Introduction
Mutations of some genes cause severe defects in an organism’s health, while mutations of other genes cause subtle effects on health. Genes with modulatory effects may be involved in how well an organism can maintain its health and how long an organism can survive. One aspect of health that determines quality of life in the elderly is locomotor ability [1]. Age-related decline in locomotor ability is a predictor of loss of independence, depressive symptoms, morbidity, and mortality [2]. A better understanding of the factors that contribute to the maintenance of locomotor ability may enable novel approaches to prevent or delay age-related decline in locomotor ability and general health.
The nematode Caenorhabditis elegans provides a unique opportunity to study locomotor healthspan due to the availability of genetic techniques and its short lifespan of approximately two weeks [3]. From a candidate-based screen of genes that extend the swimming ability of aged C. elegans worms, the epidermal growth factor signalling pathway was found to extend locomotor healthspan and lifespan [4]. Recently, loss-of-function mutations in the calcium-activated potassium channel, SLO-1, were found to improve locomotor healthspan and lifespan in C. elegans [5].
In Sydney Brenner’s seminal forward genetic screen using C. elegans, he isolated uncoordinated (unc) mutants and identified genes that are required for the proper development of locomotor ability [3]. In order to identify novel genetic factors that play modulatory roles in the maintenance of locomotor healthspan, we previously carried out an unbiased forward genetic screen and isolated five C. elegans mutants that show progressive decline in motility [6].
In the present study, we analyzed one of the isolated strains from the screen, OF1262, to identify genes that modulate the maintenance of locomotor healthspan. From analyzing this strain, we found that a missense mutation in HDA-3 leads to reduced health, including progressive decline in maximum velocity, travel distance, and reduced longevity. HDA-3 is a C. elegans ortholog of human class I histone deacetylases (HDACs) [7]. C. elegans class I HDACs have been found to play roles in both development and aging-related pathologies. Loss of HDA-1 function leads to disorganized gonad structures and aberrant vulval development such as protruded vulva or the development of multiple vulvae [8, 9]. C. elegans class I HDACs have also been found to play a role in polyglutamine toxicity and longevity [10, 11]. The missense mutation identified in the OF1262 strain is a glycine to glutamic acid substitution at the 271st amino acid (G271E) in HDA-3. This mutation causes a gene cluster carrying BATH domains to be transcriptionally repressed. Proper regulation of gene expression by HDA-3 may be required for full maintenance of locomotor healthspan and longevity during adulthood.
Results
OF1262 strain that shows shortened adult locomotor healthspan carries hda-3(ix241) and dys-1(ix259) mutations
Previously, we carried out a forward genetic screen for C. elegans mutants with a shortened adult locomotor healthspan [6]. One of the isolated strains was the OF1262 strain which showed a slight developmental deficit and a marked progressive decline in the ability to reach a food source located on the outer edge of an agar plate (Figure 1A, 1B) [6]. The OF1262 strain shows a progressive decline in maximum velocity and travel distance during adulthood as measured by 1.0-minute video recordings of worms moving on an agar plate without food. OF1262 worms show a progressive decline in maximum velocity and travel distance even when measured on a plate without a food cue, suggesting the decline is likely not due to defects in the sensory system. Maximum velocity is the highest velocity that the worm reaches during the 1.0-minute video recording, while travel distance is the total distance that the worm covers. Together, these measures estimate the maximum neuromuscular capacity of worms, as well as the capacity to maintain a certain level of locomotion. In all of the strains we tested, we observed similar trends between maximum velocity and travel distance, and thus display only the maximum velocity measurements in the main figures and show the travel distance measurements in the supplementary figures. We observed variations in both measurements among separate experiments, suggesting sensitivity of the assay to slight differences in culture conditions such as temperature and humidity.
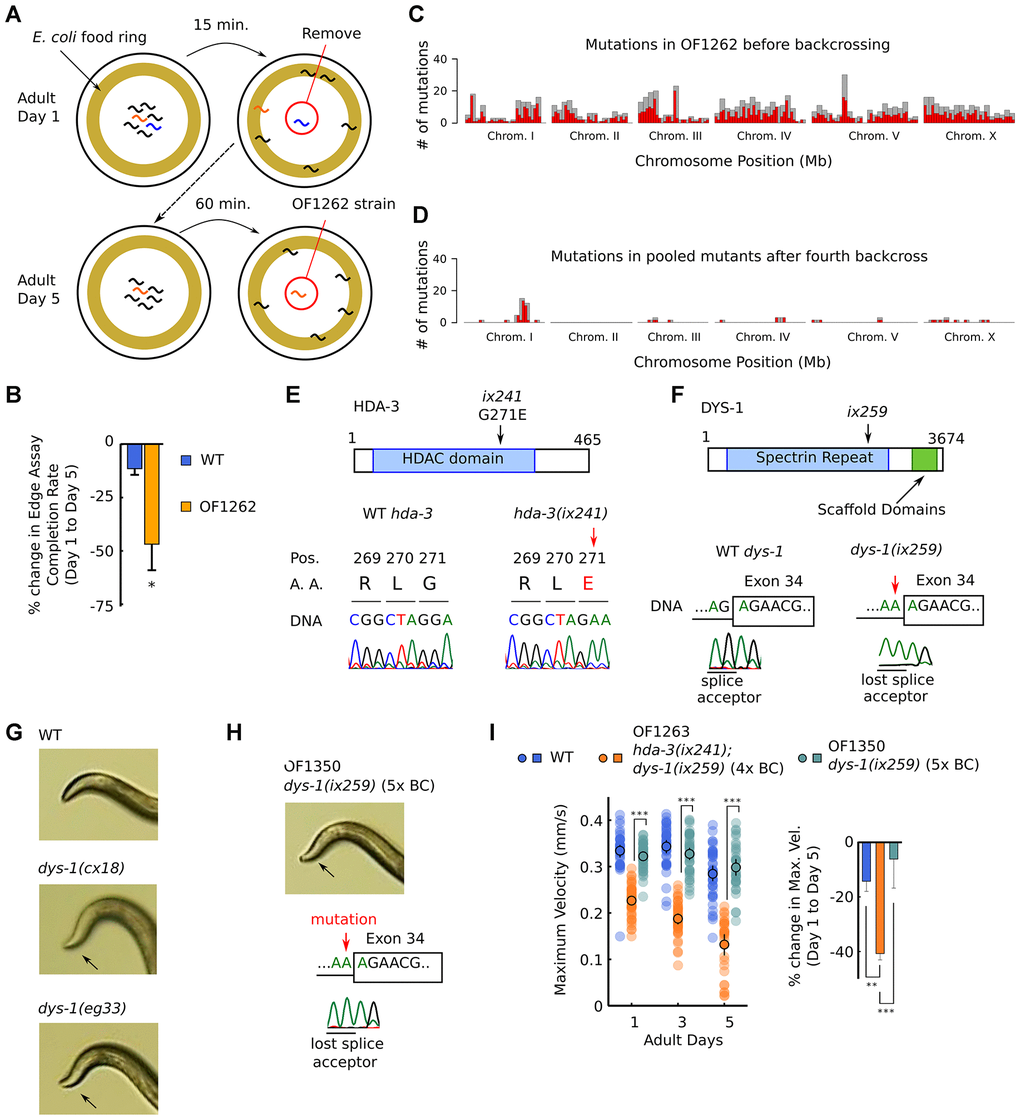
Figure 1. OF1262 strain shows locomotor deficit and carries hda-3(ix241) and dys-1(ix259) mutations. (A) Schematic of forward genetic screen to isolate OF1262 strain, a strain that shows progressive decline in adult motility. (B) Percent change in completion rate of the Edge Assay on adult day 5 compared to adult day 1. N = 3 biological replicate plates (100 worms or more per plate on adult day 1). (C) Mutation frequency along each chromosome for the OF1262 strain. Red bars indicate 0.5-Mb bins and grey bars indicate 1.0-Mb bins. (D) Mutation frequency along each chromosome for remaining mutations after subtracting mutations found in backcrossed strains that did not show a progressive decline in locomotor ability from mutations found in backcrossed strains that showed a progressive decline in locomotor ability. (E) Effect of hda-3(ix241) mutation on DNA sequence and amino acid sequence. (F) Effect of dys-1(ix259) mutation on DNA sequence. (G) Photos of head curvature during forward crawling in WT, dys-1(cx18) and dys-1(eg33) worms. Exaggerated head bending indicated by arrows. (H) (Top) Photo of head curvature during forward crawling in OF1350 dys-1(ix259) worms. (Bottom) DNA sequence of dys-1(ix259) mutation site. (I) (Left) Maximum velocities of WT, OF1263 hda-3(ix241);dys-1(ix259) (4x BC), and OF1350 dys-1(ix259) (5x BC) worms. N = 30–45 worms per strain for each day (10–15 worms from 3 biological replicate plates). (Right) Percent change in maximum velocity of WT, OF1263, and OF1350 worms on adult day 5 compared to adult day 1. N = 3 biological replicate plates. ***p < 0.001; **p < 0.01.
In order to identify the causative mutation site that leads to progressive decline in maximum velocity and travel distance in the OF1262 strain, we carried out whole genome sequencing in strains that showed and did not show the progressive decline after backcrossing. We identified mutations that were shared among the genomes of backcrossed strains that showed the progressive decline in maximum velocity, and subtracted the shared mutations among backcrossed strains that did not show the progressive decline in maximum velocity. A peak of mutations remained on Chromosome I (Figure 1C, 1D, Supplementary Table 1).
