Introduction
In recent years, the Deptor protein (DEP-domain containing mTOR-interacting protein), also known as DEPDC6 (DEP-domain containing protein 6), with gene ID number Q8TB45, has been described. It is a negative regulator of the mTORC1 and mTORC2 signaling pathways, inhibiting kinase activity of both complexes [1]. The Deptor protein is encoded by the DEPTOR gene located on chromosome 8 in the long arm in the 8q24 region [2], of which at least two isoforms have been reported by alternative splicing [3]. As mentioned, Deptor is a negative regulator of mTORC1 and mTORC2. mTORTC1 is involved in anabolism, growth and proliferation signaling pathways [4]. In addition to the growth factors and nutrients that can activate mTORC1, it can also be activated by other factors such as hypoxia, inflammation, energy deficiency, cholesterol and nucleotides. Therefore, mTORC1, through its activation, promotes metabolism and participates in the synthesis pathways of proteins, lipids and nucleotides, contributing to cell growth and proliferation [4, 5]. Interestingly, the activation of mTORC1 inhibits catabolism, blocking autophagy and lysosomal biogenesis. This allows Deptor, which inhibits mTORC1 activity, to activate catabolism. Similarly, activation of this complex can trigger negative feedback regulation of growth factor signaling pathways by regulating IRS1 (Insulin receptor substrate-1) [6, 7] as well as also of the GRB10 factor (growth factor receptor-bound 10) [8, 9]. This negative regulation by mTORC1-mediated feedback can also be positively regulated by the inhibitory effect of Deptor on mTORC1
mTORC2 integrates the signaling pathways of growth factors that regulate processes such as metabolism, survival, organization of the cytoskeleton and cell mobility [4, 5] Activation of mTORC2 regulates SGK1 (Serum and glucocorticoid (GC) -induced protein kinase-1), PKC-α (protein kinase C-α), and AKT (protein kinase B/AKT) [10]. Much like mTORC1, Deptor plays an inhibitory role in the activation of these pathways dependent on the activation of mTORC2.
Due to its ability to inhibit mTOR pathway, studies in Deptor have been of great interest regarding cancer development and progression. Studies reveal a low expression of Deptor in most tumors [1], with some exceptions such as multiple myeloma (MM), thyroid carcinoma and lung cancer, where its expression is high [11]. So, it is suggested that Deptor has a dual role in human malignancies, acting either as an oncogene or as a tumor suppressor [12].
The focus of this review is to discuss current knowledge of Deptor regarding its structure, function and regulation, as well as its role specifically in hematological malignancies such as Multiple Myeloma, Leukemia and Lymphoma.
Structure of Deptor
Deptor is a 409 amino acid protein, weighing 48 kDa and is only expressed in vertebrates [1]. It is a ubiquitously expressed and highly conserved protein that contains two tandems of DEP domains in the amino-terminal region and one PDZ domain at the Carboxyl-terminal region. (Figure 1). The two DEP domains are 84 and 74 aa respectively and are located at residues T36-K119 and S145-M219 respectively of the amino-terminal region [12, 13]. Although their role in these DEP domains is not fully understood, they are believed to play an important role in associating membrane and signaling proteins. These DEP domains have been identified in a large number of proteins [14] and they are highly conserved in sequence and structure. DEP domains function as a membrane anchor [15], signaling regulators at G protein-coupled receptors [14]. Although tandem of two DEP domains are infrequent in proteins containing DEP domains, it’s believed that the tandem of DEP domains may have an important role in mediating the connection to PI3K, just as the inhibitory role of Deptor on mTOR activity can be mediated by the structural organization of these tandem DEP domains [4]. The PDZ domain in Deptor contains 78 aa located in the carboxyl terminal region between residues T330-L407 and is responsible for the protein-protein interaction [1, 13]. This domain is responsible for binding with mTOR, likewise PDZ domains are present in a wide variety of proteins and are highly conserved sequences of G-L-G-F, which participate in folding and interaction with the target protein [16]. Interestingly, some of the proteins that have PDZ domains belong to the Central Nervous System, involved in the neuronal synapse [17]. The PDZ domain can regulate cell signaling by inhibiting phosphorylation of residues in the interaction domain in the carboxyl terminal region on its ligand or in the PDZ domain by itself [16]. Studies by Peterson and Co [1] have demonstrated Deptor phosphorylation at 14 different residues (T and S) located between the DEP2 and PDZ domain junction and comprising between residues T241-S299, determined by spectrometric studies. These studies reveal that the stability of Deptor and its binding to mTOR is in part regulated by Deptor phosphorylation. Additionally, studies by mass spectrometry in our group have identified two additional Deptor phosphorylation residues (S235, and S260), where preliminary studies show that at least one of these residues participates in the stability and degradation of Deptor (Figure 1) [18]. A sequence located in the junction region between the C-terminus of the sequence and the amino-terminal region of PDZ, designated as Degron and comprising the S286-S291 region (SSGYFS), has also been characterized. Degron is recognized and degraded by βTrCP1, in which the phosphorylation of three sites susceptible to phosphorylation is necessary for their interaction with βTrCP1 (S286, S287 and S291) [19]. This triggers their ubiquitylation and degradation, which may play a role in Deptor function (Figure 1) [20]. Studies have reported that Deptor undergoes ubiquitin ligase-mediated ubiquitination by the SCFβTrCP E3. Upon stimulation of growth factors, Deptor is rapidly degraded by the ubiquitin-proteasome pathway to ensure proper activation of the mTOR pathway [20]. This can be counteracted by the interaction with UBTOR [21], which interacts with the PDZ domain of Deptor, promoting its stability and inhibiting its ubiquitination and consequent Deptor degradation [21].
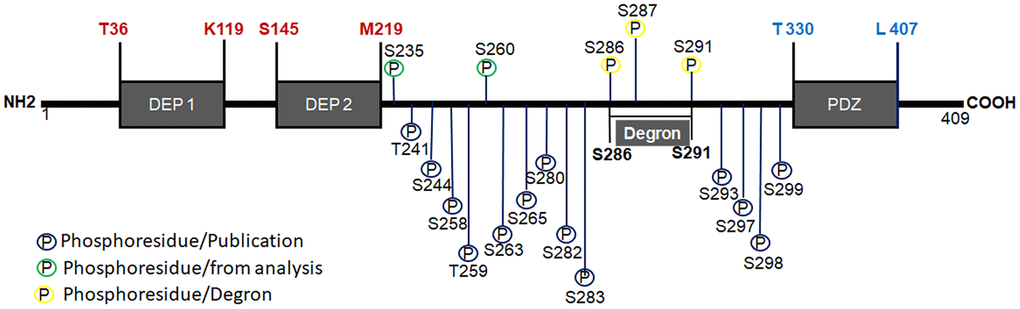
Figure 1. Structure of Deptor. Schematic representation of Deptor and his two DEP domains are indicated as well PDZ domain. Degron motif and phosphorylation residues are indicated. phosphorylation at 15 different residues (T and S) located between the DEP2 and PDZ domain junction and comprising between residues T241-S299, which was determined by spectrometric studies, are also indicated.
As we mentioned above, at least two Deptor isoforms originated by alternative splicing have been reported. Isoform 1 has been chosen as the canonical sequence. Isoform 2 differs from the canonical sequence in that it lacks the L42-K142 region (Q8TB45-2) situated in the C-terminal region, encoding a short form of 308 aa (L143-C409) [3]. Therefore, it lacks the DEP1 domain and part of the DEP2 domain, and its possible function is unknown.
As we mentioned above, like another proteins, many of their biological and regulatory functions are controlled by phosphorylation events on Deptor. Since its phosphorylation determines its binding and regulation with the mTOR complex [1], as well as the stability of the protein. To date, at least 18 sites susceptible to phosphorylation of Deptor are known, which suggest of the importance of the phosphorylation events in this protein.
Obviously, the observed phosphorylation events on Deptor as well as other posttranscriptional modification, may have significance in relation to oncogenic pathobiology, such relevance of phosphorylation events has been clearly demonstrated in other hematologic malignances [22–29] It remaining to be seen of these events also regulate contribution of Deptor to carcinogenesis. For example, our recently data, which demonstrate of ERK-dependent phosphorylation of Deptor which maintains its stability, suggests a critical effect in Myeloma. ERK activation by growth factors (i.e., IL-6, EGF-1), as well as mutated RAS, may promote Myeloma progression, in part, via stabilization of Deptor.
Deptor expression and localization
Important levels of Deptor expression in different tissues have been reported, as well as high levels of Deptor mRNA. An important expression of Deptor in serum, tonsils, bone marrow cell stroma, frontal cortex, spinal cord, stomach, colon, rectum, liver, kidney, spleen, salivary glands, thyroid, adrenal, pancreas, islets of Langerhans, gallbladder, prostate, bladder, skin, placenta, uterus, cervix, ovary, testis, seminal vesicles, as well as in different cell lines, is reported through an analysis of integrated proteomic protein expression (www.proteomicsbd.org) [30]. Of these cell lines, the most important Deptor expression is in breast cancer cell lines (LCC2), Lung cancer (NCI-H522), colon cancer (CCK-81 and HCA-46), cervical cancer (Hela) and multiple myeloma (8226). At the intracellular level, Deptor is expressed in cytosol, mitochondria and nucleus, with less expression in the plasma membrane, cytoskeleton, endoplasmic reticulum, endosome and lysosomes (according to an analysis in COMPONENTS Subcellular location data base: (https://compartments.jensenlab.org) [31] and the Atlas of Human Proteins [www.proteinatlas.org]) [32]. Different studies describe that the location of Deptor correlates with its function [1, 4, 5, 10, 33, 34].
Deptor regulation
Studies have demonstrated the different regulatory mechanisms of Deptor, including diverse and complicated epigenetic, post-transcriptional and transcriptional mechanisms. Different studies involve the mTORC1 and mTORC2 complex in downregulation of Deptor at the post-transcriptional level (e.g., phosphorylation) [1]. However, recent study has focused on knowing the transcriptional regulation of Deptor.
Epigenetic factors
Deptor regulation has been associated with epigenetic processes, as reported in rat kidney cells (NRK-52E), in which inhibition of histone methyltransferase EZH2, responsible for the trimethylation of histone H3 lysine 27 (H3K27me3), was related to an increase in Deptor expression [35]. Effect of inhibition of EZH2 on Deptor expression was confirmed in HCT116 colorectal carcinoma cells, in which interfering RNA treatment as well as a specific EZH2 inhibitor resulted in an increase in Deptor expression [36]. On the other hand, it was reported that in HEC1B cells silencing of the arginosuccinate synthase 1 gene, resulted in decreased Deptor expression as a result of altered methylation of histones, suggesting this mechanism as an epigenetic regulation of Deptor [37]. In another study in prostate cancer cells, there was the indication that treatment with the androgen receptor agonist dihydrotestosterone induces a decrease in acetylation of lysine 9 and 14 in histone 3 in the region corresponding to the 4th intron of Deptor and as a consequence, suppresses the expression of Deptor mRNA [38]. Additionally, an interaction between Deptor and Glycine-N-Methyltransferase has been reported [39]. However, the involvement of Glycine-N-methyltransferase as an epigenetic regulator of Deptor needs to be studied.
Transcription factors
In this review, we focus on reviewing Deptor’s transcriptional regulatory mechanisms.
Notch1: Studies describe a direct regulation of Deptor expression by Notch by mediating binding to NCID in a conserved sequence near the Deptor promoter start codon [40], in which Notch1 induces a high expression of Deptor, resulting in mTORC1 inhibition. This compensates for feedback inhibition of mTORC1 signaling to PI3K and induces Akt phosphorylation at residues S437 and T308, which contributes to AKT activation, a critical factor in T-ALL leukogenesis. The Notch pathway plays an important role in various cellular processes, including stem cell self-renewal, proliferation, and differentiation. Several studies have identified recurrent mutations in hematologic malignancies that make Notch a therapeutic target in acute T-cell lymphoblastic leukemia, chronic lymphocytic leukemia, and mantle cell lymphoma [41].
GR: Other studies suggest that Glucocorticoids, by binding to their nuclear receptor (GR), are capable of inducing Deptor expression [5] where it was shown that elements of response to GC were identified upstream of the Deptor transcriptional initiation site [42], suggesting that Deptor plays a role in GR-mediated inflammation. Interestingly, studies also suggest that nuclear estrogen receptors directly regulate Deptor expression.
ER-α: This has been observed in ER-α positive prostate cell lines where ER-α antagonists represses Deptor expression in these cells [43]. While it has not been established whether ER-α directly regulates Deptor transcription, it is suggested that ER-α may interact with the response elements identified in the Deptor promoter.
VDR: Several response elements for the vitamin D receptor (VDR) have been identified, and since vitamin D inhibits the activity of mTOR [44], it is suggested that VDR can positively regulate the Deptor expression [45].
AR: Studies suggest a reverse effect for nuclear androgen receptors, since AR directly inhibits Deptor expression by binding to androgen response elements identified in an intron of the DEPTOR gene [38].
AATF: Recent studies have described that AATF is capable to regulate the expression of various genes involved in cellular processes. These genes are KLF4, C-Myc, p53, p21, mTORC1 and recently reported Deptor [46–48]. Interestingly, this regulation of Deptor by AATF could be due to the direct binding to its promoter, or it can be down-regulated by regulating the expression of miR-2909, which binds to the 5” UTR region of Deptor by inhibiting its expression (Figure 2) [49].
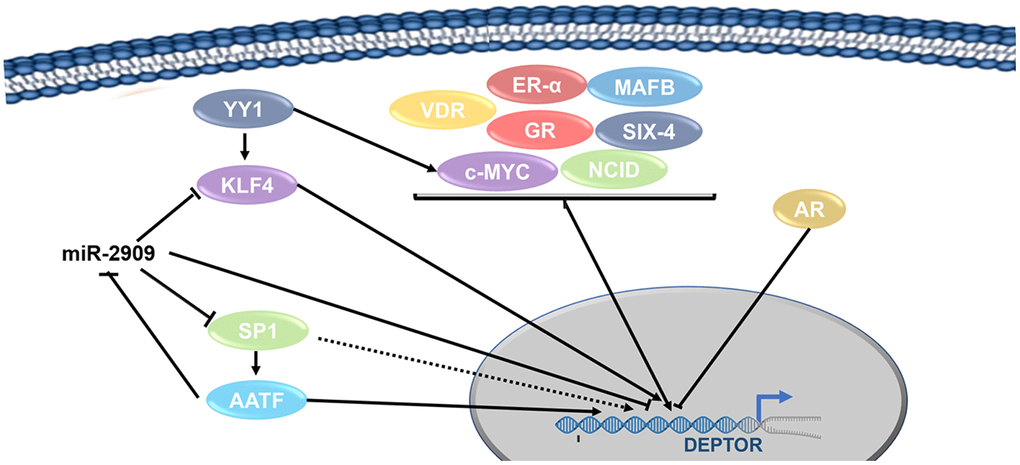
Figure 2. Transcription factors network involved in the regulation of Deptor. Transcription regulation of Deptor expression by Transcription factors are represented. Transcription factors involved in the regulation of Deptor revised here are shown.
c-MAF: Studies suggest that the c-MAF proto-oncogene and the MAFB transcription factor who regulates cellular processes are capable of regulating Deptor expression. This happens specifically in MM where high Deptor expression is shown [1], although it is unknown to date whether they do it directly or not.
Six4: Another transcriptional regulator that has been reported to be capable of regulating Deptor is Six4, which is a transcriptional factor involved in cell differentiation, cell migration, and cell survival. Consensus binding sites for Six4 have been identified in the Deptor promoter, and mutations at these sites inhibit activation of Deptor expression by Six4 and its cofactor Baf60c [50].
C-Myc: Additional studies show that Deptor is regulated by Wnt/β-Catenin/c-Myc and has implications for the pathogenesis of colorectal cancer. Where a direct transcriptional regulation of C-Myc on the Deptor promoter is demonstrated (Figure 2) [51].
Predicted transcription factors in the regulation of deptor
Our group carried out a bioinformatic study using the Jaspar platform (http://jaspar.genereg.net) [52], in order to predict the possible binding of transcription factors in the Deptor promoter. This analysis reveals according to the significant score, that transcription factors such as Arid3a, CDX1, CDX2, CREB1, CUX1, KLF4, KLF14, SP1 BCL-6 and YY1 may be involved in the transcriptional regulation of Deptor (Table 1).
Table 1. Predicted transcription factors involved in the regulation of Deptor.
| Transcription Factor | Score (Jaspar) | Position | Predictive sequence (Jaspar) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Start | End | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Arid3a | 9.84 | -1693 | -1688 | ATTAAA | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CDX1 | 12.25 | -242 | -234 | ACAATAAAA | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CDX2 | 13.312 | -244 | -234 | GTACAATAAA | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CREB1 | 10.52 | -770 | -759 | CCTGACCTAGG | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CUX1 | 12.19 | -1835 | -1826 | TTATCGATAG | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KLF4 | 11.50 | -258 | -248 | CCACACCCAA | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KLF14 | 14.45 | -82 | -69 | GGCCCCGCCCCCG | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SP1 | 8.59 | -81 | -72 | GGGGGCGGGGC | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| YY1 | 8.38 | 64 | 69 | GCCATC | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BCL6 | 9.29 | -1387 | -1372 | TAACTTTCTAGGCAGA | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted transcription factors using Jaspar Platform, with potential binding sites in the Deptor promoter. In this table we list the transcription factors binding sites with the highest score according to Jaspar Score. One or more sites with lower score can be found in the database (Data not shown). | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Arid3a: Transcription factor Arid3a is a paralog of the family of interactive AT-rich domains and is associated with the regulation of genes in the development of B cells and cancer such as ovarian cancer [53]. Interestingly we identified at least 6 different possible binding sites for this factor in the Deptor promoter (Table 1). Transcription factor ARID3A emerges as one of the most consistently and differentially expressed genes in ABC-DLBCL, compared to GC-DLBCL [54]. In B-ALL (B-cell Acute Lymphoblastic Leukemia), ARID3a is targeted by miR-125b, where ARID3a inhibits differentiation, increases proliferation and inhibits apoptosis [55]. Recent studies have also identified that miRNA-30b/c/d can regulate key factors of PCD (plasma cell differentiation) such as ARID3A. This miRNA is aberrantly over-expressed in multiple myeloma (MM) tumor plasma cells, suggesting that MM cells frequently acquired expression changes in miRNA that promote dynamic modulation of expression during normal PCD [56].
As mentioned previously, MM shows a high expression of Deptor which is probably regulated by the effect of these miRNAs on the regulation of ARD13A, who could regulate Deptor in MM.
CDX1: CDX1 is another of the factors identified by our analysis. Interestingly, recent studies indicate that the Wnt/β-Catenin/signaling pathway can regulate CDX1 through CKIP-1 in gastric cancer [57] and as we mentioned previously, Deptor can be regulated by the signaling pathway of Wnt/β-Catenin/ by c-Myc. Therefore, this pathway may also be involved in CDX1-mediated regulation of Deptor from which at least two possible binding sites are predicted in the Deptor promoter (Figure 3 and Table 1). Studies have shown that CDX binding sites are required to mediate Oct-2 ability to activate bcl-2 in lymphomas impacting cell survival [58]. In addition to the role of CDX in hematopoiesis, it has been described that CDX2 can contribute to oncogenesis since data indicates that CDX expression has a functional role in leukemia and malignant blood disease [59].
