the Hh pathway in human age-related disorders and cancers [9]. Oncogenic functions have been assigned to all PCP effectors [10–12]. Interestingly, INTU function was found to be related to the Hh pathway during carcinogenesis [12]. In basal cell carcinoma (BCC), INTU expression was aberrantly upregulated, accompanied by the induction of Hh signaling. The disruption of INTU in a BCC mouse model ameliorated tumorigenesis, and Hh activation was simultaneously suppressed [12]. Moreover, INTU was found to be functionally upstream of GLI transcription factors. Depletion of INTU attenuated the expression of GLI1 and blocked activation of Hh. However, overexpression of a constitutively activated GLI protein was capable of restoring Hh pathway activity in INTU-deficient cells [12]. These findings therefore suggest that INTU plays an important oncogenic function in BCC and highlight a synergistic mechanism involving INTU and Hh signaling in carcinogenesis. To date, the involvement of INTU in other cancer types remains elusive.
In this study, we carried out a comprehensive examination on the prognostic values of INTU in 21 different types of cancer. We found that the downregulation of INTU was associated with poor prognosis in lung adenocarcinoma (LUAD) and uterine corpus endometrial carcinoma (UCEC) patients. A group of Hh pathway-related genes, including INTU and intraflagellar transport 88 (IFT88), were enriched in LUAD and UCEC tumor samples. We further demonstrated positive correlations between INTU and IFT88 levels in both LUAD and UCEC samples, and identified multiple mechanisms spanning transcriptional, post-transcriptional and translational aspects that contribute to INTU and IFT88 downregulation in LUAD and UCEC samples. Taken together, we investigated at multifaceted levels the underlying mechanisms leading to the downregulation of Hh-related genes INTU and IFT88, and further highlighted the prognostic significance of this downregulation in LUAD and UCEC patients.
Results
Investigations on the prognostic values of INTU in multiple cancer types
The Kaplan-Meier plotter was initially used to assess the prognostic significance of INTU expression in 21 types of cancer. We found that INTU expression was significantly associated with the overall survival (OS) probabilities in patients with nine different cancer types. In liver hepatocellular carcinoma (LIHC) and lung squamous cell carcinoma (LUSC), patients with an increased level of INTU had poor OS probabilities. However, in the remaining cancer types, including esophageal adenocarcinoma (EAC), esophageal squamous cell carcinoma (ESCC), kidney renal papillary cell carcinoma (KIRP), LUAD, pancreatic ductal adenocarcinoma (PDAC), sarcoma (SARC) and UCEC, reduced levels of INTU were correlated with decreased OS probabilities in patients (Figure 1).
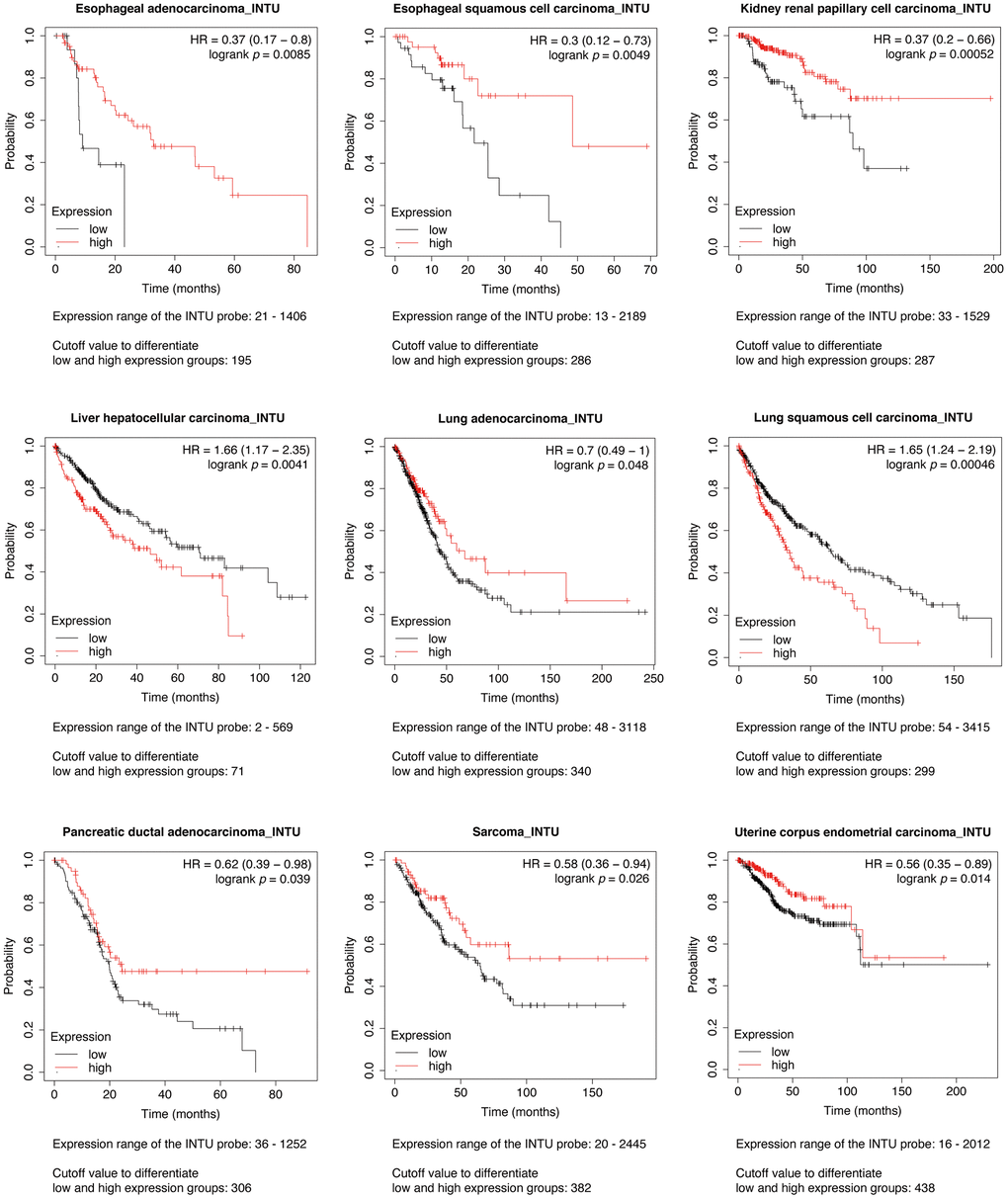
Figure 1. Evaluation of the prognostic significance of INTU mRNA level in different cancer types. Pan-cancer survival analysis was carried out to determine the relationship between INTU mRNA level and OS probabilities in 21 different types of cancer. Decreased INTU expression was found associated with poor prognosis in EAC, ESCC, KIRP, LUAD, PDAC, SARC and UCEC patients, whilst high level of INTU correlated with poor prognosis in LIHC and LUSC patients.
IFT88 downregulation was associated with the poor prognosis of LUAD and UCEC patients
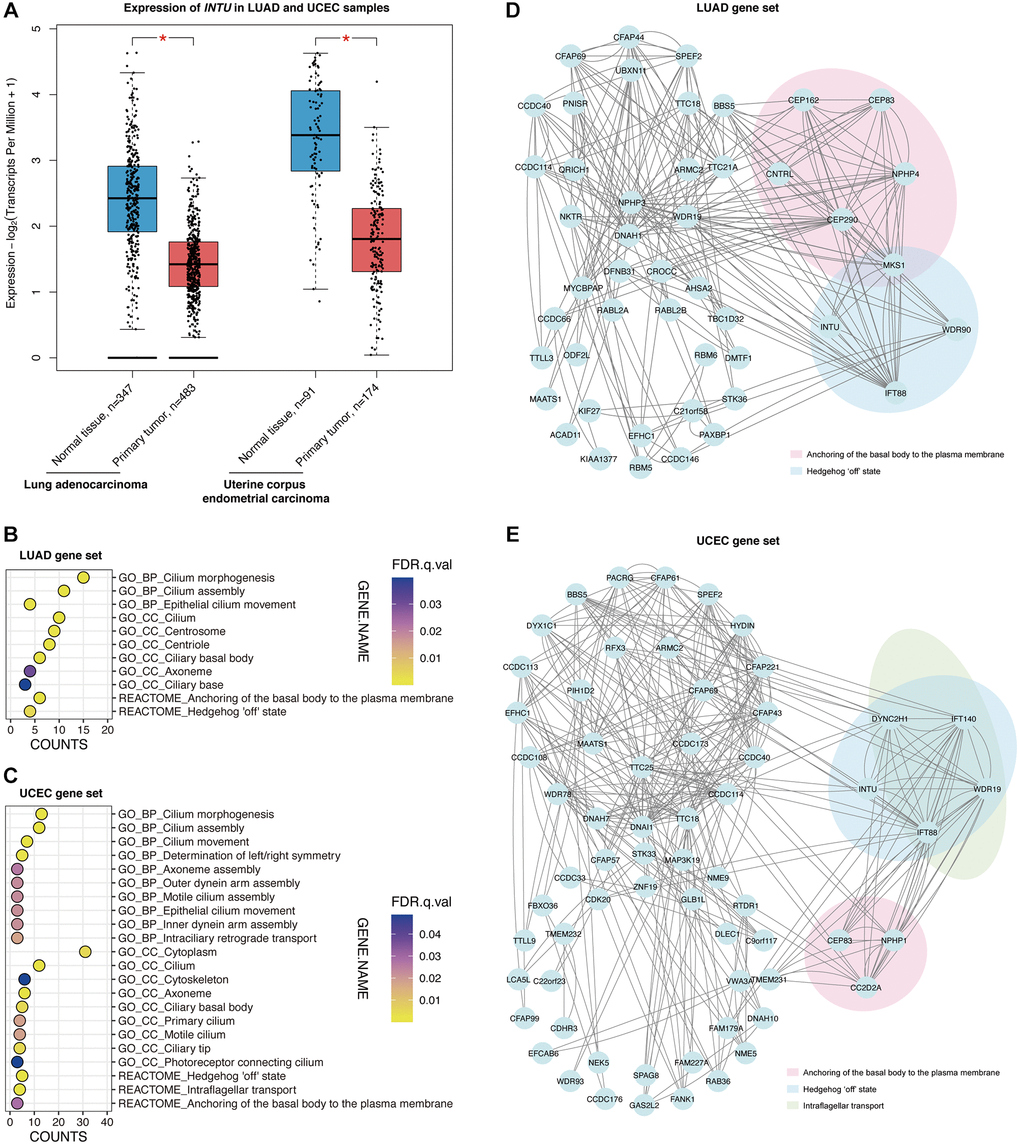
Similar to INTU, IFT88 was implicated in the “Hedgehog ‘off’ state” pathway in both the LUAD and UCEC gene sets (Figure 2D, 2E). In addition, INTU expression significantly correlated with the expression of IFT88 from LUAD and UCEC tumor samples (Figure 3A, 3B). Similar to INTU (Figure 1), we found that LUAD and UCEC patients with lowered IFT88 levels also showed decreased OS probabilities (Figure 3C, 3D). Taken together, these results highlight the significance of the downregulation of the Hh pathway-associated genes INTU and IFT88 in the prognosis of LUAD and UCEC patients. We therefore sought to delineate the underlying mechanisms of INTU and IFT88 downregulation in LUAD and UCEC tumor samples.