Two notable mutations remained after four backcrosses on Chromosome I: hda-3(ix241) and dys-1(ix259). hda-3(ix241) is a mutation in hda-3 that leads to a glycine to glutamic acid substitution at the 271st amino acid (G271E) (Figure 1E). dys-1(ix259) is a splice site mutation in dys-1 (Figure 1F). hda-3 and dys-1 are located in close proximity on Chromosome I. OF1263, a strain which was backcrossed four times to the wild-type (WT) strain, and other backcrossed strains retained an exaggerated head bending phenotype. Exaggerated head bending has previously been observed in mutants with loss-of-function mutations to dys-1, the C. elegans ortholog of human Dystrophin (Figure 1G), and to genes encoding components of the Dystrophin associated protein complex (DAPC) [12–17]. We postulated that the exaggerated head-bending and the progressive decline in maximum velocity are linked and caused by the same underlying mutation. However, after the fifth backcross, we isolated OF1350, a strain that carried the dys-1(ix259) mutation and showed exaggerated head-bending, but did not show the progressive decline in maximum velocity and travel distance (Figure 1H, 1I; Supplementary Figure 1A, 1B). This result suggested that the dys-1(ix259) mutation is not sufficient to cause the locomotor deficit. This came as a surprise since Dystrophin, the human ortholog of DYS-1 is implicated in progressive muscle weakness [17, 18].
hda-3 mutation causes progressive decline in maximum velocity and travel distance
The effect of the hda-3(ix241) mutation was tested by two strategies. In the first strategy, CRISPR-Cas9 genome editing was used to revert the hda-3(ix241) mutation in the OF1263 hda-3(ix241);dys-1(ix259) strain (Figure 2A). In order to prevent repetitive editing, a synonymous mutation was introduced that would disrupt the protospacer adjacent motif (PAM) sequence, 5 bp upstream of the editing site (Figure 2A). We refer to this reverted allele as hda-3(ix260), which has the same HDA-3 amino acid sequence as WT HDA-3. In the second strategy, the HDA-3 G271E mutation was introduced into the WT N2 background using CRISPR-Cas9 genome editing. Again, a synonymous mutation was introduced that would disrupt the PAM sequence (Figure 2A). We refer to this mutation allele as hda-3(ix261), which causes the same HDA-3 G271E mutation as the hda-3(ix241) mutation in the WT background. Strains carrying hda-3(ix261) were backcrossed twice.
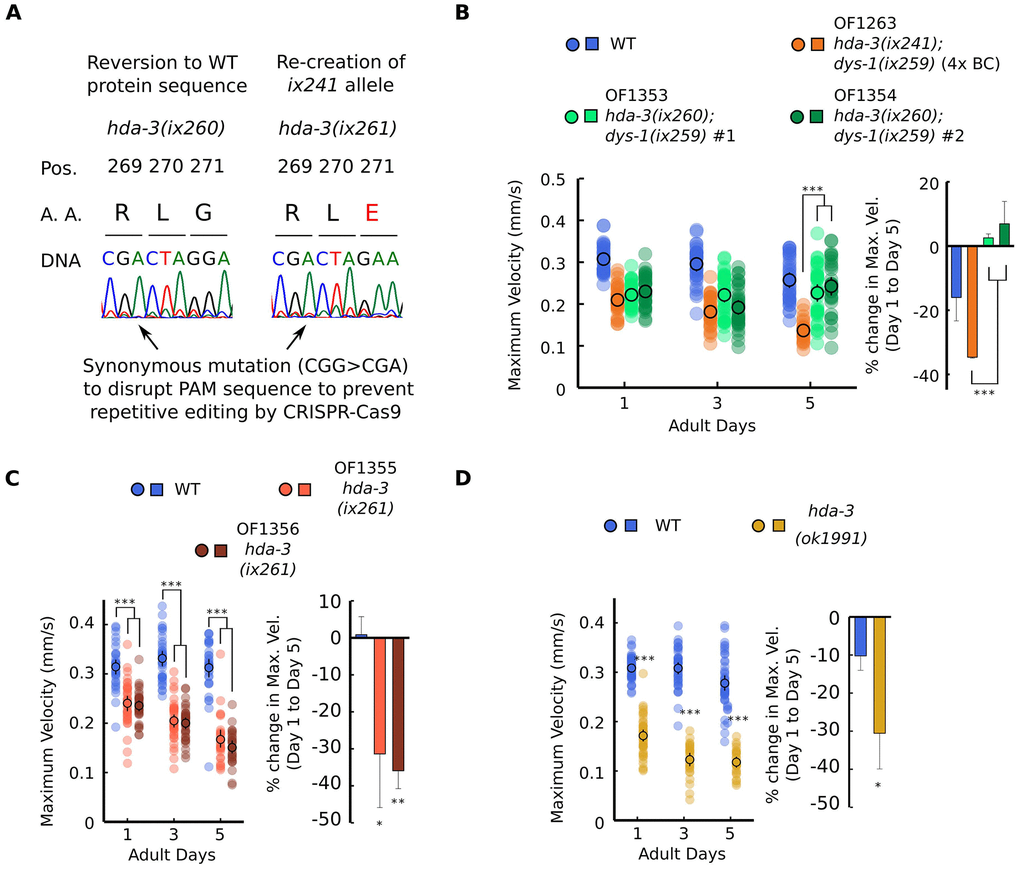
Figure 2. HDA-3 G271E missense mutation leads to progressive decline in locomotor ability. (A) (Left) Sequence of hda-3(ix260) allele which is the same amino acid sequence as WT. (Right) Sequence of hda-3(ix261) allele which is the same amino acid sequence as the hda-3(ix241) allele. Both sequences carry a synonymous mutation site to disrupt the PAM sequence to prevent repetitive editing by CRISPR-Cas9. (B) (Left) Maximum velocities of WT, OF1263 hda-3(ix241);dys-1(ix259) (4x BC), OF1353 hda-3(ix260);dys-1(ix259) and OF1354 hda-3(ix260);dys-1(ix259) worms. (Right) Percent change in maximum velocity of WT, OF1263, OF1353 and OF1354 worms on adult day 5 compared to adult day 1. (C) (Left) Maximum velocities of WT, OF1355 hda-3(ix261), OF1356 hda-3(ix261) worms. (Right) Percent change in maximum velocity of WT, OF1355 hda-3(ix261), OF1356 hda-3(ix261) worms on adult day 5 compared to adult day 1. (D) (Left) Maximum velocities of WT and hda-3(ok1991) worms. (Right) Percent change in maximum velocity of WT and hda-3(ok1991) worms on adult day 5 compared to adult day 1. For maximum velocity measurements, N = 30–45 worms per strain for each day (10–15 worms from 3 biological replicate plates). For percent change in maximum velocity graphs, N = 3 biological replicate plates. ***p < 0.001; **p < 0.01; *p < 0.05.
The reversion of the hda-3(ix241) mutation to hda-3(ix260) rescued the progressive decline in maximum velocity and travel distance (Figure 2B, Supplementary Figure 2A). This result indicated that the hda-3(ix241) mutation is necessary for the progressive decline in maximum velocity and travel distance. In the hda-3(ix261) strain that carries the G271E mutation in the N2 WT background, progressive decline in maximum velocity and travel distance was observed (Figure 2C, Supplementary Figure 2B). This result indicated that the HDA-3 G271E mutation alone is sufficient to cause progressive decline in maximum velocity and travel distance. In addition, an independently isolated hda-3(ok1991) deletion strain showed progressive decline in maximum velocity and travel distance (Figure 2D, Supplementary Figure 2C). The reversion of the hda-3(ix241) mutation did not however reverse the minor locomotor deficit observed on the first day of adulthood (Figure 2B). Given that OF1350 dys-1(ix259) did not show a locomotor deficit on the first day of adulthood (Figure 1I), there may be other mutations in the background of OF1353 and OF1354 that lead to minor deficits on the first day of adulthood. Another possibility is that the synonymous codon mutation we introduced in the PAM site (CGG>CGA) for CRISPR genome editing may have unanticipated effects on the locomotor ability of worms.
The G271 residue is evolutionarily conserved from C. elegans to humans and is located in the variable loop region (Figure 3A, 3B; Supplementary Figure 3A). The variable loop region is suggested to play a role in substrate recognition and binding to the HDAC cofactors zinc and inositol phosphate [19, 20] (Figure 3B). The C. elegans G271 amino acid aligns with human HDAC3 G267. The HDAC3 G267 amino acid is in close proximity to R265, which is an amino acid that mediates the interaction between HDAC3 and its coactivator, inositol phosphate [19, 21]. HDAC3 D259 is also in close proximity which is predicted to mediate the interaction between HDAC family proteins with the cofactor zinc [20]. The G271E mutation may alter activities of HDA-3 that play a role in the maintenance of locomotor ability.