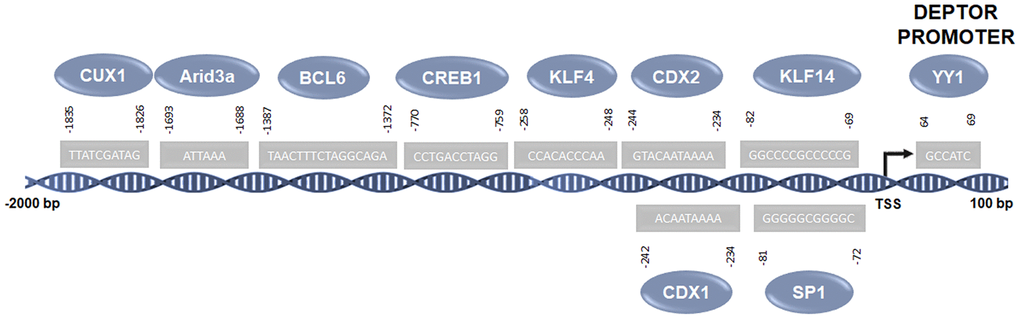
Figure 3. Predicted transcription factors involved in the regulation of Deptor. Predicted transcription factors involved in the Deptor regulation according of bioinformatic analysis using the Jaspar platform (http://jaspar.genereg.net) [52].
CREB1: CREB1 is a transcription factor member of the leucine zipper DNA binding protein family which can be activated by the TGF-β signaling pathway [60] which is linked to the regulation of Deptor expression mediated by degradation induced by the mTOR complex. However, a possible transcriptional regulation is suggested to be probably mediated by transcription factors regulated by this pathway. The bioinformatic analysis carried out by our group suggests that CREB1 can regulate Deptor transcription showing two possible binding sites in its promoter (Figure 3 and Table 1). Studies indicate that CREB has a critical role in the proliferation, survival and apoptosis of PBC-ALL cells, suggesting an oncogenic role of CREB in both ALL and CML [61, 62]. Immunohistochemical study shows an important expression of pCREB in DLBCL tissue, suggesting that high levels of pCREB could be associated with tumorigenesis in lymphoma [63]. The presence of CREB has also been associated with resistance to Lenalidomide in MM [64].
KLF4: KLF4 is a transcription factor attributed to bifunctional roles in cancer, as a potent tumor suppressor or as an oncogene [65]. Recent studies have shown that KLF4 can transcriptionally regulate Deptor expression [49]. Interestingly, our bioinformatic study reveals the presence of at least one consensus site for the binding of KLF4 in the Deptor promoter (-82) (Figure 3 and Table 1). KLF4 shows a bifunctional role in hematological malignancies, even though we and other authors have shown an oncogenic role in MM, pediatric lymphoma, and NHL cell lines [66]. Therefore, the regulation of Deptor in NHL and MM could be regulated by KLF4, contributing to pathogenesis at least in MM.
KLF14: KLF14, like KLF4, is a member of the recently identified KLF family of factors and of interest for its involvement in the field of tumorigenesis and immune regulation [67]. KLF14 expression is induced by TGF-β in intrauterine and ectodermal tissue. We identified three consensus binding sites for KLF14 in the Deptor promoter region. Thus, the known TGF-β signaling pathway mediates Deptor regulation, possibly through TGF-β-regulation of KLF14. Preliminary results by our group suggest that KLF14 plays a tumor suppressing role in lymphoma, since a low expression of KLF14 is observed in the more aggressive lymphoma phenotypes.
YY1: YY1 is a zinc finger protein, which regulates numerous genes involved in cell death, cell cycle, cell metabolism, and the inflammatory response. YY1 is highly expressed in many types of cancer which is associated with cell proliferation, survival and metabolic reprogramming [68]. In our bioinformatic analysis we identified at least one YY1 binding site in the Deptor promoter. In addition, studies in lymphoma have demonstrated the transcriptional regulation of KLF4 by YY1 [69]. Therefore, YY1 probably participates in the regulation of Deptor directly on its promoter or through the regulation of KLF4 or c-Myc, which as mentioned above, can also regulate the expression of Deptor. YY1 has been reported to play an important role in several biological processes, including the development and function of B cells [70]. Different studies have shown that YY1 is expressed in acute myeloid leukemia, B-NHL cell lines, and DLBCL tissues [71]. YY1 overexpression has also been reported to correlate with disease progression in MM and childhood acute lymphocytic leukemia [72]. Therefore, YY1 may be playing a role in these hematological malignancies either by direct regulation of Deptor, or by regulating KLF4 and/or c-Myc (Figure 2 and Table 1).
BCL6: The B-cell lymphoma 6 gene (BCL6) encodes a transcription factor belonging to the family Broad-Complex, Tramtrack and Bric-a-brac/Pox and Zinc Finger (BTB/POZ) [73] which is critical for the initiation and maintenance of GC [74, 75]. The Bcl-6 protein is capable of regulating more than 1000 target genes [76]. In normal GCB cells, Bcl-6 also suppresses the expression of the anti-apoptotic oncogene Bcl-2 by binding to its promoter region. However, in DLBCL and FL, Bcl-6-mediated deletion of Bcl-2 is often lost due to Bcl-2 translocation and Miz1 dysregulation [77]. Furthermore, although Bcl-6 is also expressed in Burkitt lymphoma, its role has not been investigated, yet it is expressed in all cases and is likely to contribute to proliferation and survival, since the BCL6 gene has long been recognized as an important oncogene in B-cell lymphoma [78, 79]. Recent studies have identified that certain subpopulations of DLBCL and BL contain MYC, Bcl-2 and/or Bcl-6 translocations and were called “double hit” lymphoma (DHL) or “triple hit lymphoma” (THL) [80]. In the most recent WHO review of lymphoma classification, the DHL/THL category is now recognized as "high-grade B-cell lymphoma (HGBL) with rearrangements of MYC and Bcl-2 and/or Bcl-6. DHL and THL have an aggressive clinical presentation and are difficult to treat with conventional chemotherapy [81, 82]
Interestingly, as discussed below, a high expression of Deptor in BL can be observed, which correlates with a high expression of Bcl-6. Because of this, it would be important to know the possible relation of regulation of Deptor by Bcl-6 in BL.
Since Multiple Myeloma is characterized by clonal accumulation of malignant plasma cells (PC) in the bone marrow, which secrete a monoclonal immunoglobulin, it has been of great interest to know the factors involved in PC differentiation. Recent studies have revealed that alterations in the expression of IRE1, XBP1, FOXP1, PAX5 or BCL6/MTA3 can reprogram PCs to previous stages of maturation [83]. Interestingly, studies in PC have shown that Deptor inhibition changed the transcriptional program associated with PC differentiation through upregulation of PAX5 and Bcl-6, which maintains the B cell program, and downregulation of IRF4, a factor that favors the differentiation of PC [84]. This suggests that Deptor is likely to down-regulate Bcl-6 and according to our analysis in sílico (Table 1), where Bcl-6 can regulate Deptor expression, it could be a loop of regulation between Deptor and Bcl-6.
SP1: Finally, our bioinformatic analysis shows at least 4 consensus sites for SP1 binding in the Deptor promoter region. SP1 is a zinc finger-containing transcription factor and its ubiquitous expression apparently mediates the maintenance of normal and malignant biological processes such as cell growth, differentiation, angiogenesis, apoptosis, cell reprogramming, and tumorigenesis [85]. Sp1 exerts its effects on cellular genes that contain putative GC-rich Sp1 binding sites in their promoters. Therefore, it is likely that SP1 can regulate direct Deptor expression. Transcriptional factor SP1 is over-expressed in pediatric acute lymphocytic leukemia B and T, which in turn increases the expression of Bcl-3, MYC, and AATF [48]. SP1 has also been described to be involved in the autocrine-paracrine loop of IL-10 in B-NHL that plays a role in cell proliferation [86]. Recent studies have shown that Sp1 regulates the expression of IQGAP1, which is a protein that plays an important role in ERK-mediated MM cell proliferation [87]. Therefore, it would be interesting to explore the possible Sp1/Deptor interaction network in MM cell proliferation.
This bioinformatic analysis suggests the possible participation of different transcription factors in the regulation of Deptor either directly or through the activation of signaling pathways mediated by growth factors such as TGF-β. However, functional interaction studies should be performed to corroborate the role of these factors in the possible regulation of Deptor (Figure 3).
Regulation of Deptor by miRNAs
miR-2909: Different studies have reported that Deptor regulation expression can be regulated by miRNAs. miR-2909 is capable of regulating crucial genes such as KLF4, c-Myc, p53, p21, SP1 and mTORC1, all essential in the regulation of cellular processes such as cell progression, cell cycle and autophagy/apoptosis [48]. Recent studies demonstrate that the expression of miR-2909 inhibits the expression of Deptor, either by inhibiting the expression of KLF4 that, as mentioned above, positively regulates the expression of Deptor or by direct binding to consensus sites in region 3'UTR of Deptor [49] (Figure 4). As well by the possible regulation of SP1 expression, who can also regulate the expression of Deptor as mentioned above.
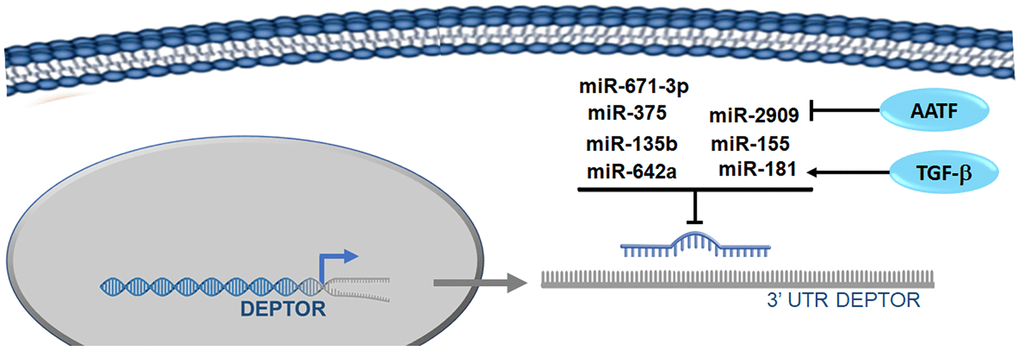
Figure 4. microRNAs involved in the regulation of Deptor. Reported microRNAs involved in the regulation of Deptor revised here are shown.
As we mentioned, miR-2909 is capable of regulating the expression of KLF4, which in pediatric ALL functions as a tumor suppressor, regulates cell cycle and apoptosis. MiR-2909-mediated down-regulation resulted in loss of KLF4 (isoform 1) activity in B-ALL as opposed to T-ALL where miR-2909 was unable to regulate KLF4 expression in T-ALL (isoform 2), due to mutations in KLF4 3'UTR, which includes the miR-2909 binding site [48]. As reviewed below, Deptor expression is higher in T-ALL than in B-ALL, so these results suggest that downregulation of KLF4 by miR-2909 may be affecting the transcriptional regulation of Deptor by KLF4 in B-ALL as reviewed above [49].
miR-155: MiR-155 is a miRNA that has been frequently studied in various types of cancer [88]. The importance of this miRNA in cancer progression is not entirely clear because in some types of cancer miR-155 exerts a tumor suppressing function, such as in the case of melanoma [89]. Recent studies have reported that miR-155 binds to the 3'-UTR region of Deptor by inducing its inhibition in breast cancer, leukemia and bladder cancer cells (Figure 4) [90–92]. Additionally, there is evidence to suggest that miR-155 acts as an "OncomiR", and is important for tumor development, such as leukemia. miR-155 is involved in various physiological processes in the cell, including hematopoiesis, immunity, inflammation and differentiation. Increased expression of miR-155 is observed in many malignant diseases, including lymphomas, acute myeloid leukemia and CLL, where miR-155 has been shown to be involved in the pathogenesis of these lymphoproliferative diseases [93]. Abnormal miR155 expression was closely associated with drug resistance to myeloma, since targeted inhibition of miR155 expression could restore sensitivity to chemotherapy by increasing FOXO3a expression in drug-resistant myeloma cells [94]. Recent studies associate miR-155 expression with increased risk of disease progression in GCB-DLBCL patients treated with R-CHOP [95]. Furthermore, recent studies report that miR-155 promote Burkitt lymphoma progression through PI3K/AKT signaling [96]. However, miR-155 is expressed at low levels in Bortezomib-resistant myeloma cells and is a direct regulator of CD47 through its 3'UTR. In addition, low miR-155 levels are associated with advanced stages of the disease, so miR-155 plays a tumor suppressor role in MM [97, 98]. This bifunctional role of miR-155 in hematological malignancies, as well as in other cancers, does not make clear the importance and relevance in the regulation of Deptor in cancer.
miR-181: miR-181 is a family of four miRNAs (a, b, c and d), which is evolutionarily conserved among vertebrates, indicating its important role in physiology. miR-181 was originally described as being highly expressed in T and B lymphocyte cell lines, although its expression is abundant in the lung, brain, and bone marrow [99]. miR-181 acts as a tumor suppressor in acute myeloid leukemia [100]. Recent studies and as previously mentioned have shown that TGF-β negatively regulates Deptor, which increases mTOR activity [101]. However, the mechanism by which Deptor is downregulated by TGF-β was not precisely known. A recent study shows that miR-181a is induced by TGF-β and this miRNA is capable of inhibiting Deptor expression by binding to the 3'UTR region of it. Demonstrating that miR-181a regulates TGFβ-stimulated mTORC1 and mTORC2 activities by direct regulation of Deptor expression (Figure 4) [102]. Recent studies show that miR-181a is overexpressed in MM cells, and may play a role in the biological function of cancer [103]. Additionally, increased levels of miR-181 have been reported in acute myeloid leukemia M1 and M2, and chronic lymphocytic leukemia (CLL) [104, 105]. While other studies in chronic B-cell lymphocytic leukemia (B-CLL) reveal that miR-181 can regulate the T-cell leukemia/lymphoma1 (TCL1) oncogene that is associated to the development of aggressive human B-CLL CD5+ [106]. While in cytogenetically normal acute myeloid leukemia (CN-AML), low expression of all miR-181 family members has been identified to have clinically favorable outcomes [107]. Thus, also in AML patients, high levels of mir-181 are expressed in the initial diagnosis compared to the first complete remission [108]. Meanwhile, in Anaplastic large cell lymphoma cells (ALK+ ALCL), a low expression of miR-181 is also observed [109].
So although it seems that miR-181 could also have a bifunctional role in hematological malignancies, its role as a tumor suppressor is dominant, which could be associated with its ability to inhibit Deptor expression, where, as we reviewed below, Deptor expression is important in MM and AML, which correlates with low miR-181 expression, while in CLL where significant miR-181 expression is observed, they have low Deptor expression. This suggests that miR-181 is probably playing some role in the regulation of Deptor in these hematological malignancies (Figure 4).
miR-375; Recent studies demonstrate that miR-375 promotes osteogenic differentiation in adipocyte-derived mesenchymal stem cells, and one of the mechanisms involved is inhibition of Deptor expression [110]. This study reports at least one consensus site in the Deptor 3'-UTR region for miR-375 binding, thus suggesting that miR-375 is capable of directly inhibiting Deptor expression. miR-375 was identified as a regulator of insulin secretion in pancreatic islets [111]. Subsequent studies revealed that miR-375 participated in multiple biological processes and, furthermore, miR-375 is significantly downregulated in different types of cancer such as multiple myeloma. Additionally, it inhibits their proliferation by inhibiting some important genes such as PDPK1, IGF1R, JAK2, and YAP1 [112–114]. Expression of miR-375 in MM has been related to myelomagenesis and subsequent progression due to its activity on oncogenes such as PDPK1 [115]. Low miR-375 expression levels predict poor outcome in AML patients as it regulates HOXB3 expression [116]. These results suggest that miR-375 in MM and AML may be playing a tumor suppressor role, interestingly in these malignancies there is a high expression of Deptor as reviewed below (Figure 4). Elevated miR-375 expression has been reported in chronic myeloid leukemia [117]. Additionally, in a study in MALT lymphoma, an elevated expression of miR-375 was demonstrated [118]. High miR-375 expression has been associated with the development and progression of pediatric AML, as it correlates with poor relapse-free survival and OS [119]. Recent studies indicate that NFKB2 may be involved in the development of Hodgkin Lymphoma by regulating miR-135a expression [120]. This suggests that miR-375 may be a potent oncomiR in MALT lymphoma, pediatric AML and HL. As reviewed further below, low Deptor expression is observed in CLL, which could be related to high expression of miR-375, while a low expression of miR-375 in MM and AML is related to a significant expression of Deptor in which miR-375 could have a tumor suppressor role through Deptor regulation (Figure 4).
miR-135b: A recent study in Multiple Myeloma reveals that Deptor expression is also controlled by two other miRNAs, miR135b and miR642a, which are down-regulated in several MM patients [121]. In other study it was shown in vitro that transfection of miR135b and miR642a decreased Deptor levels in myeloma cells. MiR135b and miR642a were observed to modulate Deptor expression through binding to consensus sites in the 3 'UTR region of Deptor [122]. Interestingly, this expression of Deptor regulated by miR135b and miR642a reversed the transcriptional program of myeloma cells and reduced cell size and tumor mass. The miR-135b has been found to play an important role in multiple life activities. For example, miR-135b inhibits the production of LPS-induced reactive oxygen species (ROS), the activation of nuclear factor-κB (NF-κB), and the mRNA expression of TNF-α in human macrophages [123]. In cancer, miR-135b acts as a double-edged factor. It promotes the development of pancreatic cancer and gastric cancer [124]. However, miR-135b can inhibit the proliferation of tumor cells in osteosarcoma [125]. Recent studies show that miR-135b regulates the expression of RBX1, significantly inhibiting the malignant behaviors of MM cells [126]. As we already mentioned, miR-135b is downregulated in MM and is associated with 14q32 rearrangement [127]. In addition, studies reveal the importance of circulating levels of miR-135b in detecting bone disease and in predicting the prognosis of patients with multiple myeloma, suggesting their possible clinical applications [128]. Studies have revealed that short-term treatment of leukemia cells with etoposide, results of transient resistance associated with the transient increase in miR-135b and miR-196b, suggesting that the increase of the expression of these miRNAs, associated with ABC1, may have a role in chemo resistance in leukemia [129]. Studies in Anaplastic Large Cell Lymphoma (ALCL) report that miR-135b is involved in anaplastic nucleophosmin-lymphoma kinase-mediated oncogenicity (NPM-ALK) and potentiates the IL-17-producing immunophenotype [130]. The NPM-ALK oncogene promotes the expression of miR-135b and its target gene LEMD1. At the same time miR-135b decreases chemosensitivity in Jurkat cells, suggesting its contribution to the oncogenic activities of NPM-ALK. In addition, another study describes that miR-135b is over-expressed in ALK+ ALCL. Thus, this association of miR135b with ALK expression may represent important downstream effectors of the ALK oncogenic pathway [131]. Interestingly, and as discussed later in the section on the role and expression of Deptor in lymphoma, ALK+ALCL vs ALK-ALCL, shows in general that there are no differences in Deptor expression. Therefore, it is not clear if miR-135b plays a relevant role in the expression of Deptor in ALK+ALCL. Additionally, the expression of miR-135b could play a critical role in the pathogenesis of gastric MALT lymphoma (GML) related to H. pylori infection [132, 133]. Therefore, in addition to the regulation of Deptor by miR-135b, it may also be associated with the regulation of KLF4 who, as previously reviewed, regulates the expression of Deptor [134].
miR-642a: There are currently very few reports regarding miR642a and its biological role. Some reports indicated that it is associated with some disease conditions including Leiomyoma, Uterine and Systemic Lupus Erythematosus [135]. As already mentioned, the low expression of miR642a has been reported in patients with MM, especially in the cytogenetic group t (14; 16) [121]. This observation suggests a possible link between the miR-642a downregulation in these MM subsets and an advantage in myeloma cell growth. Also, as previously stated, miR-642a inhibits the expression of Deptor in MM (Figure 4), which leads to the regulation of IRF4, regulating the terminal stages of differentiation [122]. miR642a was downregulated in classical Hodgkin Lymphoma compared to normal B cells, although no difference was found between stage, response to treatment, Disease Free Survival, and Overall Survival. However, there was no clinical association between clinical variables and with miR642a expression. Therefore, more studies are required to evaluate the miRNA profile and the clinical result in Hodgkin Lymphoma [136]. However, as we review below, the low expression of miR-642a in HL may be associated with high levels of Deptor expression in HL (Figure 4).
miR-671-3p: A recent study showed that miR-671-3p is capable of regulating Deptor expression in breast cancer (Figure 4) [137]. In breast cancer, it was observed that there is a significantly lower miR-671-3p expression than in normal breast epithelial cells. The forced expression of miR-671-3p induces a decrease in cell proliferation and invasion, whereas inhibition of miR-671-3p obviously promoted cell proliferation and invasion. Deptor was shown by a reporter plasmid to be a target gene for miR-671-3p and overexpression of miR-671-3p caused significant downregulation of Deptor protein expression. These findings demonstrate that miR-671-3p suppresses the proliferation and cell invasion of breast cancer by directly regulating the expression of Deptor [137].