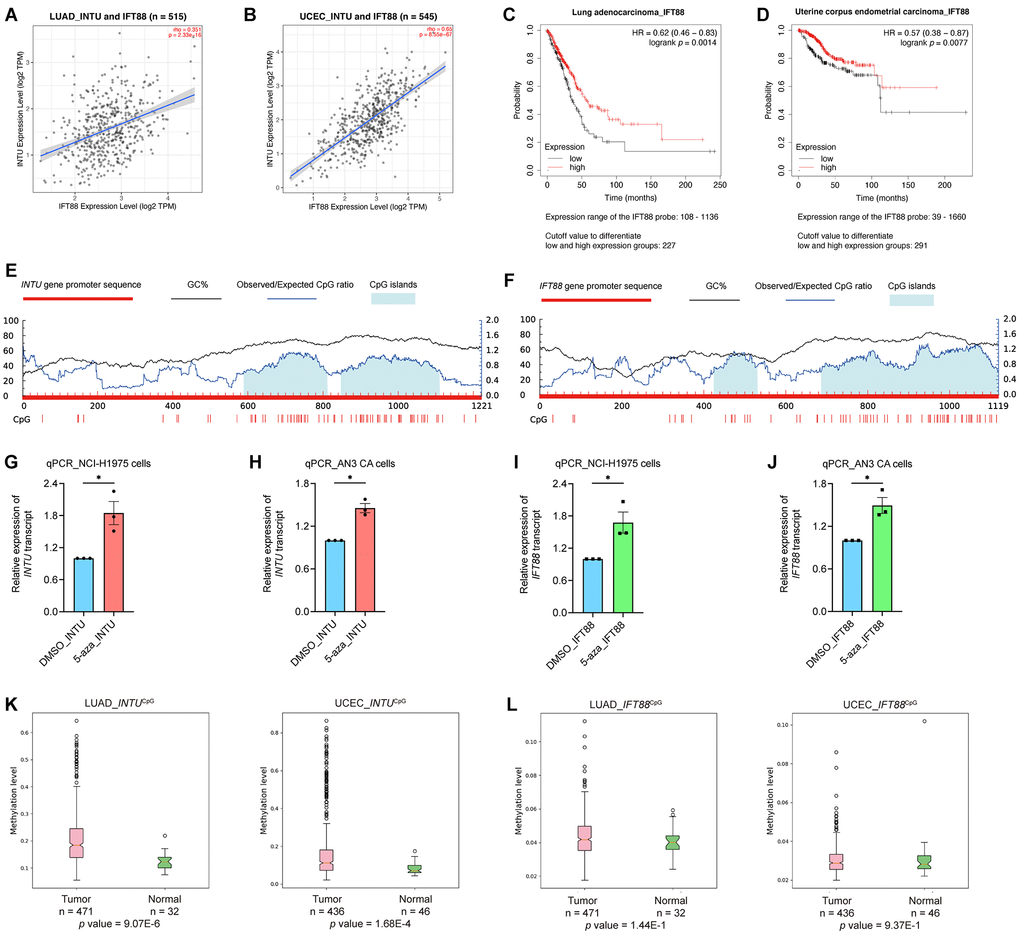
Figure 3. The expression of IFT88 correlated with INTU expression in LUAD and UCEC samples, and hypermethylation of INTUCpG was detected in LUAD and UCEC samples. (A, B) The expression of IFT88 positively correlated with the expression of INTU in LUAD (A) and UCEC (B) tumor samples. (C, D) Decreased expression of IFT88 was found associated with reduced OS probabilities in LUAD (C) and UCEC (D) patients. (E) Two putative CpG islands were identified in INTU gene promoter. (F) Two CpG islands were predicted in IFT88 gene promoter. (G, H) Treatment of 5-azacytidine induced INTU expression in NCI-H1975 (G) and AN3 CA (H) cells. n = 3 biological replicates. Each n represents an independent preparation of cell RNA samples. Error bars represent S.E.M. Statistical analysis was performed using two-tailed unpaired Student’s t-test. * denotes p < 0.05. (I, J) Treatment of 5-azacytidine induced IFT88 expression in NCI-H1975 (I) and AN3 CA (J) cells. n = 3 biological replicates. Each n represents an independent preparation of cell RNA samples. Error bars represent S.E.M. Statistical analysis was performed using two-tailed unpaired Student’s t-test. * denotes p < 0.05. (K) Hypermethylation of the INTUCpG was detected in LUAD and UCEC tumor samples. (L) No difference in IFT88CpG methylation levels were detected in LUAD and UCEC samples when compared to their respective normal control samples.
The methylation level of INTU gene promoter CpG islands was upregulated in LUAD and UCEC tumor samples
DNA methylation is a typical epigenetic modification through which gene expression is modulated [13]. In gene promoter region, CpG islands are regions with densely-accumulated CG dinucleotides, and methylation of CpG islands leads to gene silencing [14]. In each of the INTU and IFT88 gene promoter regions, two putative CpG islands were identified (Figure 3E, 3F). To investigate the association between DNA methylation and INTU and IFT88 gene expression, the LUAD (NCI-H1975) and UCEC (AN3 CA) cell lines were treated with 5-azacytidine, a DNA methyltransferase inhibitor [15], the INTU and IFT88 levels were subsequently detected. We found that upon treatment of 5-azacytidine, both INTU and IFT88 levels were upregulated in NCI-H1975 and AN3 CA cells, suggesting a negative correlation between DNA methylation and INTU and IFT88 gene expression (Figure 3G–3J). The methylation levels of INTUCpG and IFT88CpG were further examined in LUAD and UCEC tumor samples. We found that the methylation level of INTUCpG was significantly upregulated in both LUAD and UCEC tumor samples (Figure 3K). When compared with normal tissues, no significant change in IFT88CpG methylation levels was detected in LUAD and UCEC tumor samples (Figure 3L). This suggests that hypermethylation of INTUCpG potentially contributes to the reduction of INTU levels in LUAD and UCEC samples.
Involvement of the transcriptional factor TCF4 in the modulation of INTU and IFT88 levels
Transcription factors are a set of regulatory proteins that bind to gene promoter DNA sequences and modulate gene transcription [16]. A total of nine communal transcription factors, including core-binding factor subunit beta (CBFB), histone deacetylase 2 (HDAC2), transcription factor jun-D (JUND), serum response factor (SRF), small ubiquitin-like modifier 2 (SUMO2), TATA-box binding protein associated factor 1 (TAF1), TATA box binding protein (TBP), transcription factor 4 (TCF4) and yin yang 1 (YY1), were identified in the INTU and IFT88 gene promoters (Figure 4A). Next, we evaluated the expression of these transcription factors in LUAD and UCEC samples. Among these nine transcription factors, we found that only TCF4 (Figure 4B) expression was significantly downregulated in both LUAD and UCEC tumor samples (Figure 4C and Supplementary Figure 2). The TCF4 protein level was further found to be significantly reduced in both LUAD (Figure 4D) and UCEC (Figure 4E) samples. More importantly, the expression of TCF4 was positively correlated with the expression of INTU and IFT88 in LUAD and UCEC samples (Figure 4F, 4G). Similar to INTU (Figure 1) and IFT88 (Figure 3C, 3D), the reduced levels of TCF4 were associated with decreased OS probabilities in LUAD and UCEC patients (Figure 4H, 4I). Altogether, TCF4 was identified as a putative upstream regulator in controlling the expression of INTU and IFT88 in LUAD and UCEC tumor samples.
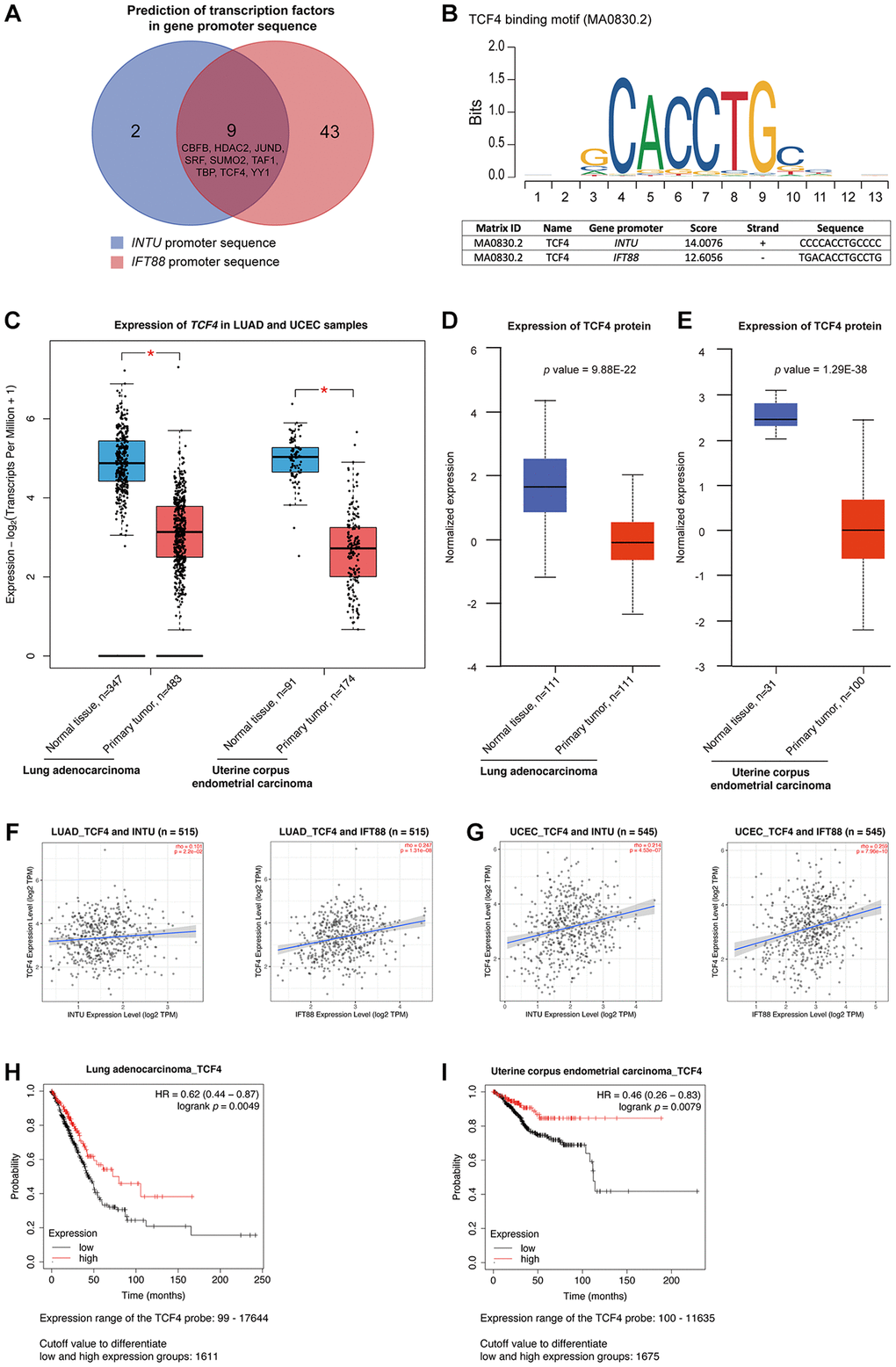
Figure 4. TCF4 was identified as a potential transcription factor that mediates INTU and IFT88 downregulation in LUAD and UCEC tumor samples. (A) Nine common transcription factors, including CBFB, HDAC2, JUND, SRF, SUMO2, TAF1, TBP, TCF4 and YY1 were predicted in INTU and IFT88 gene promoters. (B) Illustration of TCF4 binding consensus sequence and the putative TCF4 binding sites in INTU and IFT88 promoter sequence. (C) The TCF4 transcript level was downregulated in LUAD and UCEC tumor samples. * denotes p < 0.05. (D, E) The protein level of TCF4 was downregulated in LUAD (D) and UCEC (E) tumor samples. (F, G) The expression of TCF4 positively correlated with the expression of INTU and IFT88 in LUAD (F) and UCEC (G) tumor samples. (H, I) The LUAD (H) and UCEC (I) patients with lowered level of TCF4 showed reduced OS probabilities.