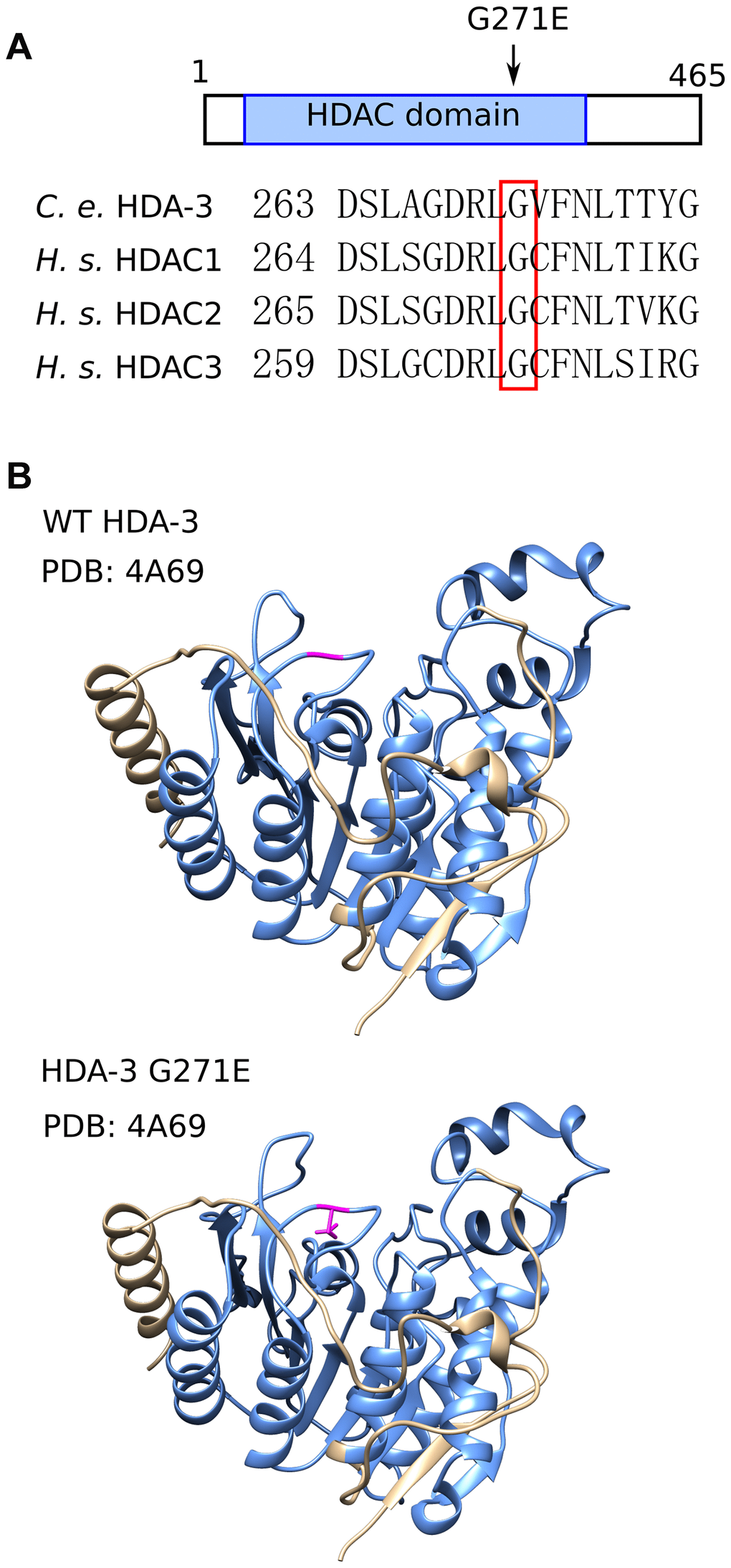
Figure 3. HDA-3 G271E missense mutation occurs at evolutionarily conserved residue. (A) (Top) Depiction of hda-3(ix241) mutation site in HDA-3 protein. (Bottom) Alignment of amino acid sequences centered around G271E mutation site in C. elegans HDA-3, H. sapiens HDAC1, HDAC2 and HDAC3. (B) (Top) Structural modeling of C. elegans WT HDA-3 based on PDB: 4A69 from H. Sapiens HDAC3. (Bottom) Structural modeling of C. elegans HDA-3 G271E based on PDB: 4A69 from H. Sapiens HDAC3. Mutated glutamic acid residue is shown in magenta. HDAC domain is indicated in blue.
Longevity is reduced in hda-3 mutants
Since HDA-3 has previously been reported to be expressed ubiquitously [10], we tested whether mutation in HDA-3 may have effects on general aspects of health and longevity. The median lifespans of hda-3(ix261) mutants were reduced from 16 days to 15 days compared to WT worms (Figure 4A). The hda-3(ix261) mutants shared a comparable reduction in lifespan as hda-3(ok1991) mutant worms that carry a deletion in hda-3. The body length of the strain OF1355 hda-3(ix261) was unchanged compared to that of WT worms, but another strain, OF1356 hda-3(ix261) (Figure 4B), was shorter and may carry additional mutations besides hda-3(ix261) that have minor effects on body size. There were no significant differences in the development times and brood sizes of the hda-3(ix261) mutant worms and WT (Figure 4C, 4D).
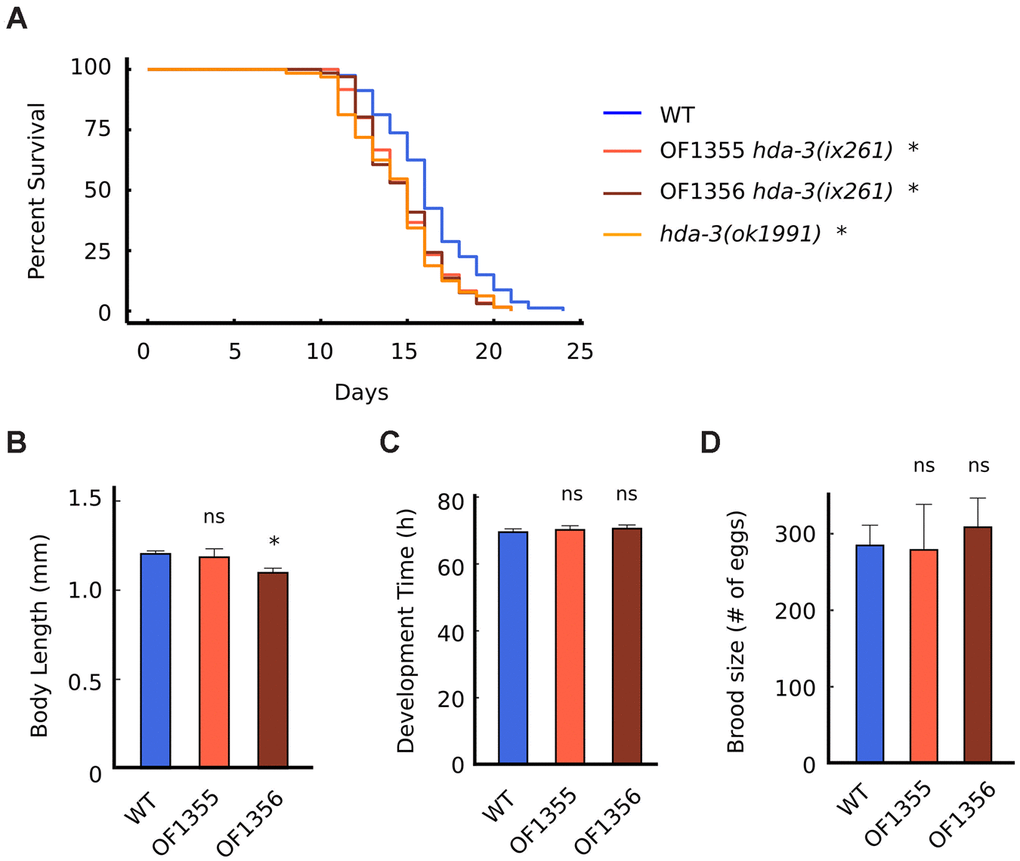
Figure 4. HDA-3 mutation leads to shortened lifespan. (A) Lifespan of WT (N = 80), OF1355 hda-3(ix261) (N=60), OF1356 hda-3(ix261) (N=66), and hda-3(ok1991) worms (N = 64). (B) Body length of WT, OF1355 hda-3(ix261), and OF1356 hda-3(ix261) worms. N = 10-17 worms per strain. (C) Development time of WT, OF1355 hda-3(ix261), and OF1356 hda-3(ix261) worms. N = 8-10 worms per strain. (D) Brood size of WT, OF1355 hda-3(ix261), and OF1356 hda-3(ix261) worms N = 5-6 worms per strain. *p < 0.05.
OF1355 and OF1356 worms showed a 25 % reduction in survival 1 day earlier than WT worms (Figure 4A), while they showed a 25 % decline in maximum velocity 3 days earlier than WT worms (Figure 2C). This suggests that the negative effects of the hda-3(ix261) mutation affect maximum velocity more strongly than lifespan.
Expression of subset of BATH genes are dysregulated in hda-3 mutant strains
In order to identify gene expression changes that occur due to the hda-3(ix241) mutation, transcriptome analysis was carried out in adult worms on the third day of adulthood. On the third day of adulthood, WT worms did not show decline in maximum velocity and travel distance whereas hda-3(ix241) worms showed significant declines compared to the first day of adulthood (Figure 1I). Therefore, we reasoned that genes that play a role in the maintenance of adult maximum velocity and travel distance may be differentially expressed between WT and hda-3(ix241) worms on the third day of adulthood.
In comparison to WT worms, OF1263 hda-3(ix241);dys-1(ix259) worms had 64 transcripts that were significantly upregulated and 47 transcripts that were significantly downregulated (FDR-adjusted q value < 0.05) (Figure 5A). In comparison to OF1350 dys-1(ix259) worms, which lost the hda-3(ix241) mutation after the fifth backcross, OF1263 worms had 27 transcripts that were significantly upregulated and 25 transcripts that were significantly downregulated (Figure 5A). Twenty-two transcripts were commonly upregulated in the OF1263 worms compared to WT and OF1350. Thirteen transcripts were commonly downregulated in the OF1263 worms compared to WT and OF1350 (Figure 5B).