Recent studies have reported that miR-671 plays an important role in multiple cancers [138–141]. The miR-671 precursor forms two mature miRNAs known as miR-671-5p and miR-671-3p. A recent study has shown that miR-671-3p is an antioncogene in breast cancer, where forced expression can inhibit cell proliferation and invasion and sensitize cells to chemotherapy in a Breast Cancer cell line [139]. Additionally, it was demonstrated that the tumor suppressor role of miR-671-3p in breast cancer is given through its influence on the Wnt signaling cascade [140].
However, in non-small-cell lung cancer, miR-671-3p has been shown to play an oncogenic role by inhibiting FOXP2. Therefore miR-671-3p has also been proposed as a potential therapeutic target in NSCLC [141]. Additionally, in glioma, miR-671-3p has been shown to promote cell proliferation and migration in vitro and to decrease apoptosis through CKAP4 regulation [142]. This study demonstrates that miR-671-3p is a predominant positive regulator of glioma progression, suggesting that the miR-671-3p/CKAP4 axis may serve as a possible therapeutic target or biomarker in glioma [142].
Predicted microRNAs in the regulation of Deptor
To address what other possible miRNAs may be regulating the post-transcriptional expression of Deptor, we looked for miRNAs-Deptor interaction in a database (mirTarBase platform; http://miRTarBase.cuhk.edu.cn/) [143], using a value of p=0.05 for miRNAs whose predicted binding sites in the Deptor 3’UTR region. According to the high prediction score, at least 8 miRNAs were predicted to have a binding site in the Deptor 3’UTR region and these sites were consistent in at least four other databases (miRMap, PITA, RNA22 and RNAhybrid). From these, we revised the following: miR-190a-3p, miR-4796-5p, miR-654-3p, miR-5011-5p, miR-1277-5p. miR-4803, miR-384 and miR-375 (Table 2). These miRNAs were validated by the Next-Generation Sequencing (NGS) method on at least one study, so it is necessary to expand the experimental evidence of these miRNAs.
Table 2. Predicted microRNAs involved in the regulation of Deptor.
| MIcroRNA | Position in DEPTOR 3'UTR | Score (miRTarBase) | Pairing of microRNAs and 3’ UTR Deptor | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| miR-375 | 5480 - 5498 | 125 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| miR-190 | 1275 - 1299 | 164 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| miR-4796-5p | 4289 - 4311 | 152 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| miR-654-3p | 4161 - 4183 | 143 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| miR-5011-5p | 1474 - 1493 | 155 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| miR-1277-5p | 1302 - 1325 | 194 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| miR-4803 | 4186 - 4206 | 135 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| miR-384 | 2436 - 2455 | 134 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Using the mirTarbase database, we found a potential binding sites of microRNAs in the 3’UTR of Deptor. Here we show the site of miRNA binding, with the highest score in accordance with mirTarbase. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Very little is currently known about miR-190a-3p. Recent studies demonstrate that miR-190a-3p regulates glioblastoma tumorigenesis by regulating PTEN [144]. The possible binding site of miR-190a-3p in Deptor mRNA is shown in Figure 5. The biological role of miR-4796-5p is currently unknown.
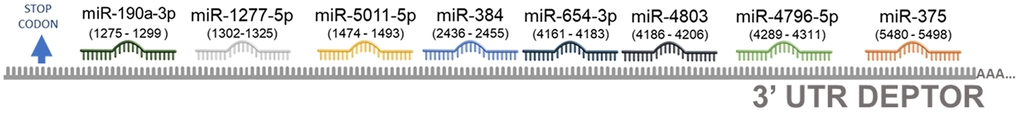
Figure 5. Predicted miRNAs involved in the regulation of Deptor. Schematic representation of predicted miRNAS involved in the Deptor regulation using the database (mirTarBase platform; http://miRTarBase.cuhk.edu.cn/) [143]. A value of p=0.05 for miRNAs whose predicted binding sites in the Deptor 3’UTR region was considerate.
Low expression of miR-654-3p has been demonstrated in natural extranodal killer/T-cell lymphoma NKTCL [145]. Recent studies reveal that the expression of miR-654-3p in hepatocellular carcinoma (HCC) is low and is associated with metastasis and is a predictor of poor prognosis. Overexpression of this inhibits the proliferation, migration and invasion of HCC, which proposes miR-654-3p as a tumor suppressor [146]. However, in colon cancer it is observed as an oncomiR, since its high expression correlates with poor survival in these patients [147]. Its role in hematologic malignancies is currently unknown with precision. The role of miR-5011-5p in biological functions of homeostasis or cancer is currently unknown. Studies in Parkinson's disease reveal that miR-1277-5p overexpression correlates with cell viability and inhibition of apoptosis in human neuroblastoma cells [148]. Our bioinformatic analysis predicts that miR-1277-5p has Deptor as a potential target gene (Figure 5). Like other miRNAs identified in our bioinformatic analysis that may have Deptor as target gene, the biological role of miR-4803, in cancer is unknown. Although the functional role of miR-384 in hematologic malignancies is unknown to date, previous studies have shown that miR-384 was abnormally expressed in various tumors. miR-384 can inhibit the proliferation and invasion of tumor cells in tumor tissues such as gastric cancer, colon cancer and liver cancer and in osteosarcoma [149–151]. Furthermore, miR-384 is involved in the phenotypic properties of cancer cells, such as cell proliferation, apoptosis, and the cell cycle, and plays a vital role in the expression of gene products [152]. These studies suggest a potential role for miR-384 as a tumor suppressor. Our bioinformatic analysis suggested that Deptor could be a possible target gene for miR-384 (Figure 5). Interestingly, our bioinformatic analysis shows the miR-375 which, as already mentioned, is involved in the regulation of Deptor [110].
Role of Deptor in hematological malignances
Deptor in Multiple Myeloma
Multiple Myeloma (MM) is a hematologic disorder which is characterized by a proliferation of malignant monoclonal plasma cells in the bone marrow (BM) and/or extramodular sites. Progress in the first PFS and overall survival (OS) has been achieved through the introduction of high-dose therapy (HDT) with autologous stem cell transplantation (SCT), and by the introduction of thalidomide, bortezomib and lenalidomide. Despite the recent progress in OS rates, MM remains an incurable disease and the majority of patients will relapse and will require treatment [153]. Recently studies have been shown that Deptor can be implicated in the chemosensitivity to melphalan in vitro. Deptor knockdown was associated with melphalan-induced growth inhibition due to an induction of apoptosis associated with the inhibition of p-AKT protein level, probably due to the fact that Deptor acts as an oncogene, compensating for the inhibition of feedback from p-P70S6K to PI3K, thus activating to AKT [154]. It is known that the inhibition of Deptor results in the inhibition of proliferation and induction of apoptosis in myeloma cells. In addition, high levels of Deptor protein are predictive of a poor response to thalidomide in myeloma, which indicates that the levels of Deptor expression is important as a prognosis marker for myeloma patients and a possible target for a new therapeutic strategy as a small chemicals inhibitor alone or in combination with conventional treatments [155].
Deptor is a 48 kDa protein that binds to mTOR and inhibits this kinase in TORC1 and TORC2 complexes specifically in MM [156]. Different reports have associated Deptor with different pathways and mechanisms in myeloma. Interestingly, Deptor expression is required to maintain myeloma cell differentiation and that high level of its expression are associated with better outcome [122]. Two GEO repository were analyzed in a study [121, 157], finding that the PFS was significantly longer in MM patients with high expression levels of Deptor that in those with low Deptor expression levels (p<0.05) [122].
In another study, the silencing of Deptor with a new drug prevents the Deptor-mTOR binding, promoting the activation of MTORC1 and MTORC2 and induction of cytotoxicity, suggesting that Deptor is a potential therapeutic target and implicating the critical role of Deptor and mTORC1 in MM [156]. Additionally, Deptor plays a crucial role in the proliferation pathway. This was related to p21 because the knockdown of Deptor induces p21 expression independent of p53, and p21 knockdown prevents the cytotoxic effects of Deptor silencing [158]. According to this the silencing of Deptor with a shRNA resulted in an inhibition of proliferation an increase in cleaved caspase 3 and PARP. This increase was related to apoptosis induction, chemo sensitization, suppression of autophagy [159], melphalan chemo sensitization effects, and reduced levels of phosphor-AKT. These factors caused an inhibition of PI3K-AKT pathway [154]. In this context, several drugs have been proposed for a theorical treatment, such as Thalidomide. This immunomodulating agent can increase PTEN expression and suppress AKT signaling, which is fundamental for MM development and maintaining [155].
Some authors have proposed Deptor as a tumor suppressor in MM, the author suggest that this due to its regulation on mTORC2, which participates in regulation of NDRG1 through the activation of SGK1, which confers chemo resistance to Bortezomib [160]. Since the decrease in SGK1 reduces the phosphorylation of NDRG1 and promotes the induction of apoptosis or chemo sensitivity to Bortezomib. This suggests that depending on the activation pathways mediated by mTORC2; PI3K/PDK1 or PI3K/AKT and the regulation of mTORC2 by Deptor will depend on the role of Deptor as an oncogene or tumor suppressor [12]. Additionally, low levels of Deptor has been found in several cancers [1]. In conclusion, the information above presented support a possible role as tumor suppressor.
A study analyzed if Deptor is capable of regulating different signaling pathways, cells transfected with siRNA-Deptor were analyzed by a protein/kinases microarray. The results showed that Deptor effectively regulates the activity of mTORC1, but interestingly shows that Deptor affects the phosphorylation and activation of ERK1/2 as well as STAT1, p38α, Paxillin, PLCγ-1 and FAK, but each to a lesser degree than ERK1/2.
In that same study, it was analyzed whether the regulation of these kinases was associated with each other, results showing that Deptor inhibition increases ERK1/2 levels independently of the mTOR pathway. Therefore, these findings indicate that Deptor selectively regulates the endogenous activity of ERK1/2 and mTORC1 through independent mechanisms. Deptor has been suggested to show independent regulatory effects on MAPK signaling pathways [161], which is consistent with reports suggesting that it contains putative ERK1/2 binding and association sites [20, 162]. Therefore, the phosphorylation and degradation mechanisms of Deptor reported mediated by the "degron" domain [20, 162] may be a regulatory mechanism in biological processes where Deptor plays a role.
Interestingly, and according to the characteristics of MM of higher secretion of immunoglobulins, the participation and homeostasis of endoplasmic reticulum is critical for the maintenance of MM. Deptor has been identified as a protein able to bind to DNA and woks by regulating the transcription of several genes involved in the maintenance of the endoplasmic reticulum such as ERLIN2, KEAP1, PSEN2 and DERL3 [33]. Since it was demonstrated by chromatin immunoprecipitation (ChIP) analysis, that Deptor is capable of binding DNA specifically to the promoter region of these genes.
This topic is still unclear because some reports identified no association with activation of the unfolded protein response after MM cytotoxicity was induced by the silencing of Deptor with drugs. However, Deptor depletion with siRNAs can induce endoplasmic reticulum stress and synergizes the effect of the proteasome inhibitor bortezomib in MM cells [33]. This could be explained as the silencing of Deptor with shRNA produces a feedback downregulation of the IGF/IRS-1/PI3-K/PDK1 pathway [163]. In our group, a set of small chemical inhibitors for Deptor was developed in which they inhibited Deptor/mTOR binding. Therefore, mTORC1 is up-regulated, promoting feedback down-regulation of the IGF/IRS-1/PI3-K/PDK1 pathway, and activating AKT phosphorylation. On the other hand, elimination of AKT decreases activation, and this effect probably results in increased mTORC2 activity which overcomes any inhibitory effect on the PI3-K/PDK1 pathway [163]. These compounds activate mTORC1 and selectively induce MM cell apoptosis and cell cycle arrest [163].
Leucine-rich repeat containing 4 (LRRC4), also called netrin-G ligand-2, is a member of the leucine-rich repeat (LRR) superfamily [164]. Previous studies have confirmed that LRRC4 plays a central role in the development and early differentiation of the nervous system, especially during synapse formation [165]. Furthermore, a new role for LRRC4 as a tumor suppressor for glioma has been reported [166].
Recent studies have shown that LRRC4, an autophagy inhibitor, interacts directly with Deptor, which induces a decrease in the level of Deptor protein. This interaction was determined by co-immunoprecipitation assays for which plasmids with different domains of Deptor were constructed, finding an interaction of LRRC4 with the PDZ region (330-407). Additionally, the deletion of the C-terminal binding domain resulted in an abolition of the interaction of Deptor with LRRC4. This resulted in the activation of mTOR, thus decreasing the level of cellular autophagy [167]. Therefore, it has been suggested that LRRC4 could be used as a possible therapeutic strategy in malignancies associated with high expression of Deptor.
In addition, natural compounds have been proposed as a potential therapeutic treatment for MM; for example, cell lines treated with 1,2,3,4,6-Penta-O-galloyl-beta-D-glucopyranoside (PGG) show a cell cycle arrest in G1 phase and increased of apoptosis by caspase 3 cleaved expression. Interestingly, the treatment with PGG in cells was closely associated with a decreased effect on Deptor expression in a dose-dependent way in MM. In contrast with other compounds, this antagonizes the effect of bortezomib [168].
It was reported a Deptor was increased in a time dependent way at 0, 6, 12 and 18 h, and then decreased after nutrient depletion in 8226 cells. These levels were consistent with autophagy [169], confirming the participation of Deptor in apoptosis and provides a possible participation in autophagy.
The potential biological roll of Deptor in MM was predicted by our analysis, using bioinformatic tools (ONCOMINE), and the expression of Deptor was analyzed to be significantly upregulated in MM cells (P<0.001). When analyzing different studies related to the expression of Deptor in MM, it is typical to find a high expression of Deptor. In a study involving 133 MM samples, moderate expression was observed compared to plasma cells. In this same study, an analysis was made to observe if there were differences between the expression of Deptor and the stage of the disease and finding that there are no statistically significant differences [170]. Another study shows a significant expression of Deptor in MM (12 samples) compared to bone marrow cells (Figure 6A) [171].
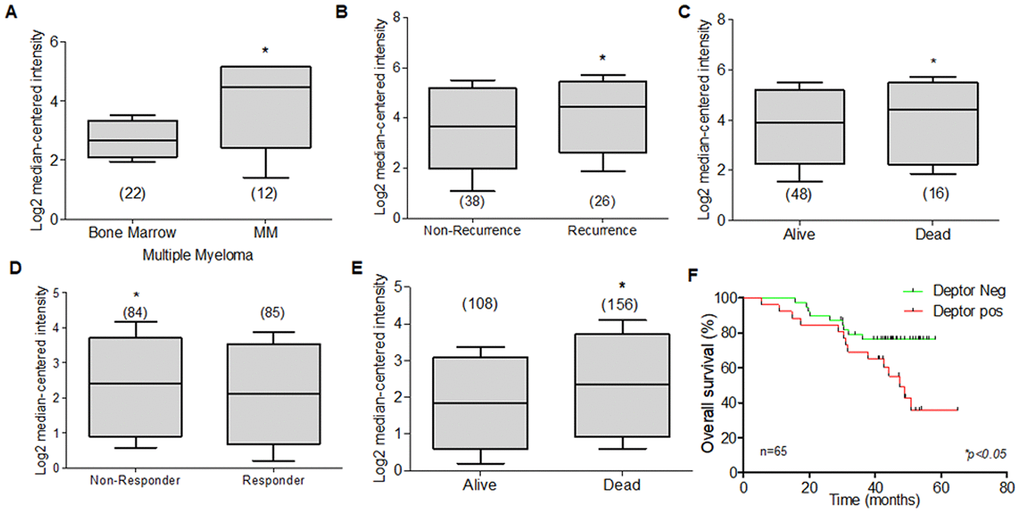
Figure 6. Expression of Deptor in Multiple Myeloma. Oncomine Deptor expression was revised in different reports; (A) Zhan et al, analysis of Deptor Expression. MM shows a higher expression compared to bone marrow cells (*p<0.001). (B) In Carrasco et al, Analysis of Deptor, relationship with disease recurrence and Deptor expression was observed with high Deptor expression compared to non-recurrence (*p<0.05). (C) in Carrasco et al Analysis, we found a moderate high expression of Deptor in Alive patients compared to Dead patients (*p<0.05). (D) in Mullygan et al, analysis shown a responder patient has a moderated high expression of Deptor compared to Non-responders (*p<0.05). (E) In the same data analysis overall survival was shown, and dead patients have an important high expression of Deptor (*p<0.001). (F) Overall survival of patients with MM according to Deptor expression. Number of patients (n) is listed next to the graph (*p<0.05).
A significant expression of Deptor was observed compared to leukocytes or bone marrow cells in a study that included 84 samples [172]. Studies presented by Chapman that included 289 MM samples [173], Broyl with 320 samples [174], Dickens with 247 samples [172], and finally Zhou with 115 samples [175] showed significant Deptor expression. Interestingly, in two studies that included 65 and 264 samples, respectively, it is observed that there is a moderate relationship with Deptor expression and Progression-free survival (PFS) at 1 year, or with OS (Figure 6B, 6C). In the first study, a relationship with disease recurrence and Deptor expression was observed (Figure 6B). In the second, a relationship with poor response to therapy (Bortezomib) was also observed (Figure 6D) [176, 177]. In a study that included 414 samples, an important Deptor expression is also observed, but it is not possible to establish a significant difference in the relationship with Deptor expression and OS [171]. Another study observed a lower expression in the samples from alive patients of MM compared with a high expression in samples from dead myeloma patients (Figure 6E) [176]. Finally, in the same study using the overall survival, we perform a Kaplan Meier analysis. We compared the overall survival of patients with high and low Deptor expression and observed that high Deptor expression correlates with a lower percent survival (*p<0.05). (Figure 6F) [176].