We further provided experimental evidence in support of our findings. Chromatin immunoprecipitation was performed to investigate the interaction between TCF4 protein and INTU and IFT88 gene promoters. We found that in both the NCI-H1975 and AN3 CA cells, the binding of TCF4 to INTU and IFT88 gene promoters was detected (Figure 5A, 5B). The transcriptional regulatory function of TCF4 on INTU and IFT88 gene expression was subsequently determined. When endogenous TCF4 was knocked down in NCI-H1975 and AN3 CA cells, the transcript levels of INTU and IFT88 were downregulated (Figure 5C, 5D). These data further support the positive correlation between TCF4 and INTU and IFT88 levels in LUAD and UCEC samples (Figure 4F, 4G).
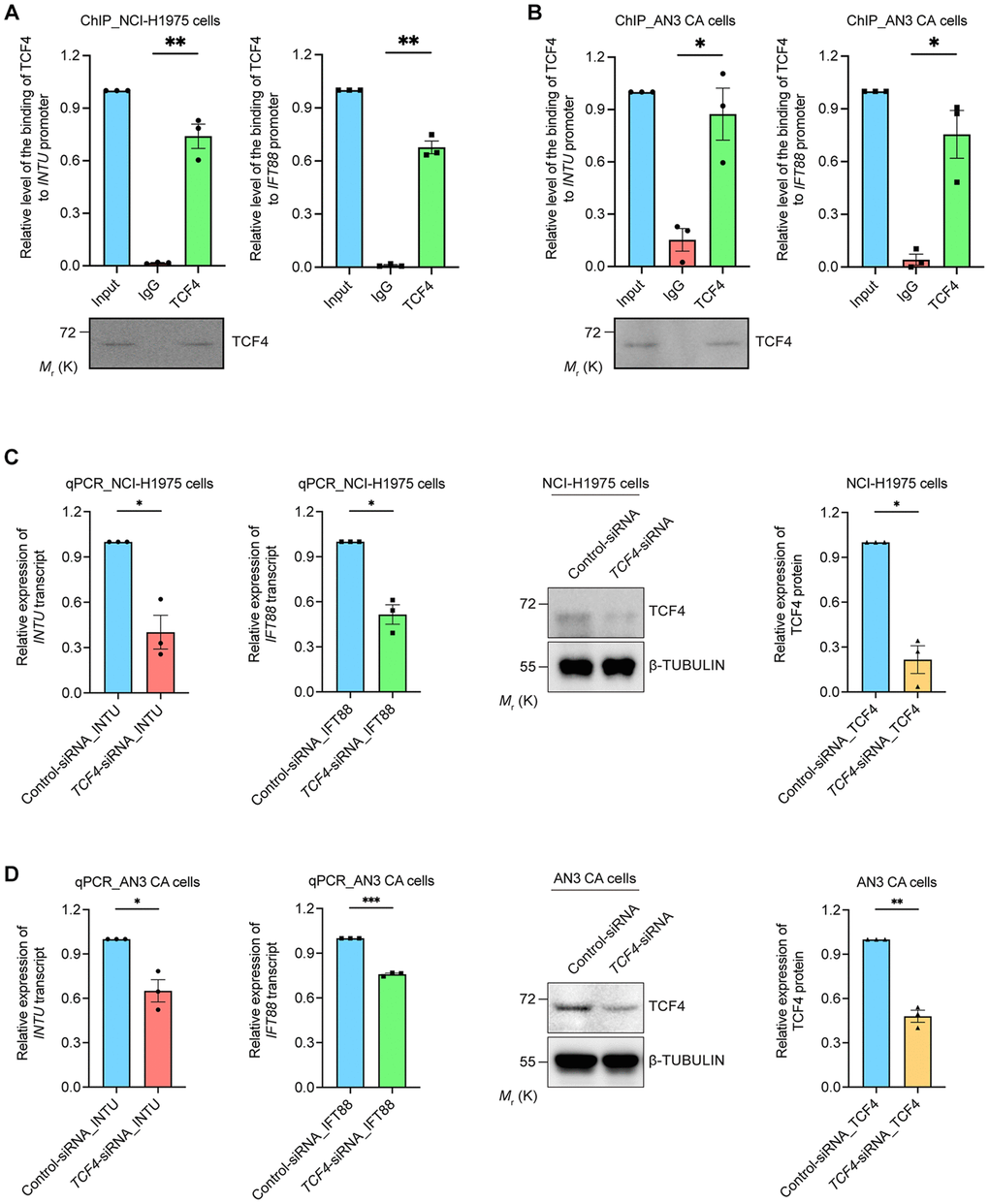
Figure 5. TCF4 interacted with INTU and IFT88 promoters and mediated their gene expression. (A, B) The binding between TCF4 and INTU and IFT88 promoters was detected in NCI-H1975 (A) and AN3 CA (B) cells. n = 3 biological replicates. Each n represents an independent preparation of ChIP samples. Error bars represent S.E.M. Statistical analysis was performed using two-tailed unpaired Student’s t-test. * denotes p < 0.05 and ** denotes p < 0.01. (C, D) Knockdown of TCF4 downregulated the transcript levels of INTU and IFT88 in NCI-H1975 (C) and AN3 CA (D) cells. n = 3 biological replicates. Each n represents an independent preparation of RNA and protein samples. Error bars represent S.E.M. Statistical analysis was performed using two-tailed unpaired Student’s t-test. *denotes p < 0.05, **denotes p < 0.01 and ***denotes p < 0.001.
Identification of MALAT1 as an upstream lncRNA
We next examined upstream lncRNAs using the DIANA-LncBase v3 database. A total of 63 lncRNAs were obtained, and their expression levels in LUAD and UCEC samples were evaluated (Figure 7A). The levels of four of the 63 lncRNAs, including HOXA transcript antisense RNA, myeloid-specific 1 (HOTAIRM1), KMT2E antisense RNA 1 (KMT2E-AS1), metastasis associated lung adenocarcinoma transcript 1 (MALAT1) and nuclear paraspeckle assembly transcript 1 (NEAT1), were significantly downregulated in LUAD and UCEC tumor samples when compared with their respective normal tissues (Figure 7B and Supplementary Figure 4A). The prognostic significance of these four lncRNAs were further evaluated. We found that MALAT1, but not HOTAIRM1 and NEAT1, showed prognostic significance in both LUAD and UCEC patients (Figure 7C, 7D and Supplementary Figure 4B). Similar to what was detected regarding INTU (Figure 1) and IFT88 (Figure 3C, 3D), the reduced level of MALAT1 contributed to poor OS probabilities in LUAD and UCEC patients (Figure 7C, 7D). Overexpression of hsa-miR-212 did not modulate MALAT1 levels in NCI-H1975 and AN3 CA cells (Figure 6G, 6H). In contrast, MALAT1 has been shown to target hsa-miR-212-3p in two independent studies (Figure 7E) [22, 23]. Moreover, the expression of MALAT1 was found to be positively associated with the mRNA levels of INTU and IFT88 in LUAD and UCEC tumor samples (Figure 7F, 7G). To further validate the regulatory function of MALAT1 on the expression of INTU and IFT88, MALAT1 was overexpressed in NCI-H1975 or AN3 CA cells (Figure 7H, 7I). We found that in MALAT1-overexpressing cells, the protein levels of INTU and IFT88 were simultaneously increased (Figure 7H, 7I). These results therefore confirm the role of MALAT1 in regulating the expression of INTU and IFT88 in lung and endometrial cancer cells.
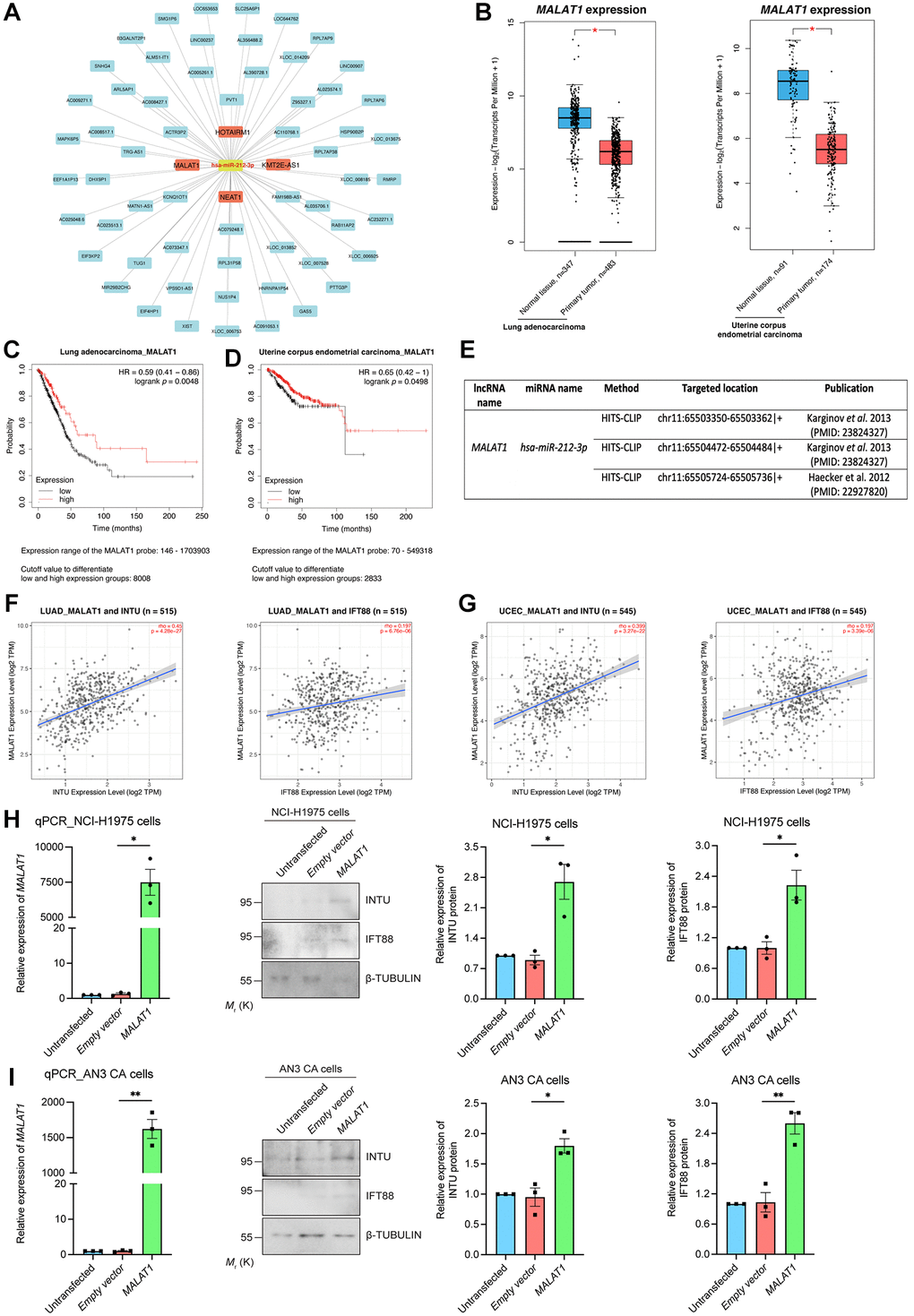
Figure 7. Identification of MALAT1 as a communal lncRNA mediating INTU and IFT88 expression in LUAD and UCEC samples. (A) Identification of the lncRNAs against hsa-miR-212-3p and construction of the lncRNA-miRNA regulatory network. (B) The expression of MALAT1 was significantly downregulated in LUAD and UCEC tumor samples. (C, D) Decreased level of MALAT1 was found associated with poor prognosis in LUAD (C) and UCEC (D) patients. (E) The MALAT1 was identified as targeted lncRNA against hsa-miR-212-3p from two independent studies. (F, G) The expression of MALAT1 positively associated with the expression of INTU and IFT88 in LUAD (F) and UCEC (G) tumor samples. (H, I) Overexpression of MALAT1 caused the upregulation of INTU and IFT88 protein levels in NCI-H1975 (H) and AN3 CA (I) cells. n = 3 biological replicates. Each n represents an independent preparation of RNA and protein samples. Error bars represent S.E.M. Statistical analysis was performed using two-tailed unpaired Student’s t-test. * denotes p < 0.05.