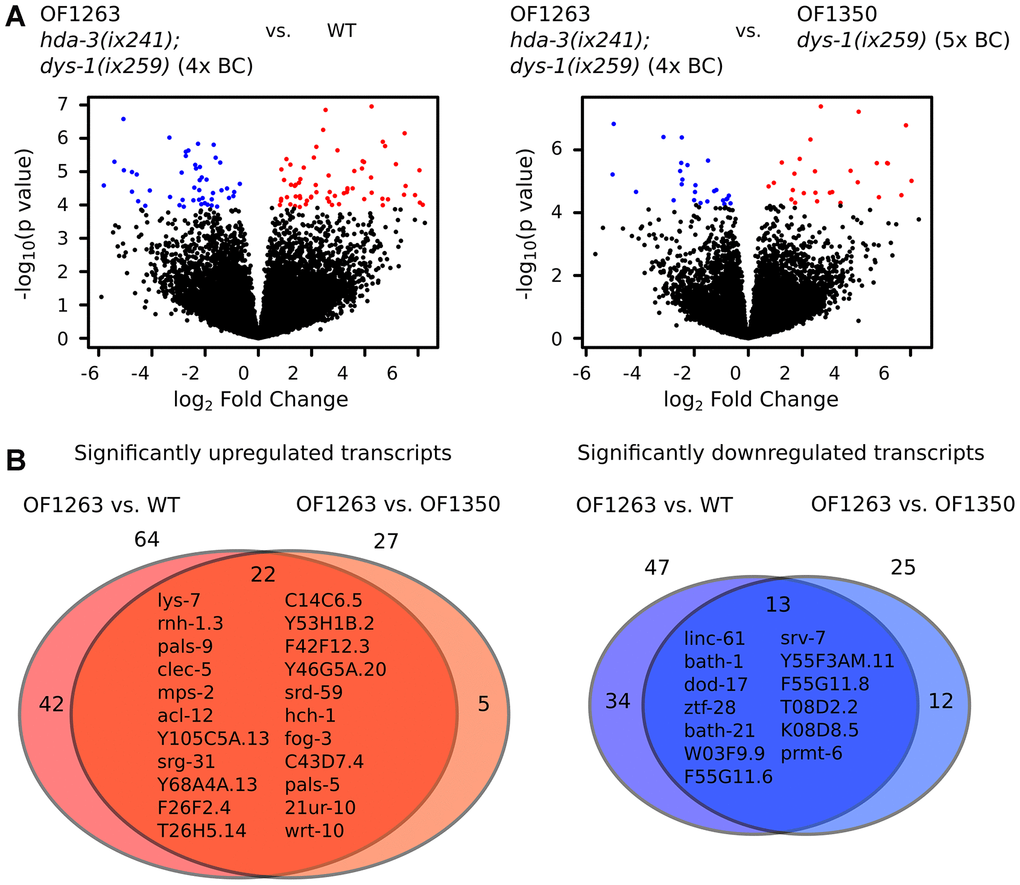
Figure 5. Gene expression is dysregulated in hda-3 mutant. (A) (Left) Volcano plot of differential expression of transcripts from OF1263 hda-3(ix241);dys-1(ix259) (4x BC) vs. WT worms. (Right) Volcano plot of differential expression of transcripts from OF1263 hda-3(ix241);dys-1(ix259) (4x BC) vs. OF1350 dys-1(ix259) (5x BC) worms. Blue points indicate downregulated genes and red points indicate upregulated genes with q value < 0.05 (FDR-adjusted p value). (B) (Left) Venn diagram of number of significantly upregulated transcripts with q < 0.05 in OF1263 vs. WT worms and OF1263 vs. OF1350 worms. (Right) Venn diagram of number of significantly upregulated and downregulated transcripts with P < 0.0001 in OF1263 vs. WT and OF1263 vs. OF1350 worms. Names of commonly upregulated or downregulated genes are indicated within the Venn diagram.
Among the downregulated transcripts, we noticed that multiple gene transcripts were downregulated within two narrow regions of the genome on Chromosome II and Chromosome IV (Figure 6A; Supplementary Figure 4A). On Chromosome II, the downregulated genes are bath-1, bath-21, and bath-24 which carry BATH domains (Figure 6A, 6B) [22, 23]. BATH domains are characterized by BTB/POZ (broad-complex, Tramtrack and bric-a-bric/Pox virus and zinc finger) and MATH (meprin-associated Traf homology) domains [23]. Downregulation of bath-1, bath-21 and bath-24 were observed in both hda-3(ix261) mutant worms and hda-3(ok1991) mutant worms (Figure 6C, 6D). HDA-3 activity is likely to play a role in promoting expression of the chromosome region which encompasses bath-1, bath-21, and bath-24. In addition, a less stringent threshold for downregulation of genes (unadjusted p value < 0.05) indicates that bath-3, bath-5, and btb-4 are also downregulated in the OF1263 strain (Supplementary Table 2). btb-4 carries the BTB-domain, one of the two domains that characterize BATH genes. INTERPRO protein domain enrichment analysis of the differentially expressed genes (p < 0.05) indicates that BTB and MATH domains are most significantly enriched among downregulated genes (Supplementary Table 3). Interestingly, histone genes (his-73, his-47, his-61, his-25, his-63, his-60) that are constituents of the nucleosome are also a category of genes that are overrepresented among downregulated genes, but not in the upregulated genes (Supplementary Table 3). Precise regulation of the composition of histone variants may be carried out by HDA-3.
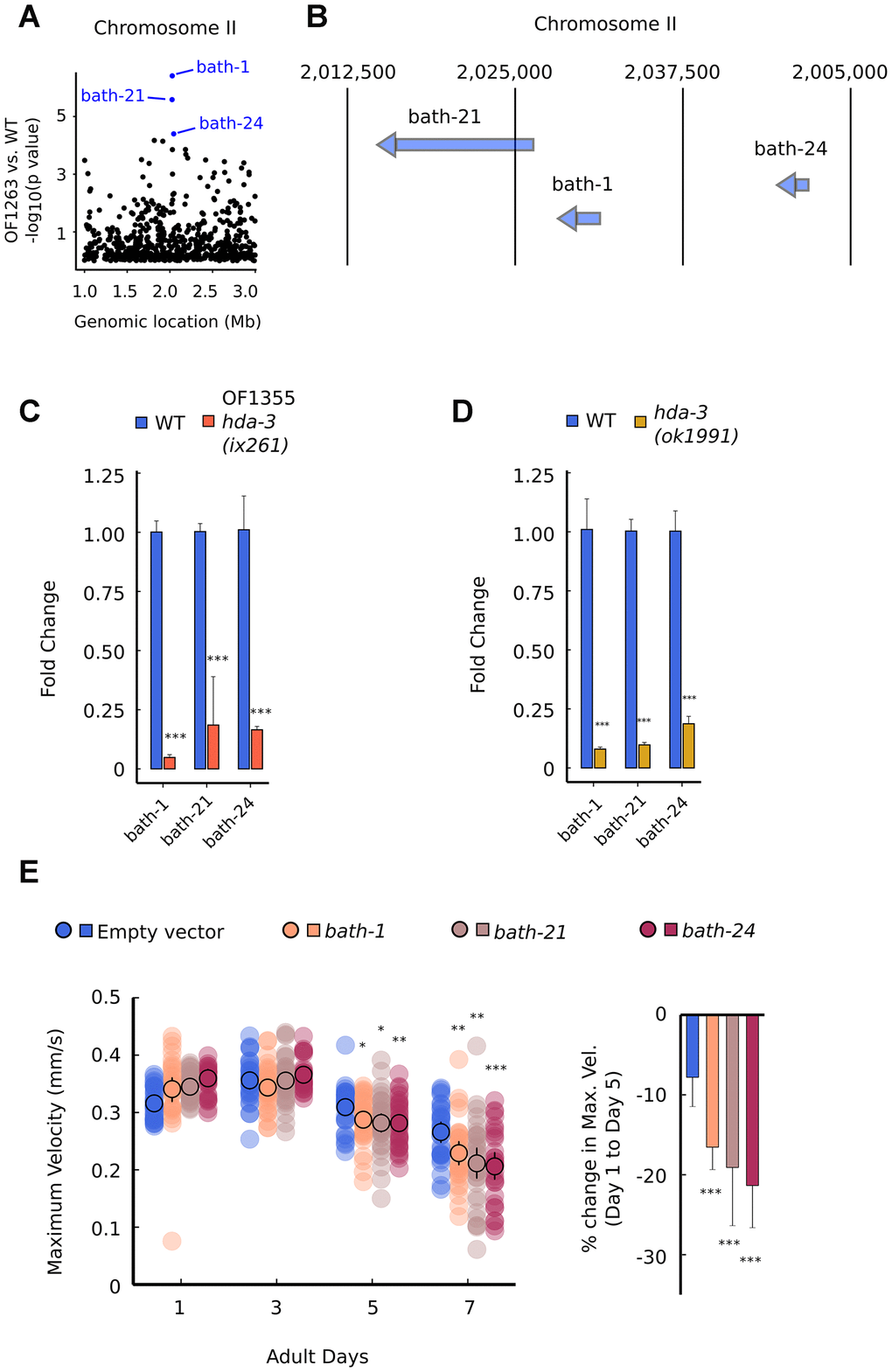
Figure 6. Specific BATH genes on Chromosome II are downregulated from HDA-3 mutations. (A) Genomic location of strongly downregulated gene transcripts on Chromosome II in OF1263 vs. WT. (B) Locations of BATH genes on Chromosome II. (C) Fold change in BATH genes in OF1355 hda-3(ix261) worms compared to WT, as measured by qPCR. (D) Fold change in BATH genes in hda-3(ok1991) worms compared to WT, as measured by qPCR. (E) Maximum velocities of worms fed with empty vector RNAi or RNAi targeted towards bath-1, bath-21, or bath-24. N = 30–45 worms per strain for each day (10–15 worms from 3 biological replicate plates). ***p < 0.001; **p < 0.01; *p < 0.05.