All these results together strongly suggest that Deptor has a tumor promoting role (oncogene) in MM, where this oncogenic role of Deptor is very specific and important in MM. Additional studies are necessary to establish more clearly the role of Deptor in MM, however, important studies have currently been carried out using Deptor with an important therapeutic target with very successful results that to date have led to pre-clinical studies [18].
Deptor in Leukemia
Another hematological malignancy in which Deptor plays an important role is Leukemia. Especially in patients with T-ALL, where the role of aberrant NOTCH1 is crucial in the pathogenesis [178]. NOTCH1 promotes leukemogenesis in cooperation with AKT [179]. In this case, NOTCH1 promotes Deptor by union in his promoter identifying a transcriptional control of Deptor and its regulation of AKT, proliferation, and leukemogenesis in T-ALL [40].
In AML, Deptor has been related to KDM4A specifically Depletion of KDM4A decreases [180]. KDM4A is an αKG-dependent enzyme of the Jumonji family of lysine demethylases [181]. KDM4A overexpression reduces the ubiquitination of Deptor while 2HG-induced KDM4A inhibition promotes mTOR activation and act with Deptor. Inhibition of KDM4A results in reduction of Deptor levels, leading to mTOR activation independently of the PI3K/AKT/TSC1-2 pathway [40]. In promyelocytic leukemia cells, the cullin-RING ligase (CRL)-NEDD8 pathway maintains essential cellular processes. The treatment with the antineoplastic ATRA promotes an up-regulated protein expression of Deptor which downregulates mTOR and could promotes the differentiation of promyelocytic cells [182].
Currently there is very little information on Deptor expression in leukemia and its biological role. However, we reviewed through Oncomine platform database, studies that included the analysis of Deptor mRNA expression in some types of leukemia. Through this analysis, we found that studies in samples of patients from Acute Myeloid Leukemia (AML) with 542 samples [183], another 23 [184], and 285 samples [185] had a significant expression of Deptor compared to peripheral blood cells or bone marrow cells respectively (Figure 7A–7C). Another study that included 43 samples of AML showed an important expression of Deptor [186]. Another important expression of Deptor is observed in another study that included 162 AML samples, in which analyzing the PFS at 3 years and the OS shows that there is no significant difference between these and the Deptor expression [187]. However, in another study that included 78 AML samples, where important Deptor expression is also shown, it is observed that Deptor expression is related to poor PFS at 3 years and total survival (Figure 7D, 7E) [187].
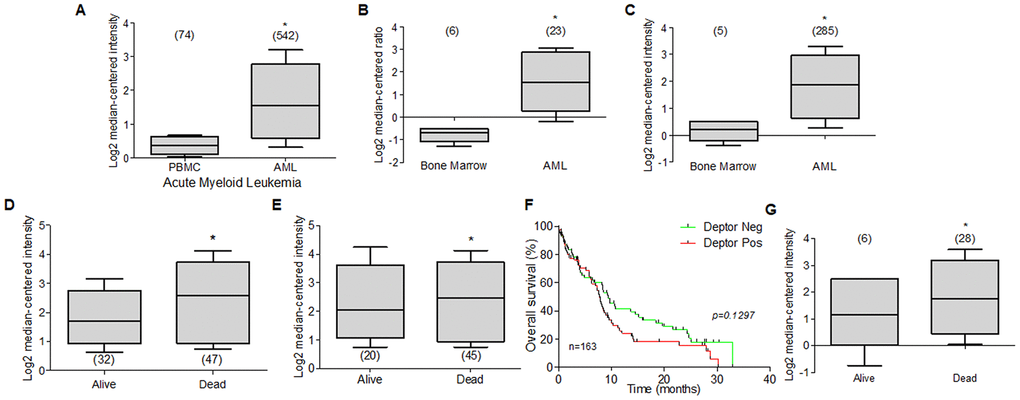
Figure 7. Deptor expression in Acute Myeloid Leukemia. Bioinformatic analysis with Oncomine of Deptor expression on AML was revised. (A–C) three different studies by Hafferlach et al, Andersson et al, and Valk et al, were revised and respectively shows an important higher expression of Deptor in patients with AML compared to controls (PBMC and bone marrow) (*p<0.001). (D–G) Metzeler et al, and Raponi et al, analysis shown overall survival status on relationship with Deptor expression (D, G), dead patients have an important high expression of Deptor (*p<0.001). and Progression free survival (E), shows a moderate relationship with high expression of Deptor in dead patients (*p>0.05). (F) Overall survival of patients with AML according to Deptor expression. Number of patients (n) is listed next to the graph (n/s p>0.05).
In the same study, we perform a Kaplan Meier analysis to compare the Deptor expression over percent survival of patients. We observe that high Deptor expression was found in patients with poor survival. (Figure 7F) [187].
Another study reported by Heuser and Co that includes 33 AML samples, a relatively low expression of Deptor is observed. However, when analyzing the response to treatment, a poor response is observed in the group of samples with higher Deptor expression (p =0.001), although it is not possible to establish whether there is a relationship between PFS 3-year or OS [188]. In other two studies carried out by Raponi and Co. they show an important Deptor expression in 58 AML samples and other with 34. Interestingly, in the last one, a poor OS is observed in relation to higher Deptor expression (Figure 7G) [189, 190].
In a study in Childhood AML (237 samples), a significant expression of Deptor is shown [191]. However, in another study that included 248 Adult AML samples and 93 Childhood AML samples, a negative expression of Deptor was observed [192]. Therefore, the expression of Deptor in childhood AML is not as clear as the case of AML where an important expression of Deptor is consist in the different studies reviewed here. Acute Myeloid Leukemia (AML) is a hematologic cancer characterized by abnormal clonal proliferation of undifferentiated blasts and by their primary infiltration of hematopoietic organs, such as the bone marrow (BM), lymph nodes, and spleen. Uncontrolled expansion of leukemic cells suppresses normal hematopoiesis [193]. Conventional AML treatment and allogeneic transplantation are the current therapeutic alternatives [193]. However, relapse and disease progression remain the main causes of treatment failure [194]. Thus, treatment of AML remains an important unmet clinical need, especially for patients lacking mutations. According to this review and the important expression of Deptor in AML, this may be of importance in studies as a potential therapeutic target, as in MM (Table 3).
Table 3. Expression of Deptor in hematological malignancies.
| Malignancy | Subtype | Deptor expression | Reference | |
| Multiple Myeloma | High | [170], [171], [172], [173], [174], [175], | ||
| Leukemia | AML | High | [183], [184], [185], [186], [187], [189] [190] [191] | |
| T- ALL | High | [184], [195], [196] | ||
| B-ALL | Low | [184] [183, 195, 196] | ||
| CML | Low | [183] | ||
| CLL | Low | [183] | ||
| Non-Hodgkin Lymphoma | DLBCL | High | [208], [209] | |
| Low | [210–212] | |||
| ABC | High | [208, 217, 218] | ||
| GCB | High | [208, 217, 218] | ||
| FL | Low | [211, 220] | ||
| High | [208] | |||
| BL | High | [209, 211] | ||
| MCL | High | [224] | ||
| MZL | Low | [212] | ||
| ALCL | High | [228] | ||
| ALK- | High | [227] | ||
| ALK+ | High | [227] | ||
| Hodgkin Lymphoma | cHL | High | [234] | |
When we reviewed the reports that show data in T-Cell Acute Lymphoblastic Leukemia (T-ALL), it is observed in several reports that the expression of Deptor is consistent in this malignancy; for example, in a study with 174 samples, a moderate expression is observed compared to peripheral blood cells [183]. In another report that included 11 T-ALL samples, a high expression of Deptor is observed compared to healthy bone marrow cells [184]. Similarly, in another study in T-ALL with 23 samples, an important expression of Deptor was observed [195]. In other studies where the Deptor expression in T-Cell childhood Acute Lymphoblastic Leukemia was reviewed, it is observed that there is also an important expression of Deptor in a report that included 46 samples, compared to normal cells or B-ALL [196] (Figure 8A). In this study, the minimum residual disease was reported at 46 days and moderate inverse relationship is shown between it and Deptor expression (p=0.05) (Figure 8B). Likewise, in another study in 10 samples of T-Cell Childhood Acute Lymphoblastic Leukemia it is observed that the Deptor expression is related to the recurrence of the disease (Figure 8C, 8D) [197]. T-ALL is a highly proliferative hematologic malignancy caused by malignant transformation of T-cell progenitors [198]. T-ALL patients generally present aggressive clinical features correlated with a poor prognosis, including inhibition of normal hematopoietic function [199]. The complete remission rate of T-ALL can reach 94%, and the long-term survival rate can reach 85% [200]. However, 20% of pediatric patients and 40% of adult patients are susceptible to recurrence and develop refractory leukemia [199]. Because of this, understanding the underlying mechanisms of T-ALL development is highly relevant. As already mentioned, a different study indicates that Deptor is importantly expressed in T-ALL and some mechanisms have been described that implicate its role in the development of this condition as an oncogene where it promotes the activity of Notch [40] and activation of the PI3K/AKT pathway. Therefore, all these results suggest Deptor with an important role in the pathophysiology of T-ALL and of interest as an important therapeutic target in T-ALL (Table 3)
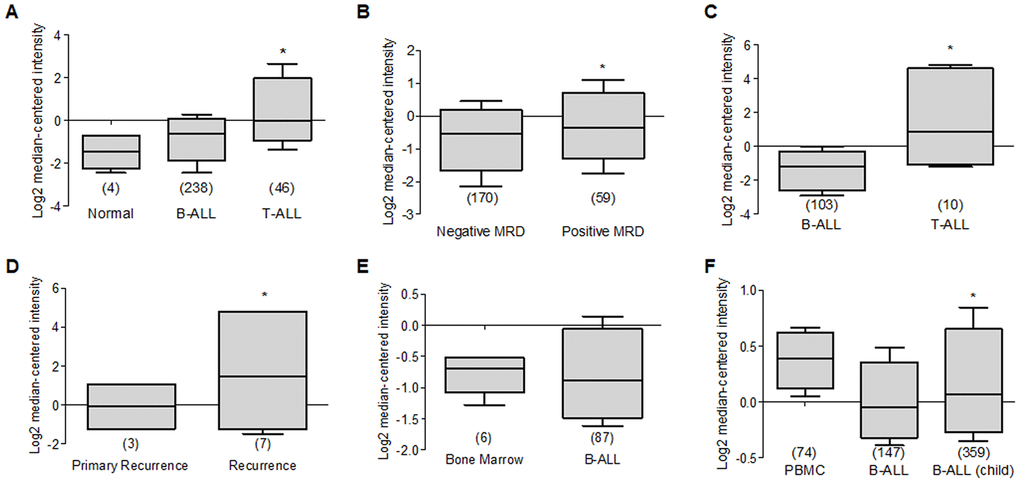
Figure 8. Deptor expression in Acute Lymphocytic Leukemia. Oncomine analysis of Deptor expression in ALL was revised. (A) in Coustan Smith et al, expression analysis shows a higher expression in T-ALL vs normal cells and B-ALL (*p=0.05). (B) analysis of minimal residual disease in relationship with Deptor expression, we observed moderate high Deptor expression in patients with positive MRD (*p>0.05). (C) Another study presents by Bhojwani et al, a significant higher Deptor expression was observed in T-ALL compared to B-ALL (*p<0.001), (D) In the same study the Deptor expression was related to recurrence of the disease compared with primary recurrence. Moderate high Deptor expression was related with recurrence (*p<0.05). (E) On the other study presented by Andersson et al, Deptor expression is low in B-ALL compared with Bone Marrow (*p= N/S). (F) On Haferlach et al. study, very low Deptor expression in B-ALL and in B-ALL childhood were observed compared with PBMC (*p= N/S).
Contrary to what was observed in T-ALL, when we analyzed Deptor expression in B-Cell Acute Lymphoblastic Leukemia (B-ALL), a decreased or negative expression is shown. Thus, in a study that included 147 samples of B-ALL, a moderate expression was observed when compared to peripheral blood cells [183]. Consistent with another study that included 87 samples of B-ALL, Deptor expression is very low (Figure 8E) [184]. This low Deptor expression is consistent in Pro-B Acute Lymphoblastic Leukemia (70 samples) [183]. When Deptor expression was reviewed in B-Cell Childhood Acute Lymphoblastic Leukemia in studies that included 359 samples, negative Deptor expression was also observed (Figure 8F) [183]. As already mentioned, ALL is a hematological neoplasm caused by the proliferation and accumulation of lymphoid progenitor cells in the bone marrow or in extramedullary sites, which can be presented as phenotypic subgroups of B cells and T cells. However, more than two thirds of all adult cases are B cell phenotypes [201]. B-ALL has a more favorable survival than T-ALL in a young population and the opposite occurs in older ages [201]. Interestingly, our analysis generally shows significant expression of Deptor in T-ALL and not in those of the B-ALL phenotype, suggesting that Deptor may have a more potential role as an oncogene in T-ALL than in B-ALL. Additional studies are necessary to determine the possible role of Deptor in B-ALL, which according to our review, could play a role as a tumor suppressor much like in other types of cancers, such as breast cancer (reviewed by Caron) [4].
Additional studies also show a negative expression of Deptor in Chronic Myelogenous Leukemia (76 samples), Myelodysplastic Syndrome (206 samples), Chronic Lymphocytic Leukemia (448 samples) [183], while observing a positive expression in Acute Promyelocytic Leukemia (12 samples) [202]. Chronic lymphocytic leukemia (CLL) is one of the most common types of leukemia in western countries [203]. CLL is mainly characterized by the accumulation of mature monoclonal CD5+ B cells in lymphoid tissues and peripheral blood [204]. Despite significant advances in its treatment, CLL remains incurable [205]. Therefore, our observations in the analysis of Deptor expression in acute and chronic hematological diseases suggest that Deptor expression is not observed in mature (chronic) phenotype leukemias, but in T-cell lymphoid (acute) leukemias and not so in B cells (Table 3).
Recently, it was revealed that mTOR plays critical roles in regulating CD4+T cell activation and proliferation [206]. In addition, Deptor as part of the mTOR signaling complex, may be playing a role in the activation, proliferation and even differentiation of CD4+T cells [207]. Deptor has been found to be expressed at high levels in non-activated CD4+T cells and its expression regulates the activation state of CD4+T cells. Low levels of Deptor are associated with an optimal mTOR activity with a differentiation of CD4+T cells [207], while the high expression of Deptor in CD4+T cells modulates the differentiation to effector cells and favors Foxp3 T cells. Thus, the mTOR pathway is involved in the differentiation and activation of CD4+ T cells and Deptor could be involved in the biological regulation of CD4+T cells and their final destination after activation [206]. Therefore, it is not surprising that Deptor is involved in the pathophysiology of hematologic malignancies of T cells.
In conclusion, and according to the studies reviewed here, an important expression of Deptor is observed in T-ALL, as well as in T-Cell Childhood Acute Lymphoblastic Leukemia, while Deptor expression is very low or negative in B-ALL and in B-Cell Childhood Acute Lymphoblastic Leukemia as well as in CLL (Table 3).
Deptor in Non-Hodgkin Lymphoma
Since there is also very few or no information about Deptor expression in lymphoma and its possible biological role, through Oncomine platform we analyzed different works that contain the mRNA Deptor expression database, finding the following result: A study by Compagno and Co. shows high Deptor expression in Diffuse Large B-Cell Lymphoma (DLBCL) (44 samples) (Figure 9A) [208], which is consistent with another study that included 166 samples [209]. While in other studies, Jais and Co. with 53 samples, Bruce and Co., and Storz and Co. with 6 samples respectively, a negative expression is observed [210–212]. This level of expression of Deptor is important but lower when compared to MM, where it is observed that there is a tendency for greater expression [213].
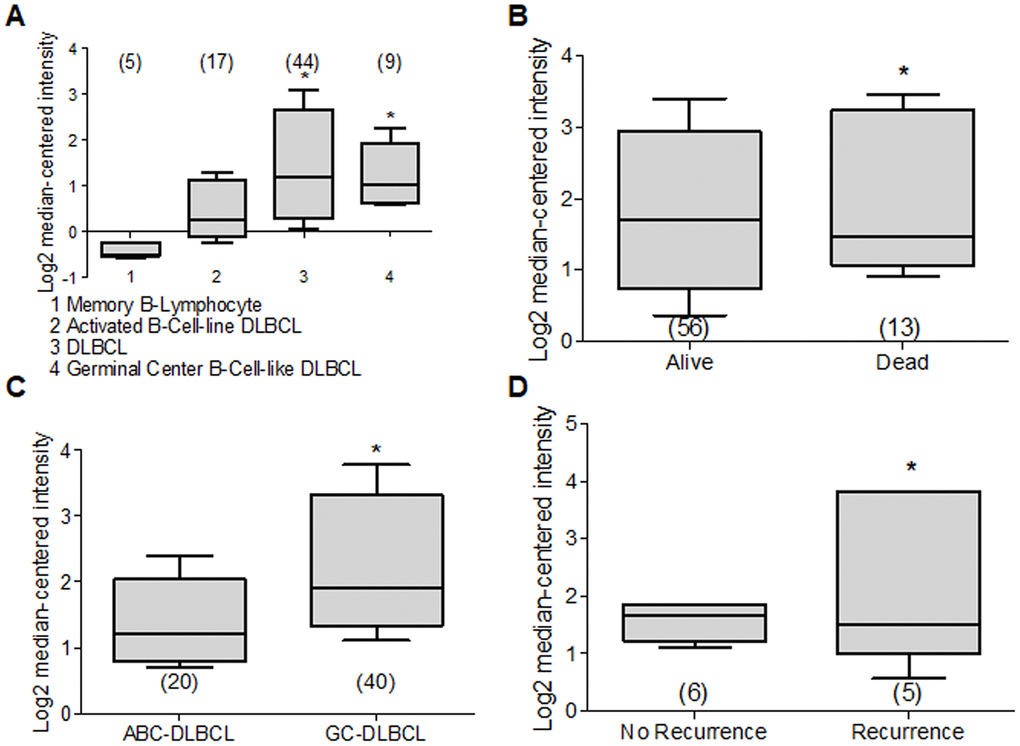
Figure 9. Deptor expression in Lymphoma. Based on the Oncomine Analysis we revised Deptor expression in different subtypes of DLBCL lymphoma. (A) in Compagno et al. analysis, we observe a differential expression of Deptor in different subtypes of DLBCL. DLBCL, GCB-DLBCL and ABC-DLBCL, shows higher Deptor expression compared with memory B-cell (*p<0.001). (B) In Shaknovich et al, Analysis, overall survival according Deptor expression was revised, alive patients show slight high Deptor expression compared with dead patients (*p=N/S). (C) in the same analysis two subtypes of DLBCL were revised; ABC-DLBCL vs GC-DLBCL shows a higher Deptor expression in GC-DLBCL (*p<0.05). (D) in the same study recurrence was analyzed and Deptor expression was no significantly high in the patients with recurrence vs patients with no recurrence (*p=N/S).