Discussion
In this study, we demonstrated that the downregulation of the INTU and IFT88 was correlated with reduced survival probabilities in LUAD and UCEC patients (Figures 1, 3C, 3D). We next sought to explore the driving forces causing this downregulation in LUAD and UCEC tumor samples, and identified multifaceted mechanisms in the DNA, RNA and protein levels contributing to INTU and IFT88 downregulation (Figures 3–7). This study provides a comprehensive mechanistic investigation regarding INTU and IFT88 downregulation in cancer, and further highlights the involvement of Hh signaling in carcinogenesis.
The dysregulation of Hh signaling has been documented in multiple types of cancer. In BCC, medulloblastoma and rhabdomyosarcoma, mutations in Hh-related genes activate Hh signaling in support of the over-proliferation and tissue invasion of cancer cells [24–26]. In addition to genetic mutation, epigenetic modification also contributes to aberrant Hh signaling in tumor samples. For example, the hypermethylation of hedgehog-interacting protein (HHIP), a gene encodes for a negative regulator of Hh signaling, was determined in pancreatic cancer samples. This leads to the reduced expression of HHIP, followed by the upregulation of Hh signaling [27]. In the majority of solid tumors, including colorectal cancer (CRC), the mutation in Hh-related genes was rarely detected [28]. Interestingly, in CRC, the stromal Hh pathway targets were found downregulated despite the increased expression of Hh ligand. This might be due to the insensitivity of stromal cells to epithelial Hh ligand, or the impairment of tissue architecture in tumor stroma [29, 30]. In addition, the restoration of stromal Hh signaling markedly alleviated tumorigenesis, whereas inhibition of Hh signaling exacerbated tumor progression [31]. These findings taken together suggest multiple mechanisms in contributing to the dysregulation of Hh signaling in different cancers, and also highlight the oncogenic role of Hh signaling. Here, we reported two subsets of enriched Hh pathway-associated genes with similar downregulation expression patterns in LUAD and UCEC tumor samples (Figure 2D, 2E). We further showed that lowered levels of INTU, IFT88 or MKS1 were correlated with decreased OS probabilities in LUAD patients, while UCEC patients with reduced INTU, IFT88 or IFT140 levels had a poor prognosis (Figures 1, 3C, 3D, Supplementary Figure 5A, 5B). INTU was found to be necessary for the ciliary recruitment of IFT-A proteins, and MKS1 functionally associates with IFT complexes in mediating the transport of cargo proteins to support ciliary outgrowth [8, 32]. Depletion of INTU or IFT machinery components leads to ciliogenesis defects [12, 33, 34]. Importantly, the loss of ciliary structures has been recorded in lung and endometrial cancer patient samples [35, 36]. Moreover, stimulation of ciliogenesis has been reported to combat against lung cancer cell proliferation, invasion and epithelial-mesenchymal transition [37]. These findings therefore indicate that disruption of Hh signaling components might cause ciliogenesis defects, which in favour of oncogenesis in LUAD and UCEC tissues.
TCF4 belongs to the helix–loop–helix (HLH) family of proteins ubiquitously expressed throughout different human tissues [38]. A basic residue group on the TCF4 N-terminus is critical for its DNA-binding function, while the C-terminal HLH domain mediates the dimerization of TCF4 when it binds to DNA [39]. Several studies have demonstrated the association between TCF4 function and DNA methylation [40, 41]. Specifically, TCF4 was correlated with DNA hypomethylation in mammalian epithelial stem cells. Upon conditional knockout of TCF4, the TCF4-bound differentially methylated DNA sequence was found to be strongly methylated [41]. Interestingly, when looking into the TCF4 binding site in the INTU promoter sequence, we found that the TCF4 site resides in the INTUCpG (highlighted in Supplementary Table 2). The methylation level of INTUCpG was upregulated in LUAD and UCEC samples (Figure 3G). Reduction of the TCF4 protein level was recorded in both LUAD and UCEC tumor samples (Figure 4D). Such attenuation of the TCF4 level might result in INTUCpG hypermethylation, which in turn leads to the downregulation of INTU expression in LUAD and UCEC samples. Meanwhile, TCF4 did not associate with IFT88CpG (Supplementary Table 2), and no significant change in IFT88CpG methylation level was detected in LUAD and UCEC samples (Figure 3H).
Genetic mutations in TCF4 have been reported in neurological disorders, including Fuchs’s corneal dystrophy [42], Pitt–Hopkins syndrome [43], and schizophrenia [44], as well as non-neurological diseases, including primary sclerosing cholangitis [45] and sporadic Sonic Hedgehog-associated medulloblastoma (SHH MB) [46]. Functional analysis was carried out on mutant TCF4 proteins harboring the mutations identified from SHH MB patients. Experimental findings highlighted the loss-of-function behind these TCF4 mutations, as exemplified by the fact that mutant TCF4 proteins failed to inhibit the proliferation of medulloblastoma cells, unlike the wild-type TCF4 protein [40, 47]. We also found a nonsense mutation at the arginine 174 residue (R174*) on the TCF4 protein in eight UCEC patient samples (Supplementary Figure 6A, 6B), and the R174 site was conserved across different species (Supplementary Figure 6C). This nonsense mutation generates a truncated TCF4 protein which lacks the C-terminal HLH motif that is crucial for mediating gene transcription, suggesting the loss of TCF4 transactivating function due to the presence of such a mutation. This could serve as another mechanism leading to the downregulation of Hh-related genes in UCEC tumor samples. Interestingly, a similar TCF4R174* mutation was previously reported in patients with SHH MB and Pitt–Hopkins syndrome [46, 47], suggesting that the communal loss-of-function mechanism is involved in a broad spectrum of human disorders.
miRNAs and lncRNAs are two major subtypes of ncRNAs associated with the well-documented ceRNA mechanism that is essential for controlling gene expression at a post-transcriptional level [18, 48]. We identified a novel and communal MALAT1-hsa-miR-212-3p regulatory network that downregulated INTU and IFT88 expression in LUAD and UCEC samples (Figure 5). The recurrent fusion of MALAT1 with the GLI1 gene was reported in patients with gastroblastoma and plexiform fibromyxoma [49, 50]. This MALAT1-GLI1 fusion mutation activated Hh signaling and consequently led to malignant tumor formation, suggesting a relationship between MALAT1 function and Hh signaling activity [49, 51, 52]. In this study, we further highlighted the involvement of MALAT1 in regulating Hh pathway-associated genes. We found that in addition to INTU and IFT88, other Hh-related genes with similar expression profiles as INTU were also enriched in LUAD and UCEC samples (Figure 2D, 2E). Interestingly, the expression of hsa-miR-212-3p was negatively associated with the expression of MKS1 and WDR90 in LUAD samples (Supplementary Figure 7A) and DYNC2H1, IFT140 and WDR19 in UCEC samples (Supplementary Figure 7B). Positive correlations were determined between MALAT1, MKS1 and WDR90 in LUAD samples and MALAT1, DYNC2H1 and WDR19 in UCEC samples (Supplementary Figure 7C, 7D). Taken together, these findings emphasize the MALAT1-hsa-miR-212-3p network as a master upstream regulator targeting downstream Hh pathway-associated genes in LUAD and UCEC tumor samples. In addition, MALAT1 binds to active chromatin sites and regulates gene transcription by recruiting chromatin modifiers or transcription regulators to specific genomic loci [53, 54]. This might be an alternative mechanism that accounts for MALAT1’s regulation on Hh pathway-associated gene expression and is worthy to be further investigated.
Several recent studies have reported the anti-tumorigenesis role of TCF4 in colorectal carcinoma and SHH MB [47, 55]. TCF4 is capable of attenuating the proliferation of colon cancer and medulloblastoma cells, whereas loss of TCF4 exerts the opposite effect, favoring tumorigenesis [47, 56, 57]. The effect of MALAT1 on tumor cell growth and invasion is controversial. Although the oncogenic functions of MALAT1 have been reported in malignancies such as colorectal and liver cancer [58, 59], more recent studies have highlighted the tumor suppressive role of MALAT1 against the growth and metastasis of glioma and breast cancer cells [60, 61]. These findings taken together suggest a cancer type-dependent role of MALAT1 in tumorigenesis. Given the functions of TCF4 and MALAT1 as tumor suppressors, targeting the functional elevation of TCF4 and MALAT1 could be therapeutically beneficial against tumorigenesis. In fact, the identification of small molecule drugs aimed at stimulating TCF4 function is now under investigation (Pitt Hopkins Research Foundation; https://pitthopkins.org/portfolio-item/pilot-study-to-identify-small-molecule-activators-of-tcf4-as-a-treatment-for-pitt-hopkins-syndrome/). Meanwhile, different small molecule activators for MALAT1 have been reported (Table 1). A combinatorial drug treatment has been demonstrated as an effective therapeutic strategy in combating carcinogenesis [62–64]. The use of both TCF4 and MALAT1 activators in the treatment against LUAD and UCEC would be an interesting topic worthy of further exploration.
Table 1. Summary of MALAT1 small molecule activators.
| lncRNA | Small molecules | Effect on lncRNA expression | Approved by FDA | Validated by experiments | Validation method | Experimental material | References |
| MALAT1 | Carboplatin + Docetaxel | Up-regulation | Yes | Yes | Quantitative real-time PCR | Ovarian cancer cell line | [81] |
| MALAT1 | Quercetin | Up-regulation | Yes | Yes | Quantitative real-time PCR | Rheumatoid arthritis fibroblast-like synoviocytes | [82] |
In summary, we showed that the downregulation of the Hh pathway-associated genes INTU and IFT88 was correlated with poor prognosis in LUAD and UCEC patients. Moreover, we demonstrated novel TCF4 and ncRNA-involved mechanisms that contribute to the downregulation of INTU and IFT88 in LUAD and UCEC tumor samples (Figure 8). We further propose that a treatment strategy that simultaneously targets TCF4 and MALAT1 to enrich INTU and IFT88 might be a promising therapeutic intervention against LUAD and UCEC.