In order to test whether bath genes alone could have effects on motility, we knocked down bath-1, bath-21, and bath-24 separately using feeding RNAi, and measured maximum velocity and travel distance. bath-1, bath-21, and bath-24 individually had subtle but significant effects on maximum velocity and travel distance starting at the fifth day of adulthood (Figure 6E; Supplementary Figure 5A). These results suggest that bath genes may normally act to promote locomotor healthspan during later stages of reproductive adulthood.
Discussion
The role of class I HDACs during the aging process has been difficult to study, as HDAC1, HDAC2, HDAC3, and HDAC8 have been found to play important roles during development in vertebrates [24]. For example, HDAC3 knockout mice die during embryonic development [25]. HDACs have been linked to playing a role during the aging process as HDAC inhibitors have been shown to promote longevity in C. elegans and D. melanogaster [26–28]. HDAC inhibitors are also proposed to be therapeutic targets for various age-related pathologies such as sarcopenia and neurodegeneration [29–31].
Histone deacetylases can affect the transcriptional expression of many downstream genes [32]. Generally, histone acetylation is positively associated with transcriptional activation and deacetylation is associated with transcriptional repression [33]. However, other studies have found that HDACs are involved in both repression and activation [34, 35]. Our results show that hda-3(ix241) and hda-3(ok1991) mutations can lead to reduced expression of a subset of genes, and suggest that HDA-3 normally acts to promote expression of some genes. It is also possible that histone deacetylases may affect transcriptional outputs by mechanisms other than histone acetylation. HDAC3 can directly interact with transcription factors GATA-2 and Runx2 to repress their transcriptional activity [36, 37]. Histone deacetylases may affect transcriptional outputs by direct interactions or modifications to non-histone proteins.
Transcriptome analysis and quantitative PCR analyses of mutant strains carrying the HDA-3 G271E mutation indicated a region in the genome that is transcriptionally repressed on Chromosome II. The Chromosome II region carried genes of related function, bath-1, bath-3, bath-5, bath-21, bath-24, and btb-4 that contain BTB/POZ and/or MATH domains [23]. Genes that contain both BTB and MATH domains are referred to as bath genes [23]. bath genes are reported to be involved in proteasomal degradation by association with CULLIN3 (CUL3), an E3 ubiquitin ligase [38, 39]. The BTB domain is reported to bind CUL3, while the MATH domain binds specific substrates to be degraded [38, 40]. Based on our study, bath genes may be part of the subset of genes that work to maintain locomotor healthspan during later stages of reproductive adulthood.
BATH-42 has been implicated in the regulation of nicotinic acetylcholine receptors by interacting with the CUL-3 ubiquitin ligase complex in C. elegans [41]. Expression of genes carrying BATH domains may be important for regulating the expression levels of other proteins that play a role in maintaining adult locomotor ability. As part of the ubiquitin-proteasome system, CUL-3 and BATH-domain carrying proteins may promote proteostasis by targeting misfolded proteins for degradation. bath-24 was found to be enriched in unc-4::GFP expressing motor neurons, pointing towards a possible role in motor neuron functioning and locomotion [42]. bath-1 and bath-21 are reported to be enriched in germline, suggesting possible indirect roles in regulating healthspan [43]. Although the specific mechanism in which hda-3 and bath genes regulate locomotor healthspan is yet to be determined, they may do so by direct and indirect mechanisms originating from the neuromuscular system or other tissues.
The HDA-3(G271E) mutation occurs at an evolutionarily conserved residue. The same residue is present in human HDACs 1–3. The C. elegans G271 amino acid aligns with human HDAC3 G267. From exome and whole genome sequencing of the non-diseased human population, no missense mutations were reported in the HDAC3 G267 residue [44]. The G267 residue in human HDAC3 may be intolerant to amino acid changes, and thus play a critical role in proper HDAC3 function. The HDAC3 G267 amino acid is located in Variable Loop 4, one of the four variable loop regions which is implicated in substrate recognition [45]. The variable loops are different in length and sequence among the different HDAC proteins, enabling the recognition of specific substrates for each HDAC protein. Interestingly, all HDAC proteins, except for HDAC8, carry a glycine residue at the amino acid location aligned to HDAC3 G267. HDAC8 carries a cysteine residue instead of glycine. HDAC8 is believed to be evolutionarily divergent from the other class I histone deacetylases HDAC1-3 [46]. HDAC8 may have tissue-specific roles that are mediated by unique differences in amino acid sequence from the other HDAC proteins [47].
Since C. elegans can tolerate the loss of hda-3 during development, the HDA-3 G271E mutants and hda-3(ok1991) deletion mutant may be valuable tools to study the role of a specific class I histone deacetylase during aging. Unbiased genomic screens have found that knockdown of hda-3 leads to increased genome instability and plays a role in RNAi-mediated gene silencing [48]. The HDA-3 G271E mutants and hda-3(ok1991) deletion mutant have a reduced lifespan, and therefore may be used to identify genetic regulators of longevity downstream of HDA-3. Interestingly, Edwards et al. found that RNAi knockdown of hda-3 leads to a 16% increase in lifespan for C. elegans [11]. Although the culture method in which C. elegans lifespan was measured differed between our study (agar plates) and Edwards et al. (liquid culture), partial reductions in HDA-3 function may have positive downstream gene expression changes for lifespan, while loss-of-function mutations to HDA-3 may have negative gene expression changes.
Genetic screens in C. elegans have identified various genes involved in locomotion. Upregulation of the EGF signaling pathway or downregulation of SLO-1 function have been shown to improve locomotion during adulthood [4, 5]. In a previous study, we identified a nonsense mutation in elpc-2 that leads to progressive decline in maximum velocity and travel distance [6]. The role of elpc-2 as part of the Elongator complex implicates the role of tRNA modifications for the maintenance of proteostasis and adult locomotor ability. In this study, we identify the G271E mutation in HDA-3 as the causative mutation site that leads to progressive decline in maximum velocity and travel distance. HDA-3 alters the transcription of BATH-domain genes located in close proximity on Chromosome II. In concert with the E3 ubiquitin ligase CUL-3, BATH-domain proteins may also regulate specific expression of proteins involved in motility, or affect proteostasis in a broader context. Together, these mutants provide insights into the genetic mechanisms that contribute to the maintenance of adult locomotor ability and may aid our understanding of the genetic bases of healthy aging.
Materials and Methods
Strains
C. elegans Bristol N2 strain was used as the WT strain. Worms were cultivated at 20° C on Nematode Growth Media (NGM) agar plates with Escherichia coli strain OP50 as a food source [3]. All strains used in this study are listed in Supplementary Table 4.
Edge assay and mutant isolation
The Edge Assay was performed as previously described [6]. In brief, Edge Assay plates were prepared by spreading 100 μL E. coli OP50 suspension near the edge of a circular 9 cm plate. “Synchronized egg-laying” was used to raise a batch of worms of similar age. Synchronized worms were collected with M9 buffer, and placed on the center of the Edge Assay plate. The Edge Assay completion rate was measured by counting the number of worms that reached the E. coli lawn near the edge of the plate divided by the total number of worms on the plate. The forward genetic screen to isolate worms with progressive decline in locomotor ability was performed using the Edge Assay as previously described [6]. In brief, L4 worms were mutagenized in 50 mM EMS solution for 4 h. Mutagenized F2 worms were placed on the center of Edge Assay plates on adult day 1. Worms that did not reach the edge within 15 minutes were removed by aspiration. Worms that reached the edge were maintained on the Edge Assay plate and tested again on adult day 3 and 5. The OF1262 strain was isolated as a worm that did not reach the edge on adult day 5. Isolated mutants that did not reach the edge were singled out on individual plates and maintained. Most worms produced at least a few progeny on adult day 5, but worms that did not produce progeny were abandoned.
Sanger sequencing
Target genomic regions were amplified using PCR and purified using Wizard SV Gel and PCR Clean-Up System (Promega, Madison, WI). The DNA sequence of the PCR fragment was determined using cycle sequencing with BigDye v3.1 reagents (Applied Biosystems, Foster City, CA). Sequencing products were purified by EtOH/EDTA precipitation. 5 μL of 125 mM EDTA was added and mixed thoroughly to the 20 μL sequencing reaction. 60 μL of 100% ethanol was added and incubated for 15 min. The solution was centrifuged at 3000 x g for 30 min and the supernatant was discarded. 60 μL of 70% ethanol was added to the pellet and centrifuged at 1650 x g for 15 min. The supernatant was discarded, and the pellet was resuspended in 10 μL of Hi-Di Formamide (Applied Biosystems). Sequencing was performed by capillary sequencing using ABI3100 (Applied Biosystems). Primers used for Sanger sequencing are listed in Supplementary Table 5.