DLBCL represents the most common Non-Hodgkin’s Lymphoma subtype and is within 30% to 40% of cases in the adult population [214]. DLBCL is an aggressive lymphoma subtype, where two histologically indistinguishable DLBCL subtypes have been identified by gene expression profiling, each derived from a different cell of origin (COO) [215]. The germinal center B-cell-like DLBCL (GCB) subtypes and the other activated B-cell-like subtype (ABC) have been identified. They are controlled by different oncogenic pathways, show different clinical behaviors, and have different clinical results. Therefore, the ABC-DLBCL subtype has worse prognosis and clinical evolution compared to GCB-DLBCL subtype [216, 217]. Because of this, we reviewed studies that contained Deptor expression data in ABC-DLBCL. One study with 17 samples, and others with 20 samples and 167 samples showed a significant expression of Deptor compared with normal memory B cells. In the last study, with 167 samples, an inverse relationship of Deptor expression and poor OS is observed, but is positively related to the presence of extra nodal sites (Figure 9B) [208, 217, 218]. We also reviewed studies that included GCB-DLBCL samples, finding an important expression of Deptor compared to ABC-DLBCL (Figure 9C) [218]. In other studies with 9, 40 and 183 samples respectively, where Deptor expression was analyzed [208, 217, 218], the study with 40 samples shows a moderate inverse relationship between Deptor expression and recurrence at 5 years (Figure 9D) [218].
Follicular lymphoma (FL) is the second most frequent subtype of malignant lymphomas and represents approximately 20% of all cases of lymphoma in the adult population. It is typically an indolent disease with long-term survival [219]. In this review we also analyzed Deptor expression in some studies that contained FL data. We found that in two of the three studies reviewed, one of 191 and the other with 5 samples show a low or negative expression of Deptor [211, 220]. The third study that includes 38 samples shows a high expression of Deptor [208], therefore, considering the size of the samples in general, it can be suggested that the FL shows a low or negative Deptor expression according to this analysis. This is consistent with other study where 8 cutaneous FL samples were included showing to be negative for Deptor expression (Table 3) [212].
Other frequently diagnosed aggressive B-NHLs include mantle cell lymphoma (MCL) and Burkitt's lymphoma (BL), while another prevalent indolent lymphoma includes marginal zone lymphoma (MZL). In our analysis in this review we found two studies that included 5 and 8 BL samples and are consistent with high Deptor expression [209, 211]. In the study of Brune et al, we observe a high expression of Deptor in BL compared to memory B-cell as control (Figure 10A). Interestingly in the last study, there is an inverse expression between Deptor vs Bcl-2, while there is a positive expression between Deptor and Bcl-6. Furthermore, Deptor expression is inverse related to poor 5-year PFS or OS (p =0.001). In addition, a Kaplan Meier analysis was performed to compare survival percent of patients with high Deptor expression. We observed that a high expression of Deptor correlated with a better overall survival percent (*p< 0.01). (Figure 10B–10D) [209]. Bcl-2 is an anti-apoptotic protein member of the Bcl-2 family of proteins that can cause interruption of the apoptotic process and survival of malignant cells [221]. The aberrant expression of the Bcl-2 protein contributes to the pathogenesis of many types of human malignancies, including leukemias and lymphomas. Within B-NHL, Bcl-2 overexpression commonly arises from genetic abnormalities and is substantially different in several lymphoma subtypes. Unlike the rest of B-NHL, the level of Bcl-2 expression in Burkitt lymphoma is low or undetectable, which has been used as part of the diagnostic algorithm for this subtype of lymphoma [222, 223].
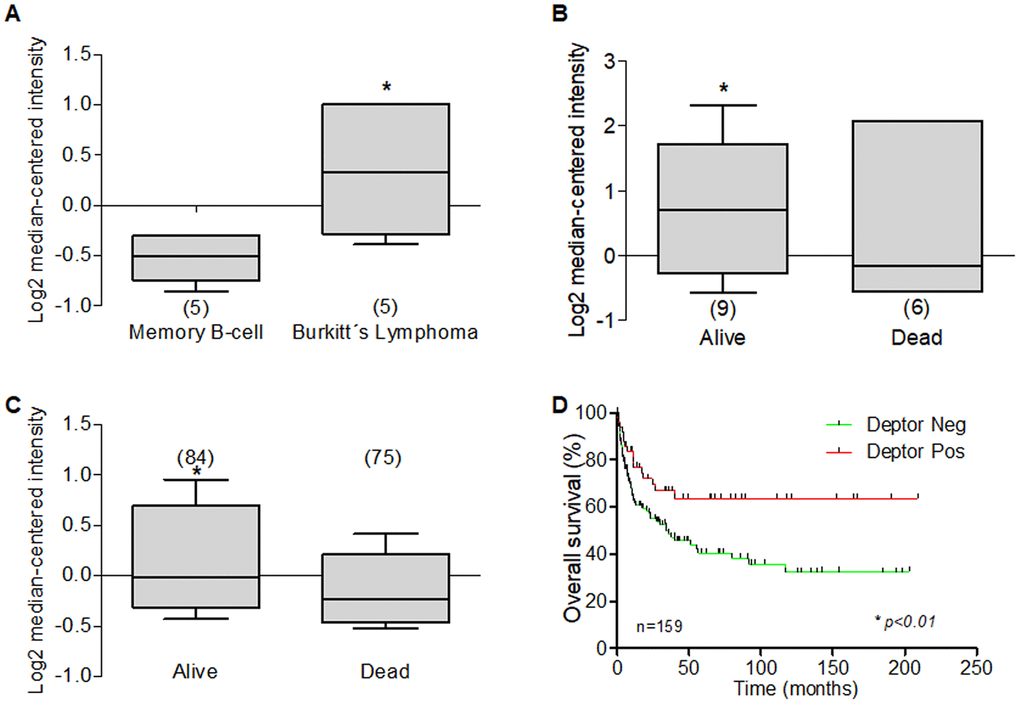
Figure 10. Deptor expression in Burkitt’s Lymphoma. Oncomine analysis was done to revised Deptor expression in Burkitt’s Lymphoma. (A) in Brune et al, analysis, show high Deptor expression Burkitt’s Lymphoma compared to memory b-cell (*p<0.05). (B) In Hummel et al, shows a moderate high Deptor expression the alive patients at 5 years or at total OS (C) (*p=N/S or *p<0.05 respectively). (D) Overall survival of patients with DLBCL according to Deptor expression. Number of patients (n) is listed next to the graph (*p>0.01).
Therefore, probably due to the low expression of Bcl2, it was not possible to establish any relationship between the positive expression of Deptor and Bcl-2. Knowing the important role of Bcl-2 in the response to treatment and the inverse expression of Deptor, it is not surprising that the expression Deptor is not related to response to treatment. So, these observations briefly suggest that Deptor may have a role in lymphoma development than in mechanisms of resistance to therapy. BCL6 gene encodes a transcription factor that is critical to the normal development of the B-cell germinal center reaction important for cell proliferation and DNA damage. Bcl-6 activity can be deregulated by a variety of mechanisms and contributes to the development of B-cell lymphoma. Therefore, finding a positive relationship between the presence of Bcl-6 and Deptor may suggest that Deptor has some role in the development of Burkitt’s Lymphoma. Furthermore, our review of the in silico analysis of possible transcription factors that regulate Deptor expression is interestingly found in Bcl-6, as described in the Deptor regulation section (Table 1 and Figure 3). Likewise, our analysis shows an important expression of Deptor in 57 MCL samples, which, as already mentioned, together with BL is another of the most aggressive B-NHL subtypes [224]. Therefore, this data suggest that Deptor expression is more related to more aggressive B-NHL subtypes like ABC-DLBCL and GCB-DLBCL. We also analyzed a report of MZL, an indolent B-NHL subtype, and found low and negative Deptor expression compared to tonsil tissue [212]. This seems to be consistent with our hypothesis of low Deptor expression in less aggressive lymphoma subtypes (Table 3).
ALK-positive anaplastic large cell lymphoma (ALK+ ALCL) is a rare subtype of peripheral T-cell Non-Hodgkin Lymphoma (NHL), and is more common in children and young adults. Patients generally have advanced stage (stage III or IV) disease and systemic symptoms (75%) [225]. Lymphoma cells show strong and uniform CD30 expression and aberrant expression of anaplastic lymphoma kinase (ALK) protein due to translocations involving the ALK gene at the 2p23 locus [80]. ALK+ ALCL has a more favorable clinical course than ALK negative cases [226]. Two studies included in our analysis show that there are no differences in Deptor expression between Anaplastic Large Cell Lymphoma, ALK-Negative (4 samples), and Anaplastic Large Cell Lymphoma, ALK-Positive (5 samples) [227], while the other study shows a very high expression of Deptor in ALCL (30 samples) [228]. A study containing 13 samples of Adult T-Cell Leukemia/Lymphoma and 37 samples of Angioimmunoblastic T-Cell Lymphoma was also analyzed, showing that both conditions show a very important expression of Deptor in the samples analyzed (Figure 11) [228]. Adult T-Cell Leukemia/Lymphoma (ATL) is an aggressive type of intractable peripheral T-cell malignancy. It is caused by infections with the human T-cell leukemia virus type I (HTLV-1). The results, after therapeutic interventions for ATL, have not been satisfactory [229], and is characterized by multiple organ invasion by ATL cells, a high frequency of opportunistic infections, resistance to chemotherapeutic drugs, and poor prognosis [230].
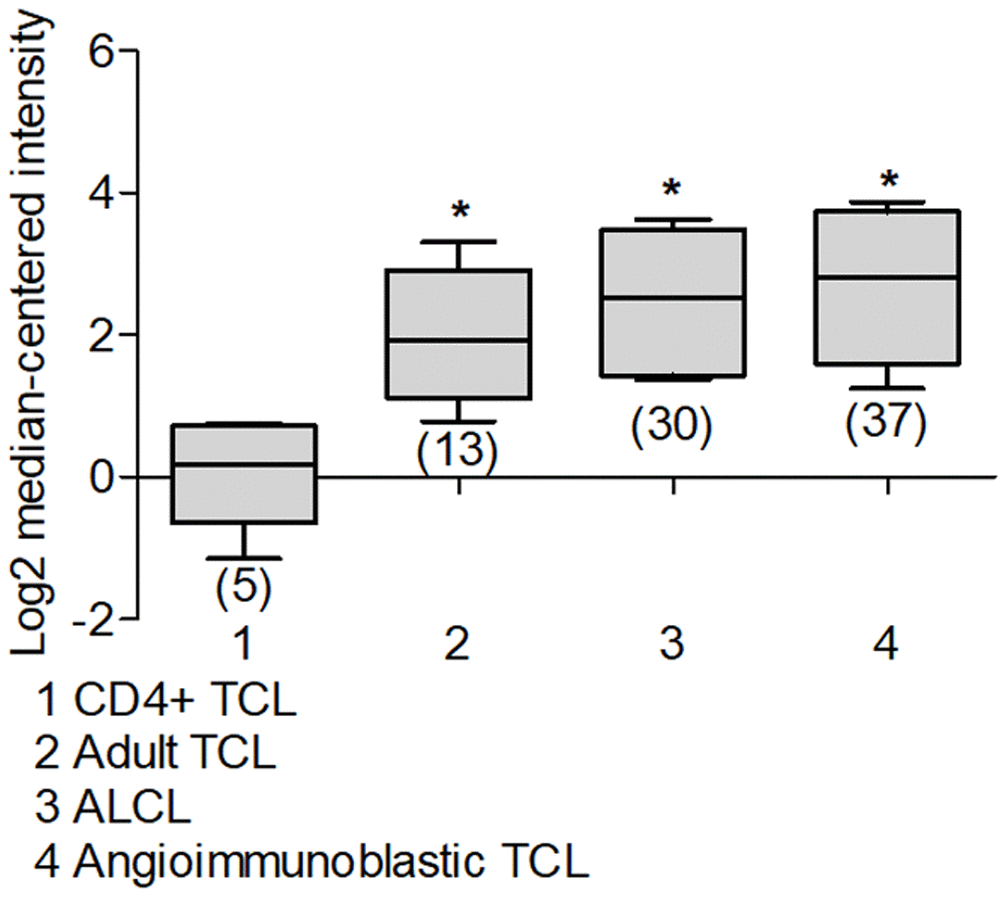
Figure 11. Deptor expression in T-cell Non-Hodgkin Lymphoma. Using Oncomine we analyzed Deptor expression on TCL and ALCL. A) a study by Iqbal et al, show high Deptor expression in Adult T-Cell leukemia, Anaplastic Large Cell Lymphoma and Angioimmunoblastic T-cell lymphoma compared with CD4+ T cell lymphoma (*p<0.001).
Angioimmunoblastic T-cell lymphoma (AITL) is another peripheral T-cell lymphoma with aggressive lymphoproliferative disorders, almost all associated with poor clinical outcomes [231]. The review of our Deptor expression analysis suggests that ATLs generally show significant Deptor expression, although some relationship of Deptor expression with some clinical characteristic cannot be established. Therefore, additional analysis and studies are necessary to establish the importance and role of Deptor in ATLs (Table 3).
Hodgkin Lymphoma (HL) is a lymphoid neoplasm and is one of the most common cancers diagnosed in adolescents. HL is classified according to immunohistochemistry and biological behavior in classical HL (cHL) and predominant HL in nodular lymphocytes. CHL is a more aggressive disease, accounting for 95% of HL cases, while Nodular lymphocyte predominant HL is rarer showing indolent and localized behavior [232, 233]. In a study where Hodgkin's Lymphoma (130 samples) was analyzed, a very important Deptor expression was observed (Figure 12A), and this expression was moderately related to the Ann Arbor stage and to the relapse in the refractory population to treatment (Figure 12B), although there was no relationship with total OS or recurrence. (Figure 12C, 12D) [234]. In another study that included Mixed Cellularity Classical Hodgkin's Lymphoma (17 samples), Nodular Sclerosis Classical Hodgkin's Lymphoma (42 samples), shows an important expression of Deptor, although this is not related to any clinical characteristic such as the outcome [235]. Therefore, according to the review of these analyzes, it is suggested that high Deptor expression in cHL and could be related to the stage and relapse of the disease (Table 3). However, further studies and analysis are necessary to establish the role of Deptor more precisely in cHL.
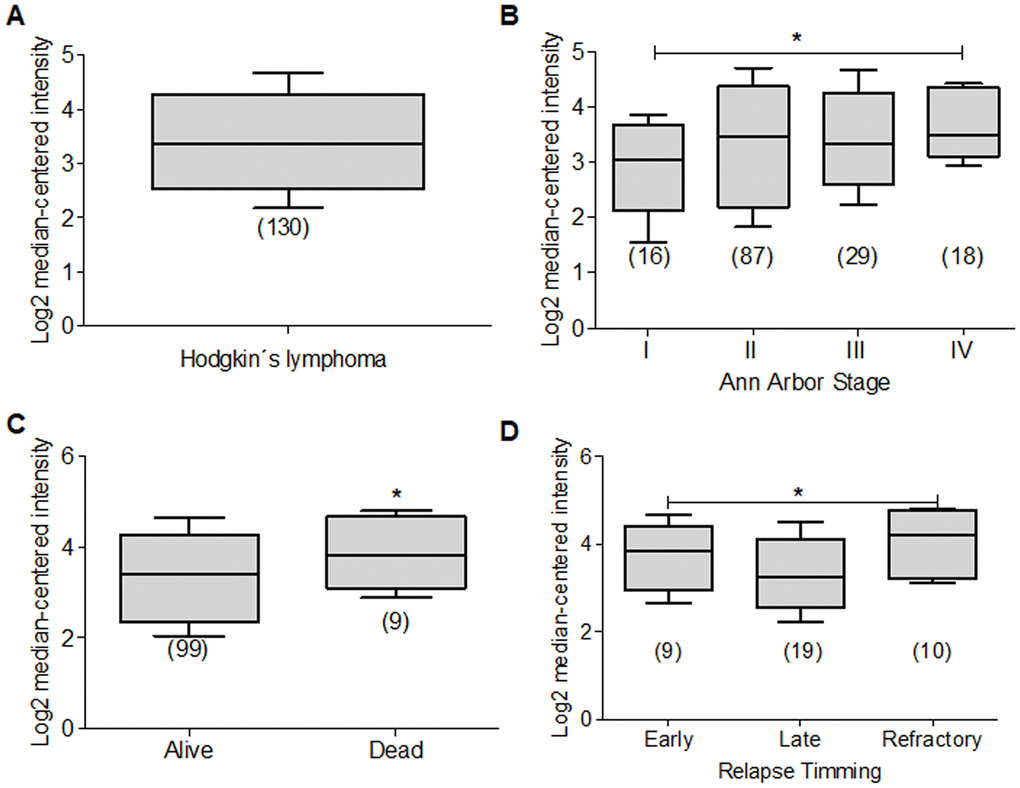
Figure 12. Deptor expression in Hodgkin lymphoma. Deptor expression in Hodgkin Lymphoma was revised by Oncomine analyzes. (A) Hodgkin Lymphoma shown a high Deptor expression. (B) Deptor expression has relationship with stage of the disease, stage IV shown higher Deptor expression compared with stage I (*p<0.05). (C) Deptor expression is higher in dead patients vs alive patients when OS was analyzed (*p<0.05). (D) Relapse timing was related to Deptor expression in refractory is higher Detour expression compare with late relapse timing (*p<0.05).
Conclusions
Since the discovery of Deptor in 2009, more than ten years ago, several a large number of information has been accumulated, revealing its importance in a large number of biological processes. As described in this review, Deptor appears to play an important role in the pathogenesis of some types of hematologic malignancies. This is due to its ability to control mTOR activity and PI3K-AKT axis activity, which has been of great importance in cancer among other biological processes. In recent years, Deptor has become an important therapeutic target not only in cancer, but also in metabolic and immunity processes. Therefore, in the coming years the development of many therapies based on the regulation of this protein will not be surprising.
Author Contributions
Performed the GEO analysis: Mario Morales-Martinez and Mario I. Vega. Analyzed the data and interpreted the results: M. Morales-Martinez, M. I. Vega. Designed, supervised, and/or obtained support for the study: Alan Lichtenstein. M. I. Vega. Wrote the manuscript: M. Morales-Martinez, M. I. Vega. All authors revised and approved the final manuscript.
Acknowledgments
Mario Morales-Martinez is a doctoral student from Programa de Doctorado en Ciencias Biomédicas, Universidad Nacional Autónoma de México (UNAM) and he has received CONACyT fellowship (596754). The authors thank Dr. Sara Huerta-Yepez and Owen Vega for constructive criticism of the paper. Anya Vega for all her support.
Conflicts of Interest
No conflicts of interest, financial or otherwise, are declared by the authors.
Funding
This study was supported in part by NIH grants RO1CA111448, RO1CA211562, R21CA168491, RO1CA217820.