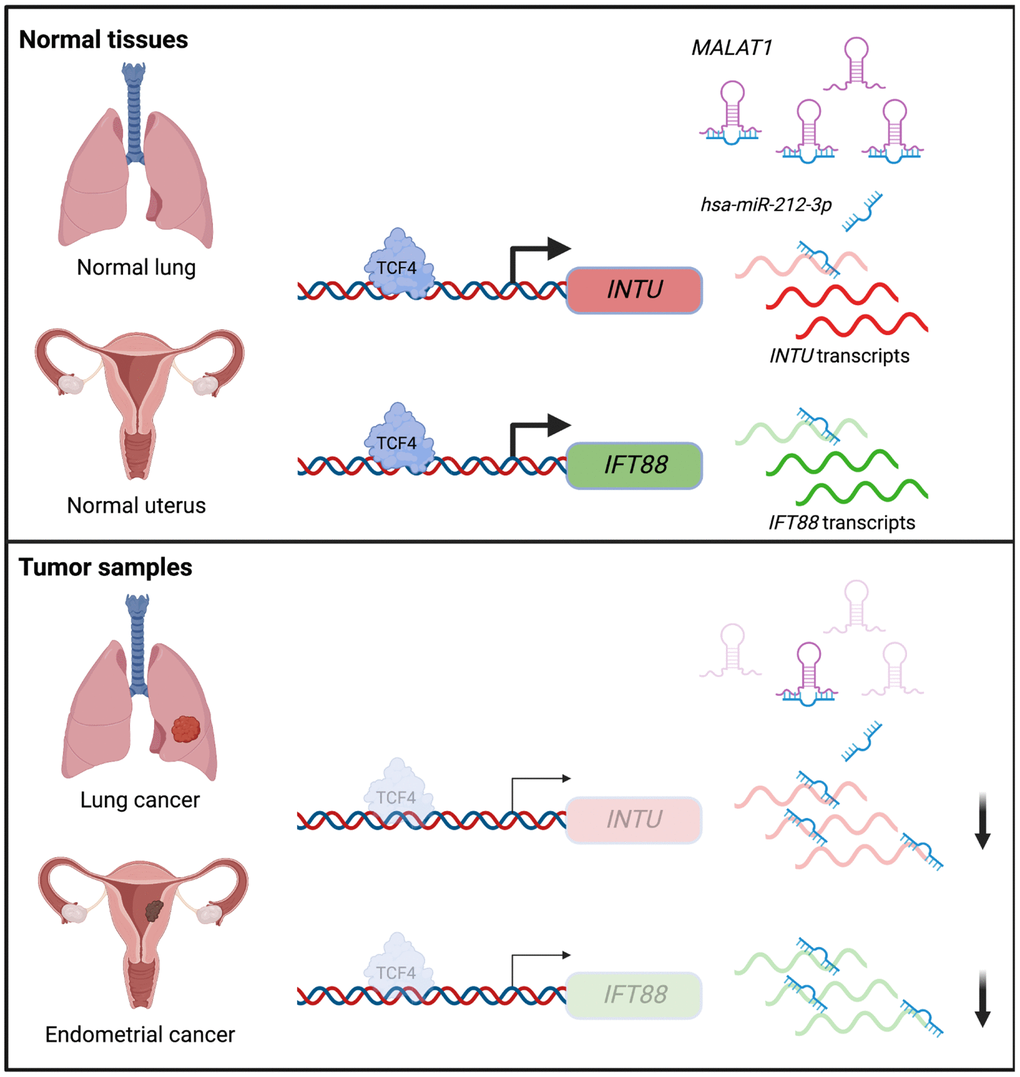
Figure 8. Illustration of the underlying mechanisms that contribute to INTU and IFT88 downregulation in the lung and endometrial cancers.
Materials and Methods
Kaplan-Meier plotter analysis
The pan-cancer analysis function (https://kmplot.com/analysis/index.php?p=service&cancer=pancancer_rnaseq) from the Kaplan-Meier plotter database was used to evaluate the prognostic significance of INTU mRNA expression in 21 different types of cancer [65]. The prognostic significance of Hh pathway-associated genes IFT88, MKS1, WDR90, IFT140, DYNC2H1 and WDR19 was also evaluated using LUAD and UCEC patient survival data from Kaplan-Meier plotter database. Similar approach was used to determine the prognostic value of TCF4 and lncRNAs. The OS probabilities of cancer patients were assessed using the Kaplan-Meier survival plots, and logrank p < 0.05 indicates that the association between gene/lncRNA expression and patient survival probability is statistically significant.
GEPIA2 database analysis
GEPIA2 (http://gepia2.cancer-pku.cn/#index) is an online database that provides gene expression profiling and interactive analyses in primary tumor and normal tissue samples on the basis of data from The Cancer Genome Atlas (TCGA) and the Genotype-Tissue Expression (GTEx) projects [66]. The expression of INTU, housekeeping genes, different transcription factors and different lncRNAs were determined in LUAD and UCEC primary tumor samples and their respective normal tissues. The p < 0.05 was considered as statistically significant. The top 100 genes with similar expression pattern as INTU in LUAD or UCEC tumor samples were also selected using the “Similar Gene Detection” function from GEPIA2. The detailed gene lists are included in Supplementary Table 1.
Gene ontology and Reactome pathway analysis
The INTU and top 100 genes with similar expression pattern obtained from GEPIA2 database were input to Database for Annotation, Visualization and Integrated Discovery (DAVID) v6.8 (https://david.ncifcrf.gov/home.jsp) to analyze their enriched Gene Ontology (GO) terms and Reactome pathways [67]. The biological processes (BP) and cellular components (CC) were included in the GO enrichment analysis. The false discovery rate (FDR) q-value < 0.05 was used as selection criteria for significantly enriched GO terms and Reactome pathways.
Protein-protein interaction (PPI) network analysis
The construction of PPI network was performed using STRING v11.5 database (https://string-db.org/) [68]. The genes with similar expression pattern obtained from GEPIA2 database were input to STRING database, and the PPI network was constructed based on the sources of “Co-expression”, “Databases”, “Experiments”, “Gene Fusion”, “Neighborhood” and “Textmining” with minimum required interaction score of medium confidence. The Cytoscape v3.8.0 was used to visualize the constructed PPI network [69].
Prediction of CpG islands and transcription factor binding sites within gene promoter sequences
INTU and IFT88 promoter sequences were withdrawn from GenBank under the accession numbers NC_000004.12 and NC_000013.11, respectively. The CpG islands were predicted using MethPrimer 2.0 (http://www.urogene.org/cgi-bin/methprimer2/MethPrimer.cgi) software [70]. The transcription factor binding sites were predicted using Animal Transcription Factor Database 3.0 (AnimalTFDB3.0; http://bioinfo.life.hust.edu.cn/AnimalTFDB/#!/) [71]. The “q-value < 0.05” and “Score > 20” were used as filtering parameters to select potential transcription factors for INTU and IFT88. The TCF4 binding site was further validated using JASPAR (https://jaspar.genereg.net/) database [72]. The relative profile score threshold equals to 90%, and “Score > 12.5” and “Relative score > 0.92” were used as filtering parameters. The detailed INTU and IFT88 promoter sequences are listed in Supplementary Table 2, the putative CpG islands are shown in blue and the TCF4 binding sites are highlighted.
MethHC2.0 database analysis
MethHC2.0 is a web-based resource that provides analysis on the methylation levels of gene regions, including CpG islands, from different types of cancer [73]. The methylation levels of INTU and IFT88 CpG islands were scrutinized using methylome data from LUAD and UCEC tumor samples and their respective normal samples.
UALCAN database analysis
UALCAN (http://ualcan.path.uab.edu/analysis-prot.html) is an interactive online resource that enables the analysis of protein expression based on the Clinical Proteomic Tumor Analysis Consortium Confirmatory/Discovery datasets [74]. In this study, the TCF4 protein expression from LUAD and UCEC tumor samples and their respective normal tissues was analyzed.
Candidate miRNA and lncRNA prediction
The list of miRNAs that target INTU and IFT88 was obtained using DIANA-Tarbase v8 database (https://dianalab.e-ce.uth.gr/html/diana/web/index.php?r=tarbasev8%2Findex) [19]. The “Species = Homo Sapiens” and “Validated as Positive” were used as filtering parameters. The miRNA-target genes regulatory network was constructed using Cytoscape v3.8.0.
The list of lncRNAs that target different miRNAs was obtained using DIANA-LncBase v3 database (https://diana.e-ce.uth.gr/lncbasev3) [75]. The “Species = Homo Sapiens”, “miRNA Conf. Level = High” and “Validated as Positive” were used as filtering parameters. The lncRNA-miRNA regulatory network was constructed using Cytoscape v3.8.0.
ENCORI database analysis
ENCORI (http://starbase.sysu.edu.cn/) is an online database that determines the correlation between miRNA level and target gene expression [76]. The expression correlation between hsa-miR-210-3p/hsa-miR-212-3p and different enriched Hh-related genes was analyzed using “miRNA-Target CoExpression” module from the “Pan-Cancer” function. The p-value < 0.05 was considered as statistically significant.
TIMER2.0 database analysis
The TIMER2.0 (http://timer.cistrome.org/) is an online web server that enables the detection of gene expression correlation [77]. The correlation among different Hh-related genes, and correlation between Hh-related genes and MALAT1 in LUAD and UCEC tumor samples were evaluated using the “Gene_Correlation” module. No adjustment was made, and p < 0.05 was considered as statistically significant.
cBioPortal database analysis
cBioPortal v3.7.3 is a comprehensive web resource that enables the visualization and analysis of cancer genomic mutation data (https://www.cbioportal.org/) [78, 79]. The missense and nonsense mutation profiles in INTU, IFT88 and TCF4 genes were obtained from Lung Adenocarcinoma (TCGA, PanCancer Atlas, 566 samples) and Uterine Corpus Endometrial Carcinoma (TCGA, PanCancer Atlas, 529 samples) datasets.
D-lnc database
The D-lnc database (http://www.jianglab.cn/D-lnc/index.jsp) is a comprehensive platform that summarizes the lncRNA-targeting drugs based on the experimental evidence and computational predictions [80]. The “Species = Homo Sapiens” and “lncRNA = MALAT1” were used to select potential small molecules that target MALAT1.
Mammalian cell culture
The human endometrial adenocarcinoma cell line AN3 CA was a kind gift from Prof. Chi Chiu Wang (Department of Obstetrics and Gynecology, The Chinese University of Hong Kong, China). The human lung adenocarcinoma cell line NCI-H1975 (CRL-5908™) was obtained from American Type Culture Collection. Both cell lines were cultured using Dulbecco’s Modified Eagle’s Medium (11995065, Thermo Fisher Scientific) supplemented with 10% fetal bovine serum (F7524, Sigma-Aldrich) and 1% penicillin-streptomycin solution (15140122, Thermo Fisher Scientific). The cells were maintained in a 37°C humidified cell culture incubator supplemented with 5% CO2.