Whole-genome DNA sequencing
C. elegans DNA was extracted by ethanol precipitation. Mixed-stage worms were collected using M9 buffer, and washed 3 times with lysis buffer (200 mM NaCl, 100 mM Tris-Hcl (pH 8.5), 50 mM EDTA, 0.5% SDS). 10 μL of 10 mg/mL proteinase K was added to the 500 μL lysis buffer solution containing the worms. The solution was incubated at 55° C for at least 3 h. Proteinase K was deactivated by incubation at 95° C for 30 min. 20 μL of 20 mg/mL RNAase A was added and incubated at 37° C for 30 min. 125 μL of saturated NaCl was added, and the solution was centrifuged for 15 min at 15,000 rpm. 500 μL of the supernatant was transferred to a new tube containing 1 mL of 100% ethanol. The solution was centrifuged at 15,000 rpm for 20 min. The supernatant was discarded and 1 mL of 70% ethanol was added to the pellet. The solution was centrifuged at 15,000 rpm for 20 min. The supernatant was discarded and the pellet containing DNA was reconstituted in 100 μL TE buffer. DNA was sequenced using the MiSeq next-generation sequencing system (Illumina, San Diego, CA) as previously described [6]. Libraries for sequencing were prepared with Illumina TruSeq Library Prep Kit. Sequenced reads were mapped using BWA software [49]. Mapped read files were converted to bam format, then to pileup format with Samtools [50]. Variant detection was carried out using VarScan and SnpEff [51–55]. Mutation effects were classified into four categories, (high, moderate, low, modifier) by SnpEff. The categorization of specific types of mutations are listed in the figure legend of Supplemetary Table 1. Mutations categorized as a high impact mutation were prioritized as most promising candidates that lead to the progressive decline in locomotion. Mutation frequencies were calculated and visualized using CloudMap [56].
Measurements of maximum velocity and travel distance
Five adult day 1 worms were placed onto an NGM plate with food, and allowed to lay eggs for 3 h. When the offspring reached adult day 1, 15 worms were randomly picked onto a 6 cm NGM plate without bacteria. After the worms moved away from the initial location with residual food, worms were again moved onto a different NGM plate without bacteria. The maximum velocity and travel distance of worms were measured on the first, third, and fifth days of adulthood as previously described by video recording [6]. Videos were analyzed using ImageJ and wrMTrck software (www.phage.dk/plugins) to produce travel distance and maximum velocity values [57]. The same plate of worms measured on the first day of adulthood were measured on the third and fifth days of adulthood. For the percent change in maximum velocity or travel distance, the percentage decline from the first to fifth day of adulthood for each plate was calculated. Maximum velocity refers to the peak velocity that a worm reached during the 1.0 minute video recording. Travel distance refers to the total distance that the worm traveled during the 1.0-minute video recording. R was used to make plots [58].
CRISPR-Cas9 genome editing
Targeted mutagenesis was carried out using CRISPR-Cas9 genome editing with single-stranded oligodeoxynucleotide (ssODN) donors as previously described [59]. First, a ribonucleoprotein complex was created by mixing together 0.5 μL of 10 μg/μL Cas9 protein, 5.0 μL of 0.4 μg/μL of tracrRNA, and 2.8 μL of 0.4 μg/μL of crRNA (Target-specific sequence: 5’-CCGAUUCACUGGCAGGAGAU-3’) and incubating at 37° C for 10 min. Following incubation, 2.2 μL of 1.0 μg/μL ssODN, 2.0 μL of 400 ng/μL pRF4::rol-6(su1006) co-injection marker, and 7.5 μL of nuclease free water was added to the mixture. This mixture was then injected into the gonad of young adult-stage worms subject to genomic editing. F1 offspring that showed the roller phenotype were singled onto individual plates, and allowed to lay eggs. Editing of the target sequence was checked by single worm PCR of the F1 worm, followed by Sanger sequencing. ssODN sequences are listed in Supplementary Table 6.
Lifespan measurements
The lifespan of each worm strain was measured at 20° C. Worms that did not respond to prodding to the head and tail were considered dead. Worms that died from an exploded vulva, bag-of-worms phenotype, or that were lost during the incubation period were censored. Lifespan measurements were carried out in the absence of Floxuridine.
Body length measurement
Photos of age-synchronized worms were taken on the first day of adulthood. Worms were mounted on agar pads in M9 buffer containing 20 mM sodium azide. A central line through the entire body of the worm was traced and measured using ImageJ [60].
Brood size
Brood size was measured by allowing individual worms to lay eggs on a plate with food. The individual worm was moved to a fresh plate every 24 h until no more eggs were laid. The number of live larvae that were produced were counted on the next day. The total number of offspring that a single worm produced was counted as the total brood size.
Development time
The time from egg to first egg-lay was considered as the development time of the worm. Gravid adults were allowed to lay eggs within a 1-h time window. Individual eggs were picked and placed on individual plates with food. Beginning at 66 h of development, worms were checked each hour to see if they had laid any eggs. The first time point in which eggs were laid was marked as the time of first egg-lay.
RNA sequencing
Worms were synchronized by placing ten adult day 1 worms onto an NGM plate with food, and allowed to lay eggs for 3 h. At the L4 stage, worms were collected and washed with M9 buffer and placed on 9 cm NGM plates with 25 μM floxuridine (FUDR). On the third day of adulthood, worms were collected with M9 buffer and RNA was extracted. RNA was extracted by the phenol-chloroform method using Trizol reagent (Thermo Fisher Scientific). A 0.4 mL slurry of glass beads in water was added to 1 mL Trizol. Synchronized adult day 3 worms were collected with 500 μL M9 buffer, and the worms were allowed to settle to the tip of the pipette tip. A drop of M9 buffer containing the majority of worms at the tip of the pipette was dropped into the Trizol solution. Worms were homogenized by 4 sets of vortexing using Micro Smash MS-100R (Tomy Seiko, Tokyo, Japan) at 5,000 rpm for 2 min at 4° C. Additional vortexing cycles were carried out if worms were not completely homogenized. The solution was allowed to settle, and 800 μL of the supernatant was transferred to a new tube. 200 μL of Trizol and 200 μL of chloroform were added to the solution. The mixture was vortexed for 30 s and centrifuged at 15,000 rpm for 15 min at 4° C. 600 μL of the supernatant was added to a tube containing 600 μL 2-propanol. The solution was mixed by pipetting and incubated at room temperature for 10 min. This solution was centrifuged at 15,000 rpm for 15 min at 4° C. The supernatant was discarded, and 1 mL of 70% ethanol was added to the pellet. This solution was centrifuged at 10,000 rpm for 5 min at 4° C. The supernatant was discarded and the pellet was air-dried. RNA was reconstituted in 30 μL ddH2O. RNA sequencing was performed on the HiSeq platform (Illumina). For bioinformatics analysis, reads were aligned using STAR [61], sorting and marking duplicates were done by Picard, and read counting was done by Featurecounts [62]. EdgeR [63] and R [58] were used to create figures to visualize differential gene expression. For differential expression, an FDR-adjusted q value < 0.05 was used. In a less stringent analysis of differentially expressed genes, an unadjusted p value < 0.05 was used.
Quantitative PCR (qPCR)
RNA was extracted from adult day-3 worms by the phenol-chloloroform method using Trizol reagent (Thermo Fisher Scientific) as described above. cDNA was synthesized using SuperScript III with oligo-dT primers (Thermo Fisher Scientific). 1 μL of oligo-dT primers, 1 μg of RNA, 1 μL of 10 mM dNTP mix and ddH20 was added to 13 μL. The solution was incubated at 65° C for 5 min and then on ice for 1 min. 4 μL of 5X First-Strand Buffer, 1 μL 0.1 M DTT, and 1 μL of SuperScript III was added to make 20 μL of solution. This solution was mixed by pipetting and incubated at 50° C for 60 min. The reaction was inactivated by heating at 70° C for 15 min. qPCR was carried out using the cDNA with Luna Universal qPCR Master Mix (New England Biolabs, Ipswich, MA) using StepOnePlus (Thermo Fisher Scientific). 100 ng of template cDNA was used for Luna Universal qPCR. The peroxisomal membrane protein pmp-3 was used as the reference control for all qPCR experiments. Primers used for qPCR are listed in Supplementary Table 5.
RNA interference
The Ahringer RNAi library was used to reduce the expression of target genes (F59H6.8, F59H6.9, B0047.3). Frozen stocks of the RNAi bacteria were streaked onto agar plates containing 100 μg/mL ampicillin. Single colonies of the RNAi bacteria were cultured for 8 h at 37° C with vigorous shaking in lysogeny broth (LB) containing 100 μg/mL ampicillin. 100 μL of bacterial culture was spread on NGM plates with 50 μg/mL ampicillin and 1.0 mM isopropyl β-D-1-thiogalactopyranoside (IPTG). Plates with RNAi bacteria were dried overnight with the lid on at room temperature (25° C). Adult wild-type worms were placed on RNAi plates and allowed to lay eggs for 3 h. The locomotor ability of the offspring was tested from the first day of adulthood. For mock control, an empty vector L4440 was used.
Statistics
All results are expressed as means with error bars representing a 95% confidence interval. For pairwise comparisons, Student’s two-tailed t test was used with Excel 2010 (Microsoft). For multiple comparisons to a control, one-way ANOVA was followed with Dunnett’s post hoc test using R [58]. For multiple comparisons, one-way ANOVA was followed with Tukey’s Honest Significant Difference test using R [58]. For lifespan comparisons, log-rank test was used with R [58]. Statistical significance was set at *P < 0.05; **P < 0.01; ***P < 0.001.
Data availability
All isolated strains are available upon request. DNA and RNA sequencing results are available on NCBI sequence read archive PRJNA530333.
Author Contributions
KK and INM designed the study. KK carried out experiments and analyzed data. KK wrote the original draft of the manuscript. INM critically revised the manuscript.
Acknowledgments
We thank H. Goto, M. Kanda, M. Kawamitsu, S. Yamasaki, N. Arakaki and other DNA sequencing section members for technical assistance with DNA and RNA sequencing. We are grateful to T. Murayama and E. Saita for technical support and advice. We thank D. Van Vactor, B. Kuhn, and members of the Maruyama unit for helpful discussions regarding this work. We are grateful to H. Ohtaki for administrative support. We thank the Caenorhabditis Genetics Center, which is funded by NIH Office of Research Infrastructure Programs (P40 OD010440), for providing worm strains.