References
- 1. Peterson TR, Laplante M, Thoreen CC, Sancak Y, Kang SA, Kuehl WM, Gray NS, Sabatini DM. DEPTOR is an mTOR inhibitor frequently overexpressed in multiple myeloma cells and required for their survival. Cell. 2009; 137:873–86. https://doi.org/10.1016/j.cell.2009.03.046 [PubMed]
- 2. Panic N, Larghi A, Amore R, Pastorino R, Bulajic M, Costamagna G, Boccia S. Single nucleotide polymorphisms within the 8Q24 region are not associated with the risk of intraductal papillary mucinous neoplasms of the pancreas. J Gastrointestin Liver Dis. 2016; 25:311–15. https://doi.org/10.15403/jgld.2014.1121.253.q24 [PubMed]
- 3. Ota T, Suzuki Y, Nishikawa T, Otsuki T, Sugiyama T, Irie R, Wakamatsu A, Hayashi K, Sato H, Nagai K, Kimura K, Makita H, Sekine M, et al. Complete sequencing and characterization of 21,243 full-length human cDNAs. Nat Genet. 2004; 36:40–45. https://doi.org/10.1038/ng1285 [PubMed]
- 4. Caron A, Briscoe DM, Richard D, Laplante M. DEPTOR at the nexus of cancer, metabolism, and immunity. Physiol Rev. 2018; 98:1765–803. https://doi.org/10.1152/physrev.00064.2017 [PubMed]
- 5. Laplante M, Sabatini DM. mTOR signaling. Cold Spring Harb Perspect Biol. 2012; 4:a011593. https://doi.org/10.1101/cshperspect.a011593 [PubMed]
- 6. Harrington LS, Findlay GM, Gray A, Tolkacheva T, Wigfield S, Rebholz H, Barnett J, Leslie NR, Cheng S, Shepherd PR, Gout I, Downes CP, Lamb RF. The TSC1-2 tumor suppressor controls insulin-PI3K signaling via regulation of IRS proteins. J Cell Biol. 2004; 166:213–23. https://doi.org/10.1083/jcb.200403069 [PubMed]
- 7. Um SH, Frigerio F, Watanabe M, Picard F, Joaquin M, Sticker M, Fumagalli S, Allegrini PR, Kozma SC, Auwerx J, Thomas G. Absence of S6K1 protects against age- and diet-induced obesity while enhancing insulin sensitivity. Nature. 2004; 431:200–05. https://doi.org/10.1038/nature02866 [PubMed]
- 8. Yu Y, Yoon SO, Poulogiannis G, Yang Q, Ma XM, Villén J, Kubica N, Hoffman GR, Cantley LC, Gygi SP, Blenis J. Phosphoproteomic analysis identifies Grb10 as an mTORC1 substrate that negatively regulates insulin signaling. Science. 2011; 332:1322–26. https://doi.org/10.1126/science.1199484 [PubMed]
- 9. Hsu PP, Kang SA, Rameseder J, Zhang Y, Ottina KA, Lim D, Peterson TR, Choi Y, Gray NS, Yaffe MB, Marto JA, Sabatini DM. The mTOR-regulated phosphoproteome reveals a mechanism of mTORC1-mediated inhibition of growth factor signaling. Science. 2011; 332:1317–22. https://doi.org/10.1126/science.1199498 [PubMed]
- 10. Zoncu R, Bar-Peled L, Efeyan A, Wang S, Sancak Y, Sabatini DM. mTORC1 senses lysosomal amino acids through an inside-out mechanism that requires the vacuolar H(+)-ATPase. Science. 2011; 334:678–83. https://doi.org/10.1126/science.1207056 [PubMed]
- 11. Wang Z, Zhong J, Inuzuka H, Gao D, Shaik S, Sarkar FH, Wei W. An evolving role for DEPTOR in tumor development and progression. Neoplasia. 2012; 14:368–75. https://doi.org/10.1593/neo.12542 [PubMed]
- 12. Catena V, Fanciulli M. Deptor: not only a mTOR inhibitor. J Exp Clin Cancer Res. 2017; 36:12. https://doi.org/10.1186/s13046-016-0484-y [PubMed]
- 13. Chen S, Hamm HE. DEP domains: more than just membrane anchors. Dev Cell. 2006; 11:436–38. https://doi.org/10.1016/j.devcel.2006.09.011 [PubMed]
- 14. Consonni SV, Maurice MM, Bos JL. DEP domains: structurally similar but functionally different. Nat Rev Mol Cell Biol. 2014; 15:357–62. https://doi.org/10.1038/nrm3791 [PubMed]
- 15. Pan WJ, Pang SZ, Huang T, Guo HY, Wu D, Li L. Characterization of function of three domains in dishevelled-1: DEP domain is responsible for membrane translocation of dishevelled-1. Cell Res. 2004; 14:324–30. https://doi.org/10.1038/sj.cr.7290232 [PubMed]
- 16. Lee HJ, Zheng JJ. PDZ domains and their binding partners: structure, specificity, and modification. Cell Commun Signal. 2010; 8:8. https://doi.org/10.1186/1478-811X-8-8 [PubMed]
- 17. Neubig RR, Siderovski DP. Regulators of G-protein signalling as new central nervous system drug targets. Nat Rev Drug Discov. 2002; 1:187–97. https://doi.org/10.1038/nrd747 [PubMed]
- 18. Vega MI, Shi Y, Frost P, Huerta-Yepez S, Antonio-Andres G, Hernandez-Pando R, Lee J, Jung ME, Gera JF, Lichtenstein A. A novel therapeutic induces DEPTOR degradation in multiple myeloma cells with resulting tumor cytotoxicity. Mol Cancer Ther. 2019; 18:1822–31. https://doi.org/10.1158/1535-7163.MCT-19-0115 [PubMed]
- 19. Duan S, Skaar JR, Kuchay S, Toschi A, Kanarek N, Ben-Neriah Y, Pagano M. mTOR generates an auto-amplification loop by triggering the βTrCP- and CK1α-dependent degradation of DEPTOR. Mol Cell. 2011; 44:317–24. https://doi.org/10.1016/j.molcel.2011.09.005 [PubMed]
- 20. Zhao Y, Xiong X, Sun Y. DEPTOR, an mTOR inhibitor, is a physiological substrate of SCF(βTrCP) E3 ubiquitin ligase and regulates survival and autophagy. Mol Cell. 2011; 44:304–16. https://doi.org/10.1016/j.molcel.2011.08.029 [PubMed]
- 21. Zhang H, Zhang Q, Gao G, Wang X, Wang T, Kong Z, Wang G, Zhang C, Wang Y, Peng G. UBTOR/KIAA1024 regulates neurite outgrowth and neoplasia through mTOR signaling. PLoS Genet. 2018; 14:e1007583. https://doi.org/10.1371/journal.pgen.1007583 [PubMed]
- 22. Dwivedi P, Muench DE, Wagner M, Azam M, Grimes HL, Greis KD. Time resolved quantitative phospho-tyrosine analysis reveals Bruton’s Tyrosine kinase mediated signaling downstream of the mutated granulocyte-colony stimulating factor receptors. Leukemia. 2019; 33:75–87. https://doi.org/10.1038/s41375-018-0188-8 [PubMed]
- 23. Myhrvold IK, Cremaschi A, Hermansen JU, Tjønnfjord GE, Munthe LA, Taskén K, Skånland SS. Single cell profiling of phospho-protein levels in chronic lymphocytic leukemia. Oncotarget. 2018; 9:9273–84. https://doi.org/10.18632/oncotarget.23949 [PubMed]
- 24. Muench DE, Olsson A, Ferchen K, Pham G, Serafin RA, Chutipongtanate S, Dwivedi P, Song B, Hay S, Chetal K, Trump-Durbin LR, Mookerjee-Basu J, Zhang K, et al. Mouse models of neutropenia reveal progenitor-stage-specific defects. Nature. 2020; 582:109–14. https://doi.org/10.1038/s41586-020-2227-7 [PubMed]
- 25. Dwivedi P, Chutipongtanate S, Muench DE, Azam M, Grimes HL, Greis KD. SWATH-proteomics of Ibrutinib’s action in myeloid leukemia initiating mutated G-CSFR signaling. Proteomics Clin Appl. 2020; 14:e1900144. https://doi.org/10.1002/prca.201900144 [PubMed]
- 26. Schumich A, Prchal-Murphy M, Maurer-Granofszky M, Hoelbl-Kovacic A, Mühlegger N, Pötschger U, Fajmann S, Haas OA, Nebral K, von Neuhoff N, Zimmermann M, Boztug H, Rasche M, et al. Phospho-profiling linking biology and clinics in pediatric acute myeloid leukemia. Hemasphere. 2019; 4:e312. https://doi.org/10.1097/HS9.0000000000000312 [PubMed]
- 27. Dwivedi P, Muench DE, Wagner M, Azam M, Grimes HL, Greis KD. Phospho serine and threonine analysis of normal and mutated granulocyte colony stimulating factor receptors. Sci Data. 2019; 6:21. https://doi.org/10.1038/s41597-019-0015-8 [PubMed]
- 28. Marmiroli S, Fabbro D, Miyata Y, Pierobon M, Ruzzene M. Phosphorylation, Signaling, and Cancer: Targets and Targeting. Biomed Res Int. 2015. https://doi.org/10.1155/2015/601543 [PubMed]
- 29. Dwivedi P, Greis KD. Granulocyte colony-stimulating factor receptor signaling in severe congenital neutropenia, chronic neutrophilic leukemia, and related malignancies. Exp Hematol. 2017; 46:9–20. https://doi.org/10.1016/j.exphem.2016.10.008 [PubMed]
- 30. Samaras P, Schmidt T, Frejno M, Gessulat S, Reinecke M, Jarzab A, Zecha J, Mergner J, Giansanti P, Ehrlich HC, Aiche S, Rank J, Kienegger H, et al. ProteomicsDB: a multi-omics and multi-organism resource for life science research. Nucleic Acids Res. 2020; 48:D1153–63. https://doi.org/10.1093/nar/gkz974 [PubMed]
- 31. Binder JX, Pletscher-Frankild S, Tsafou K, Stolte C, O’Donoghue SI, Schneider R, Jensen LJ. COMPARTMENTS: unification and visualization of protein subcellular localization evidence. Database (Oxford). 2014; 2014:bau012. https://doi.org/10.1093/database/bau012 [PubMed]
- 32. The Human Protein Atlas. https://www.proteinatlas.org/.
- 33. Catena V, Bruno T, De Nicola F, Goeman F, Pallocca M, Iezzi S, Sorino C, Cigliana G, Floridi A, Blandino G, Fanciulli M. Deptor transcriptionally regulates endoplasmic reticulum homeostasis in multiple myeloma cells. Oncotarget. 2016; 7:70546–58. https://doi.org/10.18632/oncotarget.12060 [PubMed]
- 34. Kim LC, Cook RS, Chen J. mTORC1 and mTORC2 in cancer and the tumor microenvironment. Oncogene. 2017; 36:2191–201. https://doi.org/10.1038/onc.2016.363 [PubMed]
- 35. Chen SQ, Li JQ, Wang XQ, Lei WJ, Li H, Wan J, Hu Z, Zou YW, Wu XY, Niu HX. EZH2-inhibitor DZNep enhances apoptosis of renal tubular epithelial cells in presence and absence of cisplatin. Cell Div. 2020; 15:8. https://doi.org/10.1186/s13008-020-00064-3 [PubMed]
- 36. Wei FZ, Cao Z, Wang X, Wang H, Cai MY, Li T, Hattori N, Wang D, Du Y, Song B, Cao LL, Shen C, Wang L, et al. Epigenetic regulation of autophagy by the methyltransferase EZH2 through an MTOR-dependent pathway. Autophagy. 2015; 11:2309–22. https://doi.org/10.1080/15548627.2015.1117734 [PubMed]
- 37. Ohshima K, Nojima S, Tahara S, Kurashige M, Hori Y, Hagiwara K, Okuzaki D, Oki S, Wada N, Ikeda JI, Kanai Y, Morii E. Argininosuccinate synthase 1-deficiency enhances the cell sensitivity to arginine through decreased DEPTOR expression in endometrial cancer. Sci Rep. 2017; 7:45504. https://doi.org/10.1038/srep45504 [PubMed]
- 38. Kanno Y, Zhao S, Yamashita N, Yanai K, Nemoto K, Inouye Y. Androgen receptor functions as a negative transcriptional regulator of DEPTOR, mTOR inhibitor. J Toxicol Sci. 2015; 40:753–58. https://doi.org/10.2131/jts.40.753 [PubMed]
- 39. Yen CH, Lu YC, Li CH, Lee CM, Chen CY, Cheng MY, Huang SF, Chen KF, Cheng AL, Liao LY, Lee YH, Chen YM. Functional characterization of glycine N-methyltransferase and its interactive protein DEPDC6/DEPTOR in hepatocellular carcinoma. Mol Med. 2012; 18:286–96. https://doi.org/10.2119/molmed.2011.00331 [PubMed]
- 40. Hu Y, Su H, Liu C, Wang Z, Huang L, Wang Q, Liu S, Chen S, Zhou J, Li P, Chen Z, Liu H, Qing G. DEPTOR is a direct NOTCH1 target that promotes cell proliferation and survival in T-cell leukemia. Oncogene. 2017; 36:1038–47. https://doi.org/10.1038/onc.2016.275 [PubMed]
- 41. Sorrentino C, Cuneo A, Roti G. Therapeutic Targeting of Notch Signaling Pathway in Hematological Malignancies. Mediterr J Hematol Infect Dis. 2019; 11. https://doi.org/10.4084/MJHID.2019.037 [PubMed]
- 42. Singh P, Brock CO, Volden PA, Hernandez K, Skor M, Kocherginsky M, Park JE, Brady MJ, Conzen SD. Glucocorticoid receptor ChIP-sequencing of subcutaneous fat reveals modulation of inflammatory pathways. Obesity (Silver Spring). 2015; 23:2286–93. https://doi.org/10.1002/oby.21251 [PubMed]
- 43. Parvani JG, Davuluri G, Wendt MK, Espinosa C, Tian M, Danielpour D, Sossey-Alaoui K, Schiemann WP. Deptor enhances triple-negative breast cancer metastasis and chemoresistance through coupling to survivin expression. Neoplasia. 2015; 17:317–28. https://doi.org/10.1016/j.neo.2015.02.003 [PubMed]
- 44. Lisse TS, Liu T, Irmler M, Beckers J, Chen H, Adams JS, Hewison M. Gene targeting by the vitamin D response element binding protein reveals a role for vitamin D in osteoblast mTOR signaling. FASEB J. 2011; 25:937–47. https://doi.org/10.1096/fj.10-172577 [PubMed]
- 45. Wilson RL, Buckberry S, Spronk F, Laurence JA, Leemaqz S, O’Leary S, Bianco-Miotto T, Du J, Anderson PH, Roberts CT. Vitamin D receptor gene ablation in the conceptus has limited effects on placental morphology, function and pregnancy outcome. PLoS One. 2015; 10:e0131287. https://doi.org/10.1371/journal.pone.0131287 [PubMed]
- 46. Kaul D, Mehrotra A. Functional characterization of AATF transcriptome in human leukemic cells. Mol Cell Biochem. 2007; 297:215–20. https://doi.org/10.1007/s11010-006-9317-1 [PubMed]
- 47. Kaul D, Hussain A. Cellular AATF gene encodes a novel miRNA that can contribute to HIV-1 latency. Indian J Biochem Biophys. 2009; 46:237–40. [PubMed]
- 48. Malik D, Kaul D, Chauhan N, Marwaha RK. miR-2909-mediated regulation of KLF4: a novel molecular mechanism for differentiating between B-cell and T-cell pediatric acute lymphoblastic leukemias. Mol Cancer. 2014; 13:175. https://doi.org/10.1186/1476-4598-13-175 [PubMed]
- 49. Wani S, Kaul D. Cancer cells govern miR-2909 exosomal recruitment through its 3'-end post-transcriptional modification. Cell Biochem Funct. 2018; 36:106–11. https://doi.org/10.1002/cbf.3323 [PubMed]
- 50. Meng ZX, Wang L, Xiao Y, Lin JD. The Baf60c/deptor pathway links skeletal muscle inflammation to glucose homeostasis in obesity. Diabetes. 2014; 63:1533–45. https://doi.org/10.2337/db13-1061 [PubMed]
- 51. Wang Q, Zhou Y, Rychahou P, Harris JW, Zaytseva YY, Liu J, Wang C, Weiss HL, Liu C, Lee EY, Evers BM. Deptor is a novel target of Wnt/β-catenin/c-Myc and contributes to colorectal cancer cell growth. Cancer Res. 2018; 78:3163–75. https://doi.org/10.1158/0008-5472.CAN-17-3107 [PubMed]
- 52. Fornes O, Castro-Mondragon JA, Khan A, van der Lee R, Zhang X, Richmond PA, Modi BP, Correard S, Gheorghe M, Baranašić D, Santana-Garcia W, Tan G, Chèneby J, et al. JASPAR 2020: update of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 2020; 48:D87–92. https://doi.org/10.1093/nar/gkz1001 [PubMed]
- 53. Dausinas P, Pulakanti K, Rao S, Cole JM, Dahl R, Cowden Dahl KD. ARID3A and ARID3B induce stem promoting pathways in ovarian cancer cells. Gene. 2020; 738:144458. https://doi.org/10.1016/j.gene.2020.144458 [PubMed]
- 54. Li YS, Zhou Y, Tang L, Shinton SA, Hayakawa K, Hardy RR. A developmental switch between fetal and adult B lymphopoiesis. Ann N Y Acad Sci. 2015; 1362:8–15. https://doi.org/10.1111/nyas.12769 [PubMed]
- 55. Puissegur MP, Eichner R, Quelen C, Coyaud E, Mari B, Lebrigand K, Broccardo C, Nguyen-Khac F, Bousquet M, Brousset P. B-cell regulator of immunoglobulin heavy-chain transcription (Bright)/ARID3a is a direct target of the oncomir microRNA-125b in progenitor B-cells. Leukemia. 2012; 26:2224–32. https://doi.org/10.1038/leu.2012.95 [PubMed]
- 56. Kassambara A, Jourdan M, Bruyer A, Robert N, Pantesco V, Elemento O, Klein B, Moreaux J. Global miRNA expression analysis identifies novel key regulators of plasma cell differentiation and malignant plasma cell. Nucleic Acids Res. 2017; 45:5639–52. https://doi.org/10.1093/nar/gkx327 [PubMed]
- 57. Ma L, Cao Y, Hu JJ, Chu ML. Casein kinase 2 interacting protein 1 positively regulates caudal-related homeobox 1 in intestinal-type gastric cancer. Chin Med J (Engl). 2020; 133:154–64. https://doi.org/10.1097/CM9.0000000000000604 [PubMed]
- 58. Heckman CA, Duan H, Garcia PB, Boxer LM. Oct transcription factors mediate t(14;18) lymphoma cell survival by directly regulating bcl-2 expression. Oncogene. 2006; 25:888–98. https://doi.org/10.1038/sj.onc.1209127 [PubMed]
- 59. Paczulla A, Kollotzek F, Grzywna S, Kanz L, Konantz M, Lengerke C. The role of Cdx2 in human leukemogenesis. Exp Hematol. 2013; 41:S58. https://doi.org/10.1016/j.exphem.2013.05.227
- 60. Das F, Ghosh-Choudhury N, Venkatesan B, Li X, Mahimainathan L, Choudhury GG. Akt kinase targets association of CBP with SMAD 3 to regulate TGFbeta-induced expression of plasminogen activator inhibitor-1. J Cell Physiol. 2008; 214:513–27. https://doi.org/10.1002/jcp.21236 [PubMed]
- 61. Shabestari RM, Safa M, Alikarami F, Banan M, Kazemi A. CREB knockdown inhibits growth and induces apoptosis in human pre-B acute lymphoblastic leukemia cells through inhibition of prosurvival signals. Biomed Pharmacother. 2017; 87:274–79. https://doi.org/10.1016/j.biopha.2016.12.070 [PubMed]
- 62. Perez DR, Smagley Y, Garcia M, Carter MB, Evangelisti A, Matlawska-Wasowska K, Winter SS, Sklar LA, Chigaev A. Cyclic AMP efflux inhibitors as potential therapeutic agents for leukemia. Oncotarget. 2016; 7:33960–82. https://doi.org/10.18632/oncotarget.8986 [PubMed]
- 63. Huang S, Ren Y, Wang P, Li Y, Wang X, Zhuang H, Fang R, Wang Y, Liu N, Hehir M, Zhou JX. Transcription factor CREB is involved in CaSR-mediated cytoskeleton gene expression. Anat Rec (Hoboken). 2015; 298:501–12. https://doi.org/10.1002/ar.23089 [PubMed]
- 64. Zhu YX, Yin H, Bruins LA, Shi CX, Jedlowski P, Aziz M, Sereduk C, Kortuem KM, Schmidt JE, Champion M, Braggio E, Keith Stewart A. RNA interference screening identifies lenalidomide sensitizers in multiple myeloma, including RSK2. Blood. 2015; 125:483–91. https://doi.org/10.1182/blood-2014-05-577130 [PubMed]
- 65. Evans PM, Liu C. Roles of krüpel-like factor 4 in normal homeostasis, cancer and stem cells. Acta Biochim Biophys Sin (Shanghai). 2008; 40:554–64. https://doi.org/10.1111/j.1745-7270.2008.00439.x [PubMed]
- 66. Morales-Martinez M, Franco-Cea LA, Moreno Vargas L, Martinez-Maza O, Huerta-Yepez S, Vega MI. Bifunctional role of Kruppel-like Factor 4 in hematological malignancies. For Immunopathol Dis Therap. 2016; 7:141–51. http://www.dl.begellhouse.com/journals/1a654bf03faf67ac,2451af3842dace87,0fbb4f3b40af1beb.html.