Plasmid, microRNA and siRNA transfection
The pcDNA-MALAT1 plasmid was a kind gift from Prof. Huating Wang (Department of Orthopaedics and Traumatology, The Chinese University of Hong Kong, China). Cells were transfected with 0.5 μg pcDNA-MALAT1 plasmid with 0.5 μl lipofectamine 2000 (11668019, Thermo Fisher Scientific). The RNA or protein samples were harvested 24 h post transfection. The hsa-miR-NC (4464058) and hsa-miR-212 (4464066) were synthesized by Thermo Fisher Scientific. Cells were transfected with 20 pmol microRNAs with 2 μl lipoRNAiMAX (13778150, Thermo Fisher Scientific). Cell culture medium and transfection mixtures were refreshed every 24 h, and protein samples were harvested 72 h post transfection. The Control-siRNA, 5′-UUCUCCGAACGUGUCACGUTT-3′ and TCF4-siRNA, 5′-CUAUCAGUAUUCUAGCAAUAATT-3′ were synthesized by Sangon Biotech (Shanghai) Co., Ltd. Cells were transfected with 20 pmol siRNAs with 2 μl lipoRNAiMAX. Cell culture medium and transfection mixtures were refreshed every 24 h, and RNA or protein samples were harvested 72 h post transfection.
Drug treatment
The NCI-H1975 and AN3 CA cells were treated with 2 μM 5-azacytidine (A2385, Sigma-Aldrich). The treatment lasted 72 h, with medium and drug refreshed every 24 h.
Chromatin immunoprecipitation
The chromatin immunoprecipitation (ChIP) assay was performed using Pierce™ Magnetic ChIP Kit (26157, Thermo Fisher Scientific). The experimental procedures were carried out following the manufacturer’s instructions. Two micrograms of anti-TCF4 antibody (ab217668, abcam) were used for the immunoprecipitation, while the same amount of normal rabbit IgG (I-1000, Vector Laboratories, lnc.) was used as negative control. Twenty nanograms of recovered genomic DNAs from each of input, normal rabbit IgG and TCF4 immunoprecipitated samples were used in the following real-time PCR to analyze the levels of INTU and IFT88 promoter fragments. The primers used were INTU promoter-F, 5′-CAGCCTGGACTTCGCGAG-3′; INTU promoter-R, 5′-TGAAGGCGGTGGTGTCAG-3′; IFT88 promoter-F, 5′-AAAACGGACACCTTAAGCGC-3′ and IFT88 promoter-R, 5′-CTTGTGAACCTTGGAAGCCC-3′.
RNA extraction, reverse transcription and real-time PCR
The total RNA was isolated from cultured cells using TRIzol™ reagent (15596018, Thermo Fisher Scientific). The reverse transcription was performed using ImProm-II™ Reverse Transcription System (A3803, Promega) and random hexamer (N8080127, Thermo Fisher Scientific) according to the manufacturers’ instructions. Quantitative real-time PCR was performed using SYBR™ Green PCR Master Mix (4309155, Thermo Fisher Scientific) on the Bio-Rad CFX96 system. Relative gene expression was determined via normalizing against β-ACTIN using the 2−ΔΔCT method. Primers used in this study were INTU-F, 5′-CGCATAGATGAACGGCTAGC-3′; INTU-R, 5′-AGCGTTCTTCTGCATGTTGG-3′; IFT88-F, 5′-CTGCAACCAATCTCTCAGCC-3′; IFT88-R, 5′-GCGGCCTTCTCATAATCACC-3′; MALAT1-F, 5′-ATGCGAGTTGTTCTCCGTCT-3′; MALAT1-R, 5′-TATCTGCGGTTTCCTCAAGC-3′; β-ACTIN-F, 5′-ATGTGCAAGGCCGGTTTCGC-3′ and β-ACTIN-R, 5′-CGACACGCAGCTCATTGTAG-3′.
Immunoblotting
Protein samples were harvested from cells using the SDS sample buffer (100 mM Tris-HCl, pH 6.8, 2% SDS, 40% glycerol, 5% β-mercaptoethanol, and 0.1% bromophenol blue). Samples were heated at 99°C for 10 min prior to being subjected to the immunoblot analysis. The protein samples were then transferred to a PVDF membrane (IPVH00010, pore size 0.45 μm, Merck Millipore). The membrane was blocked using 5% nonfat milk at 25°C for 1 h, followed by the incubation of primary antibodies at 4°C for 16 h. Primary antibodies used were anti-TCF4 (ab217668, 1:1,000, abcam), anti-INTU (ab229243, 1:1,000, abcam), anti-IFT88 (13967-1-AP, 1:1,000, Proteintech) and anti-β-TUBULIN (ab6046, 1:2,000, abcam). The membrane was washed three times with 1× TBST each for 10 min, before being subjected to the incubation of secondary antibodies at 25°C for 1 h. Secondary antibodies used were HRP-conjugated goat anti-rabbit IgG (H + L) (11-035-045, 1:5,000) and HRP-conjugated goat anti-mouse IgG (H + L) (115-035-062, 1:10,000) from Jackson ImmunoResearch. The membrane was washed three times with 1× TBST each for 10 min, prior to the detection of chemiluminescent signal. The signal was developed using Immobilon Forte Western HRP substrate (WBLUF0100, Merck Millipore), and the images were captured and processed using ChemiDoc™ Touch Imaging System (170-8370, Bio-Rad).
Statistical analysis
The two-tailed, unpaired Student’s t test was used for the comparison between two experimental groups. *, ** and *** represent p < 0.05, p < 0.01 and p < 0.001, respectively, which are considered statistically significant. ns indicates no significant difference. GraphPad Prism version 9.0.0 was used for statistical analysis.
Author Contributions
Z.S.C. designed research, performed experiments and analyzed the data, H.Y.E.C. and Z.S.C. wrote and revised the paper.
Acknowledgments
The human endometrial adenocarcinoma cell line AN3 CA was a kind gift from Prof. Chi Chiu Wang (Department of Obstetrics and Gynecology, The Chinese University of Hong Kong, China). The pcDNA-MALAT1 plasmid was a kind gift from Prof. Huating Wang (Department of Orthopaedics and Traumatology, The Chinese University of Hong Kong, China).
Conflicts of Interest
The authors declare no conflicts of interest related to this study.
Ethical Statement
Biological and chemical safety approval for this study (14122815) was obtained from the Chinese University of Hong Kong.
Funding
Zhefan Stephen Chen was supported by a Postdoctoral Fellowship in Clinical Neurosciences program between The Chinese University of Hong Kong and University of Oxford (Nuffield Department of Clinical Neurosciences and Pembroke College).
References
- 1. Wu F, Zhang Y, Sun B, McMahon AP, Wang Y. Hedgehog Signaling: From Basic Biology to Cancer Therapy. Cell Chem Biol. 2017; 24:252–80. https://doi.org/10.1016/j.chembiol.2017.02.010 [PubMed]
- 2. Niyaz M, Khan MS, Mudassar S. Hedgehog Signaling: An Achilles' Heel in Cancer. Transl Oncol. 2019; 12:1334–44. https://doi.org/10.1016/j.tranon.2019.07.004 [PubMed]
- 3. Goetz SC, Anderson KV. The primary cilium: a signalling centre during vertebrate development. Nat Rev Genet. 2010; 11:331–44. https://doi.org/10.1038/nrg2774 [PubMed]
- 4. Cui C, Chatterjee B, Lozito TP, Zhang Z, Francis RJ, Yagi H, Swanhart LM, Sanker S, Francis D, Yu Q, San Agustin JT, Puligilla C, Chatterjee T, et al. Wdpcp, a PCP protein required for ciliogenesis, regulates directional cell migration and cell polarity by direct modulation of the actin cytoskeleton. PLoS Biol. 2013; 11:e1001720. https://doi.org/10.1371/journal.pbio.1001720 [PubMed]
- 5. Park TJ, Haigo SL, Wallingford JB. Ciliogenesis defects in embryos lacking inturned or fuzzy function are associated with failure of planar cell polarity and Hedgehog signaling. Nat Genet. 2006; 38:303–11. https://doi.org/10.1038/ng1753 [PubMed]
- 6. Wheway G, Nazlamova L, Hancock JT. Signaling through the Primary Cilium. Front Cell Dev Biol. 2018; 6:8. https://doi.org/10.3389/fcell.2018.00008 [PubMed]
- 7. Taschner M, Lorentzen E. The Intraflagellar Transport Machinery. Cold Spring Harb Perspect Biol. 2016; 8:a028092. https://doi.org/10.1101/cshperspect.a028092 [PubMed]
- 8. Toriyama M, Lee C, Taylor SP, Duran I, Cohn DH, Bruel AL, Tabler JM, Drew K, Kelly MR, Kim S, Park TJ, Braun DA, Pierquin G, et al, and University of Washington Center for Mendelian Genomics. The ciliopathy-associated CPLANE proteins direct basal body recruitment of intraflagellar transport machinery. Nat Genet. 2016; 48:648–56. https://doi.org/10.1038/ng.3558 [PubMed]
- 9. Skoda AM, Simovic D, Karin V, Kardum V, Vranic S, Serman L. The role of the Hedgehog signaling pathway in cancer: A comprehensive review. Bosn J Basic Med Sci. 2018; 18:8–20. https://doi.org/10.17305/bjbms.2018.2756 [PubMed]
- 10. Chen ZS, Lin X, Chan TF, Chan HYE. Pan-cancer investigation reveals mechanistic insights of planar cell polarity gene Fuz in carcinogenesis. Aging (Albany NY). 2021; 13:7259–83. https://doi.org/10.18632/aging.202582 [PubMed]
- 11. Natanzon Y, Earp M, Cunningham JM, Kalli KR, Wang C, Armasu SM, Larson MC, Bowtell DD, Garsed DW, Fridley BL, Winham SJ, Goode EL. Genomic Analysis Using Regularized Regression in High-Grade Serous Ovarian Cancer. Cancer Inform. 2018; 17:1176935118755341. https://doi.org/10.1177/1176935118755341 [PubMed]
- 12. Yang N, Leung EL, Liu C, Li L, Eguether T, Jun Yao XJ, Jones EC, Norris DA, Liu A, Clark RA, Roop DR, Pazour GJ, Shroyer KR, Chen J. INTU is essential for oncogenic Hh signaling through regulating primary cilia formation in basal cell carcinoma. Oncogene. 2017; 36:4997–5005. https://doi.org/10.1038/onc.2017.117 [PubMed]
- 13. Deaton AM, Bird A. CpG islands and the regulation of transcription. Genes Dev. 2011; 25:1010–22. https://doi.org/10.1101/gad.2037511 [PubMed]