Conflicts of Interest
The authors declare that there are no conflicts of interest.
Funding
This work was partly supported by Okinawa Institute of Science and Technology Graduate University, and KK was supported by Japan Society for the Promotion of Science KAKENHI (Grant 16J06404).
References
- 1. Groessl EJ, Kaplan RM, Rejeski WJ, Katula JA, King AC, Frierson G, Glynn NW, Hsu FC, Walkup M, Pahor M. Health-related quality of life in older adults at risk for disability. Am J Prev Med. 2007; 33:214–18. https://doi.org/10.1016/j.amepre.2007.04.031 [PubMed]
- 2. Trombetti A, Reid KF, Hars M, Herrmann FR, Pasha E, Phillips EM, Fielding RA. Age-associated declines in muscle mass, strength, power, and physical performance: impact on fear of falling and quality of life. Osteoporos Int. 2016; 27:463–71. https://doi.org/10.1007/s00198-015-3236-5 [PubMed]
- 3. Brenner S. The genetics of Caenorhabditis elegans. Genetics. 1974;77: 71–94. [PubMed]
- 4. Iwasa H, Yu S, Xue J, Driscoll M. Novel EGF pathway regulators modulate C. Elegans healthspan and lifespan via EGF receptor, PLC-gamma, and IP3R activation. Aging Cell. 2010; 9:490–505. https://doi.org/10.1111/j.1474-9726.2010.00575.x [PubMed]
- 5. Li G, Gong J, Liu J, Liu J, Li H, Hsu AL, Liu J, Xu XZ. Genetic and pharmacological interventions in the aging motor nervous system slow motor aging and extend life span in C. Elegans. Sci Adv. 2019; 5:eaau5041. https://doi.org/10.1126/sciadv.aau5041 [PubMed]
- 6. Kawamura K, Maruyama IN. Forward genetic screen for Caenorhabditis elegans mutants with a shortened locomotor healthspan. G3 (Bethesda). 2019; 9:2415–23. https://doi.org/10.1534/g3.119.400241 [PubMed]
- 7. Shi Y, Mello C. A CBP/p300 homolog specifies multiple differentiation pathways in caenorhabditis elegans. Genes Dev. 1998; 12:943–55. https://doi.org/10.1101/gad.12.7.943 [PubMed]
- 8. Dufourcq P, Victor M, Gay F, Calvo D, Hodgkin J, Shi Y. Functional requirement for histone deacetylase 1 in caenorhabditis elegans gonadogenesis. Mol Cell Biol. 2002; 22:3024–34. https://doi.org/10.1128/mcb.22.9.3024-3034.2002 [PubMed]
- 9. Ranawade AV, Cumbo P, Gupta BP. Caenorhabditis elegans histone deacetylase hda-1 is required for morphogenesis of the vulva and LIN-12/Notch-mediated specification of uterine cell fates. G3 (Bethesda). 2013; 3:1363–74. https://doi.org/10.1534/g3.113.006999 [PubMed]
- 10. Bates EA, Victor M, Jones AK, Shi Y, Hart AC. Differential contributions of caenorhabditis elegans histone deacetylases to huntingtin polyglutamine toxicity. J Neurosci. 2006; 26:2830–38. https://doi.org/10.1523/JNEUROSCI.3344-05.2006 [PubMed]
- 11. Edwards C, Canfield J, Copes N, Rehan M, Lipps D, Bradshaw PC. D-beta-hydroxybutyrate extends lifespan in C. Elegans. Aging (Albany NY). 2014; 6:621–44. https://doi.org/10.18632/aging.100683 [PubMed]
- 12. Oh HJ, Abraham LS, van Hengel J, Stove C, Proszynski TJ, Gevaert K, DiMario JX, Sanes JR, van Roy F, Kim H. Interaction of α-catulin with dystrobrevin contributes to integrity of dystrophin complex in muscle. J Biol Chem. 2012; 287:21717–28. https://doi.org/10.1074/jbc.M112.369496 [PubMed]
- 13. Kim H, Rogers MJ, Richmond JE, McIntire SL. SNF-6 is an acetylcholine transporter interacting with the dystrophin complex in caenorhabditis elegans. Nature. 2004; 430:891–96. https://doi.org/10.1038/nature02798 [PubMed]
- 14. Kim H, Pierce-Shimomura JT, Oh HJ, Johnson BE, Goodman MB, McIntire SL. The dystrophin complex controls bk channel localization and muscle activity in caenorhabditis elegans. PLoS Genet. 2009; 5:e1000780. https://doi.org/10.1371/journal.pgen.1000780 [PubMed]
- 15. Grisoni K, Gieseler K, Mariol MC, Martin E, Carre-Pierrat M, Moulder G, Barstead R, Ségalat L. The stn-1 syntrophin gene of C.elegans is functionally related to dystrophin and dystrobrevin. J Mol Biol. 2003; 332:1037–46. https://doi.org/10.1016/j.jmb.2003.08.021 [PubMed]
- 16. Zhou S, Chen L. Neural integrity is maintained by dystrophin in C. Elegans. J Cell Biol. 2011; 192:349–63. https://doi.org/10.1083/jcb.201006109 [PubMed]
- 17. Bessou C, Giugia JB, Franks CJ, Holden-Dye L, Ségalat L. Mutations in the Caenorhabditis elegans dystrophin-like gene dys-1 lead to hyperactivity and suggest a link with cholinergic transmission. Neurogenetics. 1998; 2:61–72. https://doi.org/10.1007/s100480050053 [PubMed]
- 18. Hoffman EP, Brown RH
Jr , Kunkel LM. Dystrophin: the protein product of the duchenne muscular dystrophy locus. Cell. 1987; 51:919–28. https://doi.org/10.1016/0092-8674(87)90579-4 [PubMed] - 19. Watson PJ, Fairall L, Santos GM, Schwabe JW. Structure of HDAC3 bound to co-repressor and inositol tetraphosphate. Nature. 2012; 481:335–40. https://doi.org/10.1038/nature10728 [PubMed]
- 20. Schuetz A, Min J, Allali-Hassani A, Schapira M, Shuen M, Loppnau P, Mazitschek R, Kwiatkowski NP, Lewis TA, Maglathin RL, McLean TH, Bochkarev A, Plotnikov AN, et al. Human HDAC7 harbors a class IIa histone deacetylase-specific zinc binding motif and cryptic deacetylase activity. J Biol Chem. 2008; 283:11355–63. https://doi.org/10.1074/jbc.M707362200 [PubMed]
- 21. Millard CJ, Watson PJ, Celardo I, Gordiyenko Y, Cowley SM, Robinson CV, Fairall L, Schwabe JW. Class I HDACs share a common mechanism of regulation by inositol phosphates. Mol Cell. 2013; 51:57–67. https://doi.org/10.1016/j.molcel.2013.05.020 [PubMed]
- 22. Perez-Torrado R, Yamada D, Defossez PA. Born to bind: the BTB protein-protein interaction domain. Bioessays. 2006; 28:1194–202. https://doi.org/10.1002/bies.20500 [PubMed]
- 23. Thomas JH. Adaptive evolution in two large families of ubiquitin-ligase adapters in nematodes and plants. Genome Res. 2006; 16:1017–30. https://doi.org/10.1101/gr.5089806 [PubMed]
- 24. Haberland M, Montgomery RL, Olson EN. The many roles of histone deacetylases in development and physiology: implications for disease and therapy. Nat Rev Genet. 2009; 10:32–42. https://doi.org/10.1038/nrg2485 [PubMed]
- 25. Montgomery RL, Potthoff MJ, Haberland M, Qi X, Matsuzaki S, Humphries KM, Richardson JA, Bassel-Duby R, Olson EN. Maintenance of cardiac energy metabolism by histone deacetylase 3 in mice. J Clin Invest. 2008; 118:3588–97. https://doi.org/10.1172/JCI35847 [PubMed]
- 26. Kang HL, Benzer S, Min KT. Life extension in drosophila by feeding a drug. Proc Natl Acad Sci USA. 2002; 99:838–43. https://doi.org/10.1073/pnas.022631999 [PubMed]
- 27. Evason K, Collins JJ, Huang C, Hughes S, Kornfeld K. Valproic acid extends caenorhabditis elegans lifespan. Aging Cell. 2008; 7:305–17. https://doi.org/10.1111/j.1474-9726.2008.00375.x [PubMed]
- 28. Zhao Y, Sun H, Lu J, Li X, Chen X, Tao D, Huang W, Huang B. Lifespan extension and elevated hsp gene expression in drosophila caused by histone deacetylase inhibitors. J Exp Biol. 2005; 208:697–705. https://doi.org/10.1242/jeb.01439 [PubMed]
- 29. Walsh ME, Bhattacharya A, Sataranatarajan K, Qaisar R, Sloane L, Rahman MM, Kinter M, Van Remmen H. The histone deacetylase inhibitor butyrate improves metabolism and reduces muscle atrophy during aging. Aging Cell. 2015; 14:957–70. https://doi.org/10.1111/acel.12387 [PubMed]
- 30. Benito E, Urbanke H, Ramachandran B, Barth J, Halder R, Awasthi A, Jain G, Capece V, Burkhardt S, Navarro-Sala M, Nagarajan S, Schütz AL, Johnsen SA, et al. HDAC inhibitor-dependent transcriptome and memory reinstatement in cognitive decline models. J Clin Invest. 2015; 125:3572–84. https://doi.org/10.1172/JCI79942 [PubMed]
- 31. Ying M, Xu R, Wu X, Zhu H, Zhuang Y, Han M, Xu T. Sodium butyrate ameliorates histone hypoacetylation and neurodegenerative phenotypes in a mouse model for DRPLA. J Biol Chem. 2006; 281:12580–86. https://doi.org/10.1074/jbc.M511677200 [PubMed]