- 67. Chen X, Shi W, Zhang H. The role of KLF14 in multiple disease processes. Biofactors. 2020; 46:276–82. https://doi.org/10.1002/biof.1612 [PubMed]
- 68. Meliala IT, Hosea R, Kasim V, Wu S. The biological implications of Yin Yang 1 in the hallmarks of cancer. Theranostics. 2020; 10:4183–200. https://doi.org/10.7150/thno.43481 [PubMed]
- 69. Morales-Martinez M, Valencia-Hipolito A, Vega GG, Neri N, Nambo MJ, Alvarado I, Cuadra I, Duran-Padilla MA, Martinez-Maza O, Huerta-Yepez S, Vega MI. Regulation of krüppel-like factor 4 (KLF4) expression through the transcription factor Yin-Yang 1 (YY1) in non-Hodgkin B-cell lymphoma. Oncotarget. 2019; 10:2173–88. https://doi.org/10.18632/oncotarget.26745 [PubMed]
- 70. Kleiman E, Jia H, Loguercio S, Su AI, Feeney AJ. YY1 plays an essential role at all stages of B-cell differentiation. Proc Natl Acad Sci USA. 2016; 113:E3911–20. https://doi.org/10.1073/pnas.1606297113 [PubMed]
- 71. Castellano G, Torrisi E, Ligresti G, Nicoletti F, Malaponte G, Traval S, McCubrey JA, Canevari S, Libra M. Yin yang 1 overexpression in diffuse large B-cell lymphoma is associated with B-cell transformation and tumor progression. Cell Cycle. 2010; 9:557–63. https://doi.org/10.4161/cc.9.3.10554 [PubMed]
- 72. Bonavida B, Huerta-Yepez S, Baritaki S, Vega M, Liu H, Chen H, Berenson J. Overexpression of Yin Yang 1 in the pathogenesis of human hematopoietic Malignancies. Crit Rev Oncog. 2011; 16:261–67. https://doi.org/10.1615/critrevoncog.v16.i3-4.90 [PubMed]
- 73. Ye BH, Lista F, Lo Coco F, Knowles DM, Offit K, Chaganti RS, Dalla-Favera R. Alterations of a zinc finger-encoding gene, BCL-6, in diffuse large-cell lymphoma. Science. 1993; 262:747–50. https://doi.org/10.1126/science.8235596 [PubMed]
- 74. Basso K, Dalla-Favera R. BCL6: master regulator of the germinal center reaction and key oncogene in B cell lymphomagenesis. Adv Immunol. 2010; 105:193–210. https://doi.org/10.1016/S0065-2776(10)05007-8 [PubMed]
- 75. Hatzi K, Melnick A. Breaking bad in the germinal center: how deregulation of BCL6 contributes to lymphomagenesis. Trends Mol Med. 2014; 20:343–52. https://doi.org/10.1016/j.molmed.2014.03.001 [PubMed]
- 76. Hatzi K, Jiang Y, Huang C, Garrett-Bakelman F, Gearhart MD, Giannopoulou EG, Zumbo P, Kirouac K, Bhaskara S, Polo JM, Kormaksson M, MacKerell AD
Jr , Xue F, et al. A hybrid mechanism of action for BCL6 in B cells defined by formation of functionally distinct complexes at enhancers and promoters. Cell Rep. 2013; 4:578–88. https://doi.org/10.1016/j.celrep.2013.06.016 [PubMed] - 77. Saito M, Novak U, Piovan E, Basso K, Sumazin P, Schneider C, Crespo M, Shen Q, Bhagat G, Califano A, Chadburn A, Pasqualucci L, Dalla-Favera R. BCL6 suppression of BCL2 via Miz1 and its disruption in diffuse large B cell lymphoma. Proc Natl Acad Sci USA. 2009; 106:11294–99. https://doi.org/10.1073/pnas.0903854106 [PubMed]
- 78. Ye BH, Rao PH, Chaganti RS, Dalla-Favera R. Cloning of bcl-6, the locus involved in chromosome translocations affecting band 3q27 in B-cell lymphoma. Cancer Res. 1993; 53:2732–35. [PubMed]
- 79. Chen W, Iida S, Louie DC, Dalla-Favera R, Chaganti RS. Heterologous promoters fused to BCL6 by chromosomal translocations affecting band 3q27 cause its deregulated expression during B-cell differentiation. Blood. 1998; 91:603–07. [PubMed]
- 80. Swerdlow SH, Campo E, Pileri SA, Harris NL, Stein H, Siebert R, Advani R, Ghielmini M, Salles GA, Zelenetz AD, Jaffe ES. The 2016 revision of the World Health Organization classification of lymphoid neoplasms. Blood. 2016; 127:2375–90. https://doi.org/10.1182/blood-2016-01-643569 [PubMed]
- 81. Iqbal J, Meyer PN, Smith LM, Johnson NA, Vose JM, Greiner TC, Connors JM, Staudt LM, Rimsza L, Jaffe E, Rosenwald A, Ott G, Delabie J, et al. BCL2 predicts survival in germinal center B-cell-like diffuse large B-cell lymphoma treated with CHOP-like therapy and rituximab. Clin Cancer Res. 2011; 17:7785–95. https://doi.org/10.1158/1078-0432.CCR-11-0267 [PubMed]
- 82. Savage KJ, Johnson NA, Ben-Neriah S, Connors JM, Sehn LH, Farinha P, Horsman DE, Gascoyne RD. MYC gene rearrangements are associated with a poor prognosis in diffuse large B-cell lymphoma patients treated with R-CHOP chemotherapy. Blood. 2009; 114:3533–37. https://doi.org/10.1182/blood-2009-05-220095 [PubMed]
- 83. Van Keimpema M, Grüneberg LJ, Mokry M, van Boxtel R, van Zelm MC, Coffer P, Pals ST, Spaargaren M. The forkhead transcription factor FOXP1 represses human plasma cell differentiation. Blood. 2015; 126:2098–109. https://doi.org/10.1182/blood-2015-02-626176 [PubMed]
- 84. Nutt SL, Hodgkin PD, Tarlinton DM, Corcoran LM. The generation of antibody-secreting plasma cells. Nat Rev Immunol. 2015; 15:160–71. https://doi.org/10.1038/nri3795 [PubMed]
- 85. Szpirer J, Szpirer C, Riviere M, Levan G, Marynen P, Cassiman JJ, Wiese R, DeLuca HF. The Sp1 transcription factor gene (SP1) and the 1,25-dihydroxyvitamin D3 receptor gene (VDR) are colocalized on human chromosome arm 12q and rat chromosome 7. Genomics. 1991; 11:168–73. https://doi.org/10.1016/0888-7543(91)90114-t [PubMed]
- 86. Vega MI, Huerta-Yepez S, Martinez-Paniagua M, Martinez-Miguel B, Hernandez-Pando R, González-Bonilla CR, Chinn P, Hanna N, Hariharan K, Jazirehi AR, Bonavida B. Rituximab-mediated cell signaling and chemo/immuno-sensitization of drug-resistant B-NHL is independent of its Fc functions. Clin Cancer Res. 2009; 15:6582–94. https://doi.org/10.1158/1078-0432.CCR-09-1234 [PubMed]
- 87. Jin Z, Zhou S, Ye H, Jiang S, Yu K, Ma Y. The mechanism of SP1/p300 complex promotes proliferation of multiple myeloma cells through regulating IQGAP1 transcription. Biomed Pharmacother. 2019; 119:109434. https://doi.org/10.1016/j.biopha.2019.109434 [PubMed]
- 88. Chen Z, Ma T, Huang C, Hu T, Li J. The pivotal role of microRNA-155 in the control of cancer. J Cell Physiol. 2014; 229:545–50. https://doi.org/10.1002/jcp.24492 [PubMed]
- 89. Wang J, Yu F, Jia X, Iwanowycz S, Wang Y, Huang S, Ai W, Fan D. MicroRNA-155 deficiency enhances the recruitment and functions of myeloid-derived suppressor cells in tumor microenvironment and promotes solid tumor growth. Int J Cancer. 2015; 136:E602–13. https://doi.org/10.1002/ijc.29151 [PubMed]
- 90. Martin EC, Krebs AE, Burks HE, Elliott S, Baddoo M, Collins-Burow BM, Flemington EK, Burow ME. miR-155 induced transcriptome changes in the MCF-7 breast cancer cell line leads to enhanced mitogen activated protein kinase signaling. Genes Cancer. 2014; 5:353–64. https://doi.org/10.18632/genesandcancer.33 [PubMed]
- 91. Jablonska E, Gorniak P, Prusisz W, Kiliszek P, Szydlowski M, Sewastianik T, Bialopiotrowicz E, Polak A, Prochorec-Sobieszek M, Szumera-Cieckiewicz A, Warzocha K, Juszczynski P. Downregulation of Deptor By MiR-155 Promotes Cell Survival through Activation of PI3K/AKT and NFkB Signaling in ABC-Type Diffuse Large B-Cell Lymphomas. Blood. 2016; 128:1761. https://doi.org/10.1182/blood.V128.22.1761.1761
- 92. Lu JJ, Yang WM, Li F, Zhu W, Chen Z. Tunneling nanotubes mediated microRNA-155 intercellular transportation promotes bladder cancer cells’ invasive and proliferative capacity. Int J Nanomedicine. 2019; 14:9731–43. https://doi.org/10.2147/IJN.S217277 [PubMed]
- 93. Stolyar MA, Gorbenko AS, Bakhtina VI, Martynova EV, Moskov VI, Mikhalev MA, Olkhovik TI, Khazieva AS, Olkhovskiy IA. [Investigation of miR-155 level in the blood of patients with chronic lymphocytic leukemia and Ph-negative myeloproliferative neoplasms]. Klin Lab Diagn. 2020; 65:258–64. https://doi.org/10.18821/0869-2084-2020-65-4-258-264 [PubMed]
- 94. Xiao CR, Hong XL, Hu JS, Chen YM, Lu QY. [Targeting miR155 restores chemotherapy sensitivity in drug-resistant myeloma cell-line RPMI8226/DOX cells]. Zhonghua Xue Ye Xue Za Zhi. 2017; 38:55–59. https://doi.org/10.3760/cma.j.issn.0253-2727.2017.01.012 [PubMed]
- 95. Due H, Brøndum RF, Young KH, Bøgsted M, Dybkær K. MicroRNAs associated to single drug components of R-CHOP identifies diffuse large B-cell lymphoma patients with poor outcome and adds prognostic value to the international prognostic index. BMC Cancer. 2020; 20:237. https://doi.org/10.1186/s12885-020-6643-8 [PubMed]
- 96. Han B, Wang S, Zhao H. MicroRNA-21 and microRNA-155 promote the progression of Burkitt’s lymphoma by the PI3K/AKT signaling pathway. Int J Clin Exp Pathol. 2020; 13:89–98. [PubMed]
- 97. Rastgoo N, Wu J, Liu M, Pourabdollah M, Atenafu EG, Reece D, Chen W, Chang H. Targeting CD47/TNFAIP8 by miR-155 overcomes drug resistance and inhibits tumor growth through induction of phagocytosis and apoptosis in multiple myeloma. Haematologica. 2019. [Epub ahead of print]. https://doi.org/10.3324/haematol.2019.227579 [PubMed]
- 98. Amodio N, Gallo Cantafio ME, Botta C, Agosti V, Federico C, Caracciolo D, Ronchetti D, Rossi M, Driessen C, Neri A, Tagliaferri P, Tassone P. Replacement of miR-155 elicits tumor suppressive activity and antagonizes bortezomib resistance in multiple myeloma. Cancers (Basel). 2019; 11:236. https://doi.org/10.3390/cancers11020236 [PubMed]
- 99. Chen CZ, Li L, Lodish HF, Bartel DP. MicroRNAs modulate hematopoietic lineage differentiation. Science. 2004; 303:83–86. https://doi.org/10.1126/science.1091903 [PubMed]
- 100. Weng H, Lal K, Yang FF, Chen J. The pathological role and prognostic impact of miR-181 in acute myeloid leukemia. Cancer Genet. 2015; 208:225–29. https://doi.org/10.1016/j.cancergen.2014.12.006 [PubMed]
- 101. Das F, Bera A, Ghosh-Choudhury N, Abboud HE, Kasinath BS, Choudhury GG. TGFβ-induced deptor suppression recruits mTORC1 and not mTORC2 to enhance collagen I (α2) gene expression. PLoS One. 2014; 9:e109608. https://doi.org/10.1371/journal.pone.0109608 [PubMed]
- 102. Maity S, Bera A, Ghosh-Choudhury N, Das F, Kasinath BS, Choudhury GG. microRNA-181a downregulates deptor for TGFβ-induced glomerular mesangial cell hypertrophy and matrix protein expression. Exp Cell Res. 2018; 364:5–15. https://doi.org/10.1016/j.yexcr.2018.01.021 [PubMed]
- 103. Liu N, Yang J, Yuan R, Peng J, Liu L, Guo X. Effects of miR-181a on the biological function of multiple myeloma. Oncol Rep. 2019; 42:291–300. https://doi.org/10.3892/or.2019.7160 [PubMed]
- 104. Vasilatou D, Papageorgiou S, Pappa V, Papageorgiou E, Dervenoulas J. The role of microRNAs in normal and Malignant hematopoiesis. Eur J Haematol. 2010; 84:1–16. https://doi.org/10.1111/j.1600-0609.2009.01348.x [PubMed]
- 105. Mraz M, Kipps TJ. MicroRNAs and B cell receptor signaling in chronic lymphocytic leukemia. Leuk Lymphoma. 2013; 54:1836–39. https://doi.org/10.3109/10428194.2013.796055 [PubMed]
- 106. Efanov A, Zanesi N, Nazaryan N, Santanam U, Palamarchuk A, Croce CM, Pekarsky Y. CD5+CD23+ leukemic cell populations in TCL1 transgenic mice show significantly increased proliferation and Akt phosphorylation. Leukemia. 2010; 24:970–75. https://doi.org/10.1038/leu.2010.46 [PubMed]
- 107. Jinlong S, Lin F, Yonghui L, Li Y, Weidong W. Identification of let-7a-2-3p or/and miR-188-5p as prognostic biomarkers in cytogenetically normal acute myeloid leukemia. PLoS One. 2015; 10:e0118099. https://doi.org/10.1371/journal.pone.0118099 [PubMed]
- 108. Lee YG, Kim I, Oh S, Shin DY, Koh Y, Lee KW. Small RNA sequencing profiles of mir-181 and mir-221, the most relevant microRNAs in acute myeloid leukemia. Korean J Intern Med. 2019; 34:178–83. https://doi.org/10.3904/kjim.2017.102 [PubMed]
- 109. Steinhilber J, Bonin M, Walter M, Fend F, Bonzheim I, Quintanilla-Martinez L. Next-generation sequencing identifies deregulation of microRNAs involved in both innate and adaptive immune response in ALK+ ALCL. PLoS One. 2015; 10:e0117780. https://doi.org/10.1371/journal.pone.0117780 [PubMed]
- 110. Chen S, Zheng Y, Zhang S, Jia L, Zhou Y. Promotion effects of miR-375 on the osteogenic differentiation of human adipose-derived mesenchymal stem cells. Stem Cell Reports. 2017; 8:773–86. https://doi.org/10.1016/j.stemcr.2017.01.028 [PubMed]
- 111. Poy MN, Eliasson L, Krutzfeldt J, Kuwajima S, Ma X, Macdonald PE, Pfeffer S, Tuschl T, Rajewsky N, Rorsman P, Stoffel M. A pancreatic islet-specific microRNA regulates insulin secretion. Nature. 2004; 432:226–30. https://doi.org/10.1038/nature03076 [PubMed]
- 112. Ding L, Xu Y, Zhang W, Deng Y, Si M, Du Y, Yao H, Liu X, Ke Y, Si J, Zhou T. MiR-375 frequently downregulated in gastric cancer inhibits cell proliferation by targeting JAK2. Cell Res. 2010; 20:784–93. https://doi.org/10.1038/cr.2010.79 [PubMed]
- 113. Zhang ZW, Men T, Feng RC, Li YC, Zhou D, Teng CB. miR-375 inhibits proliferation of mouse pancreatic progenitor cells by targeting YAP1. Cell Physiol Biochem. 2013; 32:1808–17. https://doi.org/10.1159/000356614 [PubMed]
- 114. Zhou J, Song S, He S, Zhu X, Zhang Y, Yi B, Zhang B, Qin G, Li D. MicroRNA-375 targets PDK1 in pancreatic carcinoma and suppresses cell growth through the Akt signaling pathway. Int J Mol Med. 2014; 33:950–56. https://doi.org/10.3892/ijmm.2014.1638 [PubMed]
- 115. Tatekawa S, Chinen Y, Ri M, Narita T, Shimura Y, Matsumura-Kimoto Y, Tsukamoto T, Kobayashi T, Kawata E, Uoshima N, Taki T, Taniwaki M, Handa H, et al. Epigenetic repression of miR-375 is the dominant mechanism for constitutive activation of the PDPK1/RPS6KA3 signalling axis in multiple myeloma. Br J Haematol. 2017; 178:534–46. https://doi.org/10.1111/bjh.14707 [PubMed]
- 116. Bi L, Zhou B, Li H, He L, Wang C, Wang Z, Zhu L, Chen M, Gao S. A novel miR-375-HOXB3-CDCA3/DNMT3B regulatory circuitry contributes to leukemogenesis in acute myeloid leukemia. BMC Cancer. 2018; 18:182. https://doi.org/10.1186/s12885-018-4097-z [PubMed]
- 117. Prinsloo A, Pool R, Van Niekerk C. Preliminary data on microRNA expression profiles in a group of South African patients diagnosed with chronic myeloid leukaemia. Mol Clin Oncol. 2017; 7:386–90. https://doi.org/10.3892/mco.2017.1319 [PubMed]
- 118. Gebauer N, Kuba J, Senft A, Schillert A, Bernard V, Thorns C. MicroRNA-150 is up-regulated in extranodal marginal zone lymphoma of MALT type. Cancer Genomics Proteomics. 2014; 11:51–56. [PubMed]