- 14. Jones PA. Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat Rev Genet. 2012; 13:484–92. https://doi.org/10.1038/nrg3230 [PubMed]
- 15. Christman JK. 5-Azacytidine and 5-aza-2'-deoxycytidine as inhibitors of DNA methylation: mechanistic studies and their implications for cancer therapy. Oncogene. 2002; 21:5483–95. https://doi.org/10.1038/sj.onc.1205699 [PubMed]
- 16. Lambert SA, Jolma A, Campitelli LF, Das PK, Yin Y, Albu M, Chen X, Taipale J, Hughes TR, Weirauch MT. The Human Transcription Factors. Cell. 2018; 172:650–65. https://doi.org/10.1016/j.cell.2018.01.029 [PubMed]
- 17. O'Brien J, Hayder H, Zayed Y, Peng C. Overview of MicroRNA Biogenesis, Mechanisms of Actions, and Circulation. Front Endocrinol (Lausanne). 2018; 9:402. https://doi.org/10.3389/fendo.2018.00402 [PubMed]
- 18. Statello L, Guo CJ, Chen LL, Huarte M. Gene regulation by long non-coding RNAs and its biological functions. Nat Rev Mol Cell Biol. 2021; 22:96–118. https://doi.org/10.1038/s41580-020-00315-9 [PubMed]
- 19. Karagkouni D, Paraskevopoulou MD, Chatzopoulos S, Vlachos IS, Tastsoglou S, Kanellos I, Papadimitriou D, Kavakiotis I, Maniou S, Skoufos G, Vergoulis T, Dalamagas T, Hatzigeorgiou AG. DIANA-TarBase v8: a decade-long collection of experimentally supported miRNA-gene interactions. Nucleic Acids Res. 2018; 46:D239–45. https://doi.org/10.1093/nar/gkx1141 [PubMed]
- 20. Fasanaro P, Greco S, Lorenzi M, Pescatori M, Brioschi M, Kulshreshtha R, Banfi C, Stubbs A, Calin GA, Ivan M, Capogrossi MC, Martelli F. An integrated approach for experimental target identification of hypoxia-induced miR-210. J Biol Chem. 2009; 284:35134–43. https://doi.org/10.1074/jbc.M109.052779 [PubMed]
- 21. Hollander JA, Im HI, Amelio AL, Kocerha J, Bali P, Lu Q, Willoughby D, Wahlestedt C, Conkright MD, Kenny PJ. Striatal microRNA controls cocaine intake through CREB signalling. Nature. 2010; 466:197–202. https://doi.org/10.1038/nature09202 [PubMed]
- 22. Karginov FV, Hannon GJ. Remodeling of Ago2-mRNA interactions upon cellular stress reflects miRNA complementarity and correlates with altered translation rates. Genes Dev. 2013; 27:1624–32. https://doi.org/10.1101/gad.215939.113 [PubMed]
- 23. Haecker I, Gay LA, Yang Y, Hu J, Morse AM, McIntyre LM, Renne R. Ago HITS-CLIP expands understanding of Kaposi's sarcoma-associated herpesvirus miRNA function in primary effusion lymphomas. PLoS Pathog. 2012; 8:e1002884. https://doi.org/10.1371/journal.ppat.1002884 [PubMed]
- 24. Taylor MD, Liu L, Raffel C, Hui CC, Mainprize TG, Zhang X, Agatep R, Chiappa S, Gao L, Lowrance A, Hao A, Goldstein AM, Stavrou T, et al. Mutations in SUFU predispose to medulloblastoma. Nat Genet. 2002; 31:306–10. https://doi.org/10.1038/ng916 [PubMed]
- 25. Xie J, Murone M, Luoh SM, Ryan A, Gu Q, Zhang C, Bonifas JM, Lam CW, Hynes M, Goddard A, Rosenthal A, Epstein EH
Jr , de Sauvage FJ. Activating Smoothened mutations in sporadic basal-cell carcinoma. Nature. 1998; 391:90–2. https://doi.org/10.1038/34201 [PubMed] - 26. Tostar U, Malm CJ, Meis-Kindblom JM, Kindblom LG, Toftgård R, Undén AB. Deregulation of the hedgehog signalling pathway: a possible role for the PTCH and SUFU genes in human rhabdomyoma and rhabdomyosarcoma development. J Pathol. 2006; 208:17–25. https://doi.org/10.1002/path.1882 [PubMed]
- 27. Martin ST, Sato N, Dhara S, Chang R, Hustinx SR, Abe T, Maitra A, Goggins M. Aberrant methylation of the Human Hedgehog interacting protein (HHIP) gene in pancreatic neoplasms. Cancer Biol Ther. 2005; 4:728–33. https://doi.org/10.4161/cbt.4.7.1802 [PubMed]
- 28. Cancer Genome Atlas Network. Comprehensive molecular characterization of human colon and rectal cancer. Nature. 2012; 487:330–7. https://doi.org/10.1038/nature11252 [PubMed]
- 29. Cohen M, Kicheva A, Ribeiro A, Blassberg R, Page KM, Barnes CP, Briscoe J. Ptch1 and Gli regulate Shh signalling dynamics via multiple mechanisms. Nat Commun. 2015; 6:6709. https://doi.org/10.1038/ncomms7709 [PubMed]
- 30. Shyer AE, Huycke TR, Lee C, Mahadevan L, Tabin CJ. Bending gradients: how the intestinal stem cell gets its home. Cell. 2015; 161:569–80. https://doi.org/10.1016/j.cell.2015.03.041 [PubMed]
- 31. Gerling M, Büller NV, Kirn LM, Joost S, Frings O, Englert B, Bergström Å, Kuiper RV, Blaas L, Wielenga MC, Almer S, Kühl AA, Fredlund E, et al. Stromal Hedgehog signalling is downregulated in colon cancer and its restoration restrains tumour growth. Nat Commun. 2016; 7:12321. https://doi.org/10.1038/ncomms12321 [PubMed]
- 32. Goetz SC, Bangs F, Barrington CL, Katsanis N, Anderson KV. The Meckel syndrome- associated protein MKS1 functionally interacts with components of the BBSome and IFT complexes to mediate ciliary trafficking and hedgehog signaling. PLoS One. 2017; 12:e0173399. https://doi.org/10.1371/journal.pone.0173399 [PubMed]
- 33. Liem KF
Jr , Ashe A, He M, Satir P, Moran J, Beier D, Wicking C, Anderson KV. The IFT-A complex regulates Shh signaling through cilia structure and membrane protein trafficking. J Cell Biol. 2012; 197:789–800. https://doi.org/10.1083/jcb.201110049 [PubMed] - 34. Eguether T, Cordelieres FP, Pazour GJ. Intraflagellar transport is deeply integrated in hedgehog signaling. Mol Biol Cell. 2018; 29:1178–89. https://doi.org/10.1091/mbc.E17-10-0600 [PubMed]
- 35. Auerbach O, Stout AP. Histopathological aspects of occult cancer of the lung. Ann N Y Acad Sci. 1964; 114:803–10. [PubMed]
- 36. Gould PR, Li L, Henderson DW, Barter RA, Papadimitriou JM. Cilia and ciliogenesis in endometrial adenocarcinomas. An ultrastructural analysis. Arch Pathol Lab Med. 1986; 110:326–30. [PubMed]
- 37. Bao Z, Huang WJ. Thioridazine promotes primary ciliogenesis in lung cancer cells through enhancing cell autophagy. Int J Clin Exp Med. 2017; 10:13960–9.
- 38. Jung M, Häberle BM, Tschaikowsky T, Wittmann MT, Balta EA, Stadler VC, Zweier C, Dörfler A, Gloeckner CJ, Lie DC. Analysis of the expression pattern of the schizophrenia-risk and intellectual disability gene TCF4 in the developing and adult brain suggests a role in development and plasticity of cortical and hippocampal neurons. Mol Autism. 2018; 9:20. https://doi.org/10.1186/s13229-018-0200-1 [PubMed]
- 39. Murre C. Helix-loop-helix proteins and the advent of cellular diversity: 30 years of discovery. Genes Dev. 2019; 33:6–25. https://doi.org/10.1101/gad.320663.118 [PubMed]
- 40. Kennedy AJ, Rahn EJ, Paulukaitis BS, Savell KE, Kordasiewicz HB, Wang J, Lewis JW, Posey J, Strange SK, Guzman-Karlsson MC, Phillips SE, Decker K, Motley ST, et al. Tcf4 Regulates Synaptic Plasticity, DNA Methylation, and Memory Function. Cell Rep. 2016; 16:2666–85. https://doi.org/10.1016/j.celrep.2016.08.004 [PubMed]
- 41. Kaaij LT, van de Wetering M, Fang F, Decato B, Molaro A, van de Werken HJ, van Es JH, Schuijers J, de Wit E, de Laat W, Hannon GJ, Clevers HC, Smith AD, Ketting RF. DNA methylation dynamics during intestinal stem cell differentiation reveals enhancers driving gene expression in the villus. Genome Biol. 2013; 14:R50. https://doi.org/10.1186/gb-2013-14-5-r50 [PubMed]
- 42. Baratz KH, Tosakulwong N, Ryu E, Brown WL, Branham K, Chen W, Tran KD, Schmid-Kubista KE, Heckenlively JR, Swaroop A, Abecasis G, Bailey KR, Edwards AO. E2-2 protein and Fuchs's corneal dystrophy. N Engl J Med. 2010; 363:1016–24. https://doi.org/10.1056/NEJMoa1007064 [PubMed]
- 43. Amiel J, Rio M, de Pontual L, Redon R, Malan V, Boddaert N, Plouin P, Carter NP, Lyonnet S, Munnich A, Colleaux L. Mutations in TCF4, encoding a class I basic helix-loop-helix transcription factor, are responsible for Pitt-Hopkins syndrome, a severe epileptic encephalopathy associated with autonomic dysfunction. Am J Hum Genet. 2007; 80:988–93. https://doi.org/10.1086/515582 [PubMed]
- 44. Wray NR, Ripke S, Mattheisen M, Trzaskowski M, Byrne EM, Abdellaoui A, Adams MJ, Agerbo E, Air TM, Andlauer TMF, Bacanu SA, Bækvad-Hansen M, Beekman AFT, et al, and eQTLGen, and 23andMe, and Major Depressive Disorder Working Group of the Psychiatric Genomics Consortium. Genome-wide association analyses identify 44 risk variants and refine the genetic architecture of major depression. Nat Genet. 2018; 50:668–81. https://doi.org/10.1038/s41588-018-0090-3 [PubMed]
- 45. Ellinghaus D, Folseraas T, Holm K, Ellinghaus E, Melum E, Balschun T, Laerdahl JK, Shiryaev A, Gotthardt DN, Weismüller TJ, Schramm C, Wittig M, Bergquist A, et al. Genome-wide association analysis in primary sclerosing cholangitis and ulcerative colitis identifies risk loci at GPR35 and TCF4. Hepatology. 2013; 58:1074–83. https://doi.org/10.1002/hep.25977 [PubMed]
- 46. Kool M, Jones DT, Jäger N, Northcott PA, Pugh TJ, Hovestadt V, Piro RM, Esparza LA, Markant SL, Remke M, Milde T, Bourdeaut F, Ryzhova M, et al, and ICGC PedBrain Tumor Project. Genome sequencing of SHH medulloblastoma predicts genotype-related response to smoothened inhibition. Cancer Cell. 2014; 25:393–405. https://doi.org/10.1016/j.ccr.2014.02.004 [PubMed]