- 32. Struhl K. Histone acetylation and transcriptional regulatory mechanisms. Genes Dev. 1998; 12:599–606. https://doi.org/10.1101/gad.12.5.599 [PubMed]
- 33. Eberharter A, Becker PB. Histone acetylation: a switch between repressive and permissive chromatin. Second in review series on chromatin dynamics. EMBO Rep. 2002; 3:224–29. https://doi.org/10.1093/embo-reports/kvf053 [PubMed]
- 34. Wang A, Kurdistani SK, Grunstein M. Requirement of Hos2 histone deacetylase for gene activity in yeast. Science. 2002; 298:1412–14. https://doi.org/10.1126/science.1077790 [PubMed]
- 35. Nusinzon I, Horvath CM. Histone deacetylases as transcriptional activators? role reversal in inducible gene regulation. Sci STKE. 2005; 2005:re11. https://doi.org/10.1126/stke.2962005re11 [PubMed]
- 36. Ozawa Y, Towatari M, Tsuzuki S, Hayakawa F, Maeda T, Miyata Y, Tanimoto M, Saito H. Histone deacetylase 3 associates with and represses the transcription factor GATA-2. Blood. 2001; 98:2116–23. https://doi.org/10.1182/blood.v98.7.2116 [PubMed]
- 37. Schroeder TM, Kahler RA, Li X, Westendorf JJ. Histone deacetylase 3 interacts with runx2 to repress the osteocalcin promoter and regulate osteoblast differentiation. J Biol Chem. 2004; 279:41998–2007. https://doi.org/10.1074/jbc.M403702200 [PubMed]
- 38. Pintard L, Willis JH, Willems A, Johnson JL, Srayko M, Kurz T, Glaser S, Mains PE, Tyers M, Bowerman B, Peter M. The BTB protein MEL-26 is a substrate-specific adaptor of the CUL-3 ubiquitin-ligase. Nature. 2003; 425:311–16. https://doi.org/10.1038/nature01959 [PubMed]
- 39. Xu L, Wei Y, Reboul J, Vaglio P, Shin TH, Vidal M, Elledge SJ, Harper JW. BTB proteins are substrate-specific adaptors in an SCF-like modular ubiquitin ligase containing CUL-3. Nature. 2003; 425:316–21. https://doi.org/10.1038/nature01985 [PubMed]
- 40. Zhuang M, Calabrese MF, Jiang L, Waddell MB, Nourse A, Hammel M, Miller DJ, Walden H, Duda DM, Seyedin SN, Hoggard T, Harper JW, White KP, Schulman BA. Structures of SPOP-Substrate Complexes: Insights into Molecular Architectures of BTB-Cul3 Ubiquitin Ligases. Mol Cell. 2009; 36:39–50. https://doi.org/10.1016/j.molcel.2009.09.022 [PubMed]
- 41. Shteingauz A, Cohen E, Biala Y, Treinin M. The BTB-MATH protein BATH-42 interacts with RIC-3 to regulate maturation of nicotinic acetylcholine receptors. J Cell Sci. 2009; 122:807–12. https://doi.org/10.1242/jcs.036343 [PubMed]
- 42. Fox RM, Von Stetina SE, Barlow SJ, Shaffer C, Olszewski KL, Moore JH, Dupuy D, Vidal M, Miller DM
3rd . A gene expression fingerprint of C. Elegans embryonic motor neurons. BMC Genomics. 2005; 6:42. https://doi.org/10.1186/1471-2164-6-42 [PubMed] - 43. Kaletsky R, Yao V, Williams A, Runnels AM, Tadych A, Zhou S, Troyanskaya OG, Murphy CT. Transcriptome analysis of adult caenorhabditis elegans cells reveals tissue-specific gene and isoform expression. PLoS Genet. 2018; 14:e1007559. https://doi.org/10.1371/journal.pgen.1007559 [PubMed]
- 44. Karczewski KJ, Francioli LC, Tiao G, Cummings BB, Alföldi J, Wang Q, Collins RL, Laricchia KM, Ganna A, Birnbaum DP, Gauthier LD, Brand H, Solomonson M, et al, and Genome Aggregation Database Consortium. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature. 2020; 581:434–43. https://doi.org/10.1038/s41586-020-2308-7 [PubMed]
- 45. Schapira M. Structural Biology of Human Metal-Dependent Histone Deacetylases. In: Yao TP, Seto E, editors. Histone Deacetylases: the Biology and Clinical Implication. Berlin, Heidelberg: Springer Berlin Heidelberg; 2011. pp. 225–240. https://doi.org/10.1007/978-3-642-21631-2_10
- 46. Gregoretti IV, Lee YM, Goodson HV. Molecular evolution of the histone deacetylase family: functional implications of phylogenetic analysis. J Mol Biol. 2004; 338:17–31. https://doi.org/10.1016/j.jmb.2004.02.006 [PubMed]
- 47. Hu E, Chen Z, Fredrickson T, Zhu Y, Kirkpatrick R, Zhang GF, Johanson K, Sung CM, Liu R, Winkler J. Cloning and characterization of a novel human class I histone deacetylase that functions as a transcription repressor. J Biol Chem. 2000; 275:15254–64. https://doi.org/10.1074/jbc.M908988199 [PubMed]
- 48. Pothof J, van Haaften G, Thijssen K, Kamath RS, Fraser AG, Ahringer J, Plasterk RH, Tijsterman M. Identification of genes that protect the C. Elegans genome against mutations by genome-wide RNAi. Genes Dev. 2003; 17:443–48. https://doi.org/10.1101/gad.1060703 [PubMed]
- 49. Li H, Durbin R. Fast and accurate short read alignment with burrows-wheeler transform. Bioinformatics. 2009; 25:1754–60. https://doi.org/10.1093/bioinformatics/btp324 [PubMed]
- 50. Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R, and 1000 Genome Project Data Processing Subgroup. The sequence alignment/map format and SAMtools. Bioinformatics. 2009; 25:2078–79. https://doi.org/10.1093/bioinformatics/btp352 [PubMed]
- 51. Blankenberg D, Von Kuster G, Coraor N, Ananda G, Lazarus R, Mangan M, Nekrutenko A, Taylor J. Galaxy: a web-based genome analysis tool for experimentalists. Curr Protoc Mol Biol. 2010. https://doi.org/10.1002/0471142727.mb1910s89 [PubMed]
- 52. Cingolani P, Platts A, Wang le L, Coon M, Nguyen T, Wang L, Land SJ, Lu X, Ruden DM. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin). 2012; 6:80–92. https://doi.org/10.4161/fly.19695 [PubMed]
- 53. Giardine B, Riemer C, Hardison RC, Burhans R, Elnitski L, Shah P, Zhang Y, Blankenberg D, Albert I, Taylor J, Miller W, Kent WJ, Nekrutenko A. Galaxy: a platform for interactive large-scale genome analysis. Genome Res. 2005; 15:1451–55. https://doi.org/10.1101/gr.4086505 [PubMed]
- 54. Goecks J, Nekrutenko A, Taylor J, and Galaxy Team. Galaxy: a comprehensive approach for supporting accessible, reproducible, and transparent computational research in the life sciences. Genome Biol. 2010; 11:R86. https://doi.org/10.1186/gb-2010-11-8-r86 [PubMed]
- 55. Koboldt DC, Chen K, Wylie T, Larson DE, McLellan MD, Mardis ER, Weinstock GM, Wilson RK, Ding L. VarScan: variant detection in massively parallel sequencing of individual and pooled samples. Bioinformatics. 2009; 25:2283–85. https://doi.org/10.1093/bioinformatics/btp373 [PubMed]
- 56. Minevich G, Park DS, Blankenberg D, Poole RJ, Hobert O. CloudMap: a cloud-based pipeline for analysis of mutant genome sequences. Genetics. 2012; 192:1249–69. https://doi.org/10.1534/genetics.112.144204 [PubMed]
- 57. Nussbaum-Krammer CI, Neto MF, Brielmann RM, Pedersen JS, Morimoto RI. Investigating the spreading and toxicity of prion-like proteins using the metazoan model organism C. Elegans. J Vis Exp. 2015; 52321. https://doi.org/10.3791/52321 [PubMed]
- 58. Team RC. R: A language and environment for statistical computing [Internet]. Vienna, Austria: R Foundation for Statistical Computing.; 2015. 2015.
- 59. Dokshin GA, Ghanta KS, Piscopo KM, Mello CC. Robust genome editing with short single-stranded and long, partially single-stranded DNA donors in Caenorhabditis elegans. Genetics. 2018; 210:781–87. https://doi.org/10.1534/genetics.118.301532 [PubMed]
- 60. Schneider CA, Rasband WS, Eliceiri KW. NIH image to ImageJ: 25 years of image analysis. Nat Methods. 2012; 9:671–75. https://doi.org/10.1038/nmeth.2089 [PubMed]
- 61. Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, Batut P, Chaisson M, Gingeras TR. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013; 29:15–21. https://doi.org/10.1093/bioinformatics/bts635 [PubMed]
- 62. Liao Y, Smyth GK, Shi W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics. 2014; 30:923–30. https://doi.org/10.1093/bioinformatics/btt656 [PubMed]
- 63. Robinson MD, McCarthy DJ, Smyth GK. edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010; 26:139–40. https://doi.org/10.1093/bioinformatics/btp616 [PubMed]