- 47. Hellwig M, Lauffer MC, Bockmayr M, Spohn M, Merk DJ, Harrison L, Ahlfeld J, Kitowski A, Neumann JE, Ohli J, Holdhof D, Niesen J, Schoof M, et al. TCF4 (E2-2) harbors tumor suppressive functions in SHH medulloblastoma. Acta Neuropathol. 2019; 137:657–73. https://doi.org/10.1007/s00401-019-01982-5 [PubMed]
- 48. Gebert LFR, MacRae IJ. Regulation of microRNA function in animals. Nat Rev Mol Cell Biol. 2019; 20:21–37. https://doi.org/10.1038/s41580-018-0045-7 [PubMed]
- 49. Graham RP, Nair AA, Davila JI, Jin L, Jen J, Sukov WR, Wu TT, Appelman HD, Torres-Mora J, Perry KD, Zhang L, Kloft-Nelson SM, Knudson RA, et al. Gastroblastoma harbors a recurrent somatic MALAT1-GLI1 fusion gene. Mod Pathol. 2017; 30:1443–52. https://doi.org/10.1038/modpathol.2017.68 [PubMed]
- 50. Spans L, Fletcher CD, Antonescu CR, Rouquette A, Coindre JM, Sciot R, Debiec-Rychter M. Recurrent MALAT1-GLI1 oncogenic fusion and GLI1 up-regulation define a subset of plexiform fibromyxoma. J Pathol. 2016; 239:335–43. https://doi.org/10.1002/path.4730 [PubMed]
- 51. Antonescu CR, Agaram NP, Sung YS, Zhang L, Swanson D, Dickson BC. A Distinct Malignant Epithelioid Neoplasm With GLI1 Gene Rearrangements, Frequent S100 Protein Expression, and Metastatic Potential: Expanding the Spectrum of Pathologic Entities With ACTB/MALAT1/PTCH1-GLI1 Fusions. Am J Surg Pathol. 2018; 42:553–60. https://doi.org/10.1097/PAS.0000000000001010 [PubMed]
- 52. Prall OWJ, McEvoy CRE, Byrne DJ, Iravani A, Browning J, Choong DY, Yellapu B, O'Haire S, Smith K, Luen SJ, Mitchell PLR, Desai J, Fox SB, et al. A Malignant Neoplasm From the Jejunum With a MALAT1-GLI1 Fusion and 26-Year Survival History. Int J Surg Pathol. 2020; 28:553–62. https://doi.org/10.1177/1066896919900548 [PubMed]
- 53. Engreitz JM, Sirokman K, McDonel P, Shishkin AA, Surka C, Russell P, Grossman SR, Chow AY, Guttman M, Lander ES. RNA-RNA interactions enable specific targeting of noncoding RNAs to nascent Pre-mRNAs and chromatin sites. Cell. 2014; 159:188–99. https://doi.org/10.1016/j.cell.2014.08.018 [PubMed]
- 54. West JA, Davis CP, Sunwoo H, Simon MD, Sadreyev RI, Wang PI, Tolstorukov MY, Kingston RE. The long noncoding RNAs NEAT1 and MALAT1 bind active chromatin sites. Mol Cell. 2014; 55:791–802. https://doi.org/10.1016/j.molcel.2014.07.012 [PubMed]
- 55. Brandl L, Horst D, de Toni E, Kirchner T, Herbst A, Kolligs FT. ITF-2B protein levels are correlated with favorable prognosis in patients with colorectal carcinomas. Am J Cancer Res. 2015; 5:2241–8. [PubMed]
- 56. Herbst A, Bommer GT, Kriegl L, Jung A, Behrens A, Csanadi E, Gerhard M, Bolz C, Riesenberg R, Zimmermann W, Dietmaier W, Wolf I, Brabletz T, et al. ITF-2 is disrupted via allelic loss of chromosome 18q21, and ITF-2B expression is lost at the adenoma-carcinoma transition. Gastroenterology. 2009; 137:639–48. https://doi.org/10.1053/j.gastro.2009.04.049 [PubMed]
- 57. Grill JI, Herbst A, Brandl L, Kong L, Schneider MR, Kirchner T, Wolf E, Kolligs FT. Inactivation of Itf2 promotes intestinal tumorigenesis in Apc(Min/+) mice. Biochem Biophys Res Commun. 2015; 461:249–53. https://doi.org/10.1016/j.bbrc.2015.04.009 [PubMed]
- 58. Malakar P, Shilo A, Mogilevsky A, Stein I, Pikarsky E, Nevo Y, Benyamini H, Elgavish S, Zong X, Prasanth KV, Karni R. Long Noncoding RNA MALAT1 Promotes Hepatocellular Carcinoma Development by SRSF1 Upregulation and mTOR Activation. Cancer Res. 2017; 77:1155–67. https://doi.org/10.1158/0008-5472.CAN-16-1508 [PubMed]
- 59. Ji Q, Zhang L, Liu X, Zhou L, Wang W, Han Z, Sui H, Tang Y, Wang Y, Liu N, Ren J, Hou F, Li Q. Long non-coding RNA MALAT1 promotes tumour growth and metastasis in colorectal cancer through binding to SFPQ and releasing oncogene PTBP2 from SFPQ/PTBP2 complex. Br J Cancer. 2014; 111:736–48. https://doi.org/10.1038/bjc.2014.383 [PubMed]
- 60. Kim J, Piao HL, Kim BJ, Yao F, Han Z, Wang Y, Xiao Z, Siverly AN, Lawhon SE, Ton BN, Lee H, Zhou Z, Gan B, et al. Long noncoding RNA MALAT1 suppresses breast cancer metastasis. Nat Genet. 2018; 50:1705–15. https://doi.org/10.1038/s41588-018-0252-3 [PubMed]
- 61. Han Y, Wu Z, Wu T, Huang Y, Cheng Z, Li X, Sun T, Xie X, Zhou Y, Du Z. Tumor-suppressive function of long noncoding RNA MALAT1 in glioma cells by downregulation of MMP2 and inactivation of ERK/MAPK signaling. Cell Death Dis. 2016; 7:e2123. https://doi.org/10.1038/cddis.2015.407 [PubMed]
- 62. Conlon K, Watson DC, Waldmann TA, Valentin A, Bergamaschi C, Felber BK, Peer CJ, Figg WD, Potter EL, Roederer M, McNeel DG, Thompson JA, Gupta S, et al. Phase I study of single agent NIZ985, a recombinant heterodimeric IL-15 agonist, in adult patients with metastatic or unresectable solid tumors. J Immunother Cancer. 2021; 9:e003388. https://doi.org/10.1136/jitc-2021-003388 [PubMed]
- 63. Gilad Y, Gellerman G, Lonard DM, O'Malley BW. Drug Combination in Cancer Treatment-From Cocktails to Conjugated Combinations. Cancers (Basel). 2021; 13:669. https://doi.org/10.3390/cancers13040669 [PubMed]
- 64. Bhatia K, Bhumika, Das A. Combinatorial drug therapy in cancer - New insights. Life Sci. 2020; 258:118134. https://doi.org/10.1016/j.lfs.2020.118134 [PubMed]
- 65. Nagy Á, Munkácsy G, Győrffy B. Pancancer survival analysis of cancer hallmark genes. Sci Rep. 2021; 11:6047. https://doi.org/10.1038/s41598-021-84787-5 [PubMed]
- 66. Tang Z, Kang B, Li C, Chen T, Zhang Z. GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res. 2019; 47:W556–60. https://doi.org/10.1093/nar/gkz430 [PubMed]
- 67. Huang da W, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009; 4:44–57. https://doi.org/10.1038/nprot.2008.211 [PubMed]
- 68. Szklarczyk D, Gable AL, Lyon D, Junge A, Wyder S, Huerta-Cepas J, Simonovic M, Doncheva NT, Morris JH, Bork P, Jensen LJ, Mering CV. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019; 47:D607–13. https://doi.org/10.1093/nar/gky1131 [PubMed]
- 69. Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003; 13:2498–504. https://doi.org/10.1101/gr.1239303 [PubMed]
- 70. Li LC, Dahiya R. MethPrimer: designing primers for methylation PCRs. Bioinformatics. 2002; 18:1427–31. https://doi.org/10.1093/bioinformatics/18.11.1427 [PubMed]
- 71. Hu H, Miao YR, Jia LH, Yu QY, Zhang Q, Guo AY. AnimalTFDB 3.0: a comprehensive resource for annotation and prediction of animal transcription factors. Nucleic Acids Res. 2019; 47:D33–8. https://doi.org/10.1093/nar/gky822 [PubMed]
- 72. Castro-Mondragon JA, Riudavets-Puig R, Rauluseviciute I, Lemma RB, Turchi L, Blanc-Mathieu R, Lucas J, Boddie P, Khan A, Manosalva Pérez N, Fornes O, Leung TY, Aguirre A, et al. JASPAR 2022: the 9th release of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 2022; 50:D165–73. https://doi.org/10.1093/nar/gkab1113 [PubMed]
- 73. Huang HY, Li J, Tang Y, Huang YX, Chen YG, Xie YY, Zhou ZY, Chen XY, Ding SY, Luo MF, Jin CN, Zhao LS, Xu JT, et al. MethHC 2.0: information repository of DNA methylation and gene expression in human cancer. Nucleic Acids Res. 2021; 49:D1268–75. https://doi.org/10.1093/nar/gkaa1104 [PubMed]
- 74. Chen F, Chandrashekar DS, Varambally S, Creighton CJ. Pan-cancer molecular subtypes revealed by mass-spectrometry-based proteomic characterization of more than 500 human cancers. Nat Commun. 2019; 10:5679. https://doi.org/10.1038/s41467-019-13528-0 [PubMed]
- 75. Karagkouni D, Paraskevopoulou MD, Tastsoglou S, Skoufos G, Karavangeli A, Pierros V, Zacharopoulou E, Hatzigeorgiou AG. DIANA-LncBase v3: indexing experimentally supported miRNA targets on non-coding transcripts. Nucleic Acids Res. 2020; 48:D101–10. https://doi.org/10.1093/nar/gkz1036 [PubMed]
- 76. Li JH, Liu S, Zhou H, Qu LH, Yang JH. starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 2014; 42:D92–7. https://doi.org/10.1093/nar/gkt1248 [PubMed]
- 77. Li T, Fu J, Zeng Z, Cohen D, Li J, Chen Q, Li B, Liu XS. TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res. 2020; 48:W509–14. https://doi.org/10.1093/nar/gkaa407 [PubMed]
- 78. Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E, Antipin Y, Reva B, Goldberg AP, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012; 2:401–4. https://doi.org/10.1158/2159-8290.CD-12-0095 [PubMed]
- 79. Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, Cerami E, Sander C, Schultz N. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013; 6:pl1. https://doi.org/10.1126/scisignal.2004088 [PubMed]
- 80. Jiang W, Qu Y, Yang Q, Ma X, Meng Q, Xu J, Liu X, Wang S. D-lnc: a comprehensive database and analytical platform to dissect the modification of drugs on lncRNA expression. RNA Biol. 2019; 16:1586–91. https://doi.org/10.1080/15476286.2019.1649584 [PubMed]
- 81. Liu E, Liu Z, Zhou Y. Carboplatin-docetaxel-induced activity against ovarian cancer is dependent on up-regulated lncRNA PVT1. Int J Clin Exp Pathol. 2015; 8:3803–10. [PubMed]
- 82. Pan F, Zhu L, Lv H, Pei C. Quercetin promotes the apoptosis of fibroblast-like synoviocytes in rheumatoid arthritis by upregulating lncRNA MALAT1. Int J Mol Med. 2016; 38:1507–14. https://doi.org/10.3892/ijmm.2016.2755 [PubMed]