- 119. Wang Z, Hong Z, Gao F, Feng W. Upregulation of microRNA-375 is associated with poor prognosis in pediatric acute myeloid leukemia. Mol Cell Biochem. 2013; 383:59–65. https://doi.org/10.1007/s11010-013-1754-z [PubMed]
- 120. Zhang Y, Tong L, Chen S, Wu W, Wang L. Analysis of NFKB2-mediated regulation of mechanisms underlying the development of Hodgkin’s lymphoma. Mol Med Rep. 2018; 17:8129–36. https://doi.org/10.3892/mmr.2018.8911 [PubMed]
- 121. Gutiérrez NC, Sarasquete ME, Misiewicz-Krzeminska I, Delgado M, De Las Rivas J, Ticona FV, Fermiñán E, Martín-Jiménez P, Chillón C, Risueño A, Hernández JM, García-Sanz R, González M, San Miguel JF. Deregulation of microRNA expression in the different genetic subtypes of multiple myeloma and correlation with gene expression profiling. Leukemia. 2010; 24:629–37. https://doi.org/10.1038/leu.2009.274 [PubMed]
- 122. Quwaider D, Corchete LA, Misiewicz-Krzeminska I, Sarasquete ME, Pérez JJ, Krzeminski P, Puig N, Mateos MV, García-Sanz R, Herrero AB, Gutiérrez NC. DEPTOR maintains plasma cell differentiation and favorably affects prognosis in multiple myeloma. J Hematol Oncol. 2017; 10:92. https://doi.org/10.1186/s13045-017-0461-8 [PubMed]
- 123. Li P, Fan JB, Gao Y, Zhang M, Zhang L, Yang N, Zhao X. miR-135b-5p inhibits LPS-induced TNFα production via silencing AMPK phosphatase Ppm1e. Oncotarget. 2016; 7:77978–86. https://doi.org/10.18632/oncotarget.12866 [PubMed]
- 124. Lu M, Huang Y, Sun W, Li P, Li L, Li L. miR-135b-5p promotes gastric cancer progression by targeting CMTM3. Int J Oncol. 2018; 52:589–98. https://doi.org/10.3892/ijo.2017.4222 [PubMed]
- 125. Li ZW, Zhu YR, Zhou XZ, Zhuo BB, Wang XD. microRNA-135b expression silences Ppm1e to provoke AMPK activation and inhibit osteoblastoma cell proliferation. Oncotarget. 2017; 8:26424–33. https://doi.org/10.18632/oncotarget.15477 [PubMed]
- 126. Zhang XH, Xin ZM. MiR-135b-5p inhibits the progression of Malignant melanoma cells by targeting RBX1. Eur Rev Med Pharmacol Sci. 2020; 24:1309–15. https://doi.org/10.26355/eurrev_202002_20188 [PubMed]
- 127. Attia HR, Abdelrahman AH, Ibrahim MH, Eid MM, Eid OM, Sallam MT, El Gammal MM, Kamel MM. Altered expression of MicroRNAs in the bone marrow of multiple myeloma patients and their relationship to cytogenetic aberrations. Curr Pharm Biotechnol. 2020; 21:1394–401. https://doi.org/10.2174/1389201021666200320135139 [PubMed]
- 128. Hao M, Zang M, Zhao L, Deng S, Xu Y, Qi F, An G, Qin Y, Sui W, Li F, Yang W, Li Z, Yi S, et al. Serum high expression of miR-214 and miR-135b as novel predictor for myeloma bone disease development and prognosis. Oncotarget. 2016; 7:19589–600. https://doi.org/10.18632/oncotarget.7319 [PubMed]
- 129. Ho TT, He X, Mo YY, Beck WT. Transient resistance to DNA damaging agents is associated with expression of microRNAs-135b and -196b in human leukemia cell lines. Int J Biochem Mol Biol. 2016; 7:27–47. [PubMed]
- 130. Matsuyama H, Suzuki HI, Nishimori H, Noguchi M, Yao T, Komatsu N, Mano H, Sugimoto K, Miyazono K. miR-135b mediates NPM-ALK-driven oncogenicity and renders IL-17-producing immunophenotype to anaplastic large cell lymphoma. Blood. 2011; 118:6881–92. https://doi.org/10.1182/blood-2011-05-354654 [PubMed]
- 131. Liu C, Iqbal J, Teruya-Feldstein J, Shen Y, Dabrowska MJ, Dybkaer K, Lim MS, Piva R, Barreca A, Pellegrino E, Spaccarotella E, Lachel CM, Kucuk C, et al. MicroRNA expression profiling identifies molecular signatures associated with anaplastic large cell lymphoma. Blood. 2013; 122:2083–92. https://doi.org/10.1182/blood-2012-08-447375 [PubMed]
- 132. Floch P, Capdevielle C, Staedel C, Izotte J, Sifré E, Laur AM, Giese A, Korolik V, Dubus P, Mégraud F, Lehours P. Deregulation of MicroRNAs in gastric lymphomagenesis induced in the d3Tx mouse model of Helicobacter pylori infection. Front Cell Infect Microbiol. 2017; 7:185. https://doi.org/10.3389/fcimb.2017.00185 [PubMed]
- 133. Blosse A, Levy M, Robe C, Staedel C, Copie-Bergman C, Lehours P. Deregulation of miRNA in Helicobacter pylori-induced gastric MALT lymphoma: from mice to human. J Clin Med. 2019; 8:845. https://doi.org/10.3390/jcm8060845 [PubMed]
- 134. Wani S, Kaul D. Tumorigenic PVT-1 gene locus is governed by miR-2909 RNomics. Cell Biochem Funct. 2018; 36:408–12. https://doi.org/10.1002/cbf.3360 [PubMed]
- 135. Piletič K, Kunej T. MicroRNA epigenetic signatures in human disease. Arch Toxicol. 2016; 90:2405–19. https://doi.org/10.1007/s00204-016-1815-7 [PubMed]
- 136. Paydas S, Acikalin A, Ergin M, Celik H, Yavuz B, Tanriverdi K. Micro-RNA (miRNA) profile in Hodgkin lymphoma: association between clinical and pathological variables. Med Oncol. 2016; 33:34. https://doi.org/10.1007/s12032-016-0749-5 [PubMed]
- 137. Xia W, Gong D, Qin X, Cai Z. [MicroRNA-671-3p suppresses proliferation and invasion of breast cancer cells by targeting DEPTOR]. Nan Fang Yi Ke Da Xue Xue Bao. 2020; 40:42–48. https://doi.org/10.12122/j.issn.1673-4254.2020.01.07 [PubMed]
- 138. Yu Y, Wang Z, Sun D, Zhou X, Wei X, Hou W, Ding Y, Ma Y, Hou Y. miR-671 promotes prostate cancer cell proliferation by targeting tumor suppressor SOX6. Eur J Pharmacol. 2018; 823:65–71. https://doi.org/10.1016/j.ejphar.2018.01.016 [PubMed]
- 139. Tan X, Fu Y, Chen L, Lee W, Lai Y, Rezaei K, Tabbara S, Latham P, Teal CB, Man YG, Siegel RS, Brem RF, Fu SW. miR-671-5p inhibits epithelial-to-mesenchymal transition by downregulating FOXM1 expression in breast cancer. Oncotarget. 2016; 7:293–307. https://doi.org/10.18632/oncotarget.6344 [PubMed]
- 140. Xiong DD, Chen H, He RQ, Lan AH, Zhong JC, Chen G, Feng ZB, Wei KL. MicroRNA-671-3p inhibits the development of breast cancer: a study based on in vitro experiments, in-house quantitative polymerase chain reaction and bioinformatics analysis. Int J Oncol. 2018; 52:1801–14. https://doi.org/10.3892/ijo.2018.4339 [PubMed]
- 141. Li ZY, Zhang ZZ, Bi H, Zhang QD, Zhang SJ, Zhou L, Zhu XQ, Zhou J. Upregulated microRNA-671-3p promotes tumor progression by suppressing forkhead box P2 expression in non-small-cell lung cancer. Mol Med Rep. 2019; 20:3149–59. https://doi.org/10.3892/mmr.2019.10563 [PubMed]
- 142. Lu GF, You CY, Chen YS, Jiang H, Zheng X, Tang WW, Wang XY, Xu HY, Geng F. MicroRNA-671-3p promotes proliferation and migration of glioma cells via targeting CKAP4. Onco Targets Ther. 2018; 11:6217–26. https://doi.org/10.2147/OTT.S177325 [PubMed]
- 143. Huang HY, Lin YC, Li J, Huang KY, Shrestha S, Hong HC, Tang Y, Chen YG, Jin CN, Yu Y, Xu JT, Li YM, Cai XX, et al. miRTarBase 2020: updates to the experimentally validated microRNA-target interaction database. Nucleic Acids Res. 2020; 48:D148–54. https://doi.org/10.1093/nar/gkz896 [PubMed]
- 144. Chu L, Yu L, Liu J, Song S, Yang H, Han F, Liu F, Hu Y. Long intergenic non-coding LINC00657 regulates tumorigenesis of glioblastoma by acting as a molecular sponge of miR-190a-3p. Aging (Albany NY). 2019; 11:1456–70. https://doi.org/10.18632/aging.101845 [PubMed]
- 145. Go H, Jang JY, Kim CW, Huh J, Kim PJ, Jeon YK. Identification of microRNAs modulated by DNA hypomethylating drugs in extranodal NK/T-cell lymphoma. Leuk Lymphoma. 2020; 61:66–74. https://doi.org/10.1080/10428194.2019.1654096 [PubMed]
- 146. Yang J, Zhang Z, Chen S, Dou W, Xie R, Gao J. miR-654-3p predicts the prognosis of hepatocellular carcinoma and inhibits the proliferation, migration, and invasion of cancer cells. Cancer Biomark. 2020; 28:73–79. https://doi.org/10.3233/CBM-191084 [PubMed]
- 147. Li P, Cai JX, Han F, Wang J, Zhou JJ, Shen KW, Wang LH. Expression and significance of miR-654-5p and miR-376b-3p in patients with colon cancer. World J Gastrointest Oncol. 2020; 12:492–502. https://doi.org/10.4251/wjgo.v12.i4.492 [PubMed]
- 148. Xu X, Zhuang C, Wu Z, Qiu H, Feng H, Wu J. LincRNA-p21 inhibits cell viability and promotes cell apoptosis in Parkinson’s disease through activating α-synuclein expression. Biomed Res Int. 2018; 2018:8181374. https://doi.org/10.1155/2018/8181374 [PubMed]
- 149. Bai PS, Xia N, Sun H, Kong Y. Pleiotrophin, a target of miR-384, promotes proliferation, metastasis and lipogenesis in HBV-related hepatocellular carcinoma. J Cell Mol Med. 2017; 21:3023–43. https://doi.org/10.1111/jcmm.13213 [PubMed]
- 150. Lai YY, Shen F, Cai WS, Chen JW, Feng JH, Cao J, Xiao HQ, Zhu GH, Xu B. MiR-384 regulated IRS1 expression and suppressed cell proliferation of human hepatocellular carcinoma. Tumour Biol. 2016; 37:14165–71. https://doi.org/10.1007/s13277-016-5233-5 [PubMed]
- 151. Tan Y, Chen L, Li S, Hao H, Zhang D. MiR-384 inhibits Malignant biological behavior such as proliferation and invasion of osteosarcoma by regulating IGFBP3. Technol Cancer Res Treat. 2020; 19:1533033820909125. https://doi.org/10.1177/1533033820909125 [PubMed]
- 152. Wang F. miR-384 targets metadherin gene to suppress growth, migration, and invasion of gastric cancer cells. J Int Med Res. 2019; 47:926–35. https://doi.org/10.1177/0300060518817171 [PubMed]
- 153. Sonneveld P. Management of multiple myeloma in the relapsed/refractory patient. Hematology Am Soc Hematol Educ Program. 2017; 2017:508–17. https://doi.org/10.1182/asheducation-2017.1.508 [PubMed]
- 154. Zhang HR, Chen JM, Zeng ZY, Que WZ. Knockdown of DEPTOR inhibits cell proliferation and increases chemosensitivity to melphalan in human multiple myeloma RPMI-8226 cells via inhibiting PI3K/AKT activity. J Int Med Res. 2013; 41:584–95. https://doi.org/10.1177/0300060513480920 [PubMed]
- 155. de la Rubia J, Such E. DEPTOR expression and response to thalidomide: toward a new therapeutic target in multiple myeloma? Leuk Lymphoma. 2010; 51:1960–61. https://doi.org/10.3109/10428194.2010.527410 [PubMed]
- 156. Shi Y, Daniels-Wells TR, Frost P, Lee J, Finn RS, Bardeleben C, Penichet ML, Jung ME, Gera J, Lichtenstein A. Cytotoxic Properties of a DEPTOR-mTOR Inhibitor in Multiple Myeloma Cells. Cancer Res. 2016; 76:5822–31. https://doi.org/10.1158/0008-5472.CAN-16-1019 [PubMed]
- 157. Todoerti K, Agnelli L, Fabris S, Lionetti M, Tuana G, Mosca L, Lombardi L, Grieco V, Bianchino G, D’Auria F, Statuto T, Mazzoccoli C, De Luca L, et al. Transcriptional characterization of a prospective series of primary plasma cell leukemia revealed signatures associated with tumor progression and poorer outcome. Clin Cancer Res. 2013; 19:3247–58. https://doi.org/10.1158/1078-0432.CCR-12-3461 [PubMed]
- 158. Yang Y, Bardeleben C, Frost P, Hoang B, Shi Y, Finn R, Gera J, Lichtenstein A. DEPTOR is linked to a TORC1-p21 survival proliferation pathway in multiple myeloma cells. Genes Cancer. 2014; 5:407–19. https://doi.org/10.18632/genesandcancer.44 [PubMed]
- 159. Zhang H, Chen J, Zeng Z, Que W, Zhou L. Knockdown of DEPTOR induces apoptosis, increases chemosensitivity to doxorubicin and suppresses autophagy in RPMI-8226 human multiple myeloma cells in vitro. Int J Mol Med. 2013; 31:1127–34. https://doi.org/10.3892/ijmm.2013.1299 [PubMed]
- 160. Hoang B, Shi Y, Frost PJ, Mysore V, Bardeleben C, Lichtenstein A. SGK kinase activity in multiple myeloma cells protects against ER stress apoptosis via a SEK-dependent mechanism. Mol Cancer Res. 2016; 14:397–407. https://doi.org/10.1158/1541-7786.MCR-15-0422 [PubMed]
- 161. Bruneau S, Nakayama H, Woda CB, Flynn EA, Briscoe DM. DEPTOR regulates vascular endothelial cell activation and proinflammatory and angiogenic responses. Blood. 2013; 122:1833–42. https://doi.org/10.1182/blood-2013-03-488486 [PubMed]
- 162. Gao D, Inuzuka H, Tan MK, Fukushima H, Locasale JW, Liu P, Wan L, Zhai B, Chin YR, Shaik S, Lyssiotis CA, Gygi SP, Toker A, et al. mTOR drives its own activation via SCF(βTrCP)-dependent degradation of the mTOR inhibitor DEPTOR. Mol Cell. 2011; 44:290–303. https://doi.org/10.1016/j.molcel.2011.08.030 [PubMed]
- 163. Lee J, Shi Y, Vega M, Yang Y, Gera J, Jung ME, Lichtenstein A. Structure-activity relationship study of small molecule inhibitors of the DEPTOR-mTOR interaction. Bioorg Med Chem Lett. 2017; 27:4714–24. https://doi.org/10.1016/j.bmcl.2017.09.002 [PubMed]
- 164. Jieru W, Jun Q, Li D, Xiaoling L, Chen T, Jiang L, Bicheng Z, Jie Z, Guiyuan L. Identification of LLRC4, a Novel Member of Leucine-rich Repeat (LRR) Superfamily, and Its Expression Analysis in Brain Tumor. Sheng wu hua xue yu Sheng wu wu li jin Zhan. 2002; 29:233–9. https://europepmc.org/article/cba/384792.
- 165. Xu G, Wang R, Wang Z, Lei Q, Yu Z, Liu C, Li P, Yang Z, Cheng X, Li G, Wu M. NGL-2 is a new partner of PAR complex in axon differentiation. J Neurosci. 2015; 35:7153–64. https://doi.org/10.1523/JNEUROSCI.4726-14.2015 [PubMed]
- 166. Li P, Xu G, Li G, Wu M. Function and mechanism of tumor suppressor gene LRRC4/NGL-2. Mol Cancer. 2014; 13:266. https://doi.org/10.1186/1476-4598-13-266 [PubMed]
- 167. Feng J, Zhang Y, Ren X, Li D, Fu H, Liu C, Zhou W, Liu Q, Liu Q, Wu M. Leucine-rich repeat containing 4 act as an autophagy inhibitor that restores sensitivity of glioblastoma to temozolomide. Oncogene. 2020; 39:4551–66. https://doi.org/10.1038/s41388-020-1312-6 [PubMed]
- 168. Tseeleesuren D, Kant R, Yen CH, Hsiao HH, Chen YA. 1,2,3,4,6-Penta- O-Galloyl-Beta-D-Glucopyranoside inhibits proliferation of multiple myeloma cells accompanied with suppression of MYC expression. Front Pharmacol. 2018; 9:65. https://doi.org/10.3389/fphar.2018.00065 [PubMed]
- 169. Liu Y, Chen Y, Wen L, Cui G. Molecular mechanisms underlying the time-dependent autophagy and apoptosis induced by nutrient depletion in multiple myeloma: a pilot study. J Huazhong Univ Sci Technolog Med Sci. 2012; 32:1–8. https://doi.org/10.1007/s11596-012-0001-2 [PubMed]
- 170. Agnelli L, Mosca L, Fabris S, Lionetti M, Andronache A, Kwee I, Todoerti K, Verdelli D, Battaglia C, Bertoni F, Deliliers GL, Neri A. A SNP microarray and FISH-based procedure to detect allelic imbalances in multiple myeloma: an integrated genomics approach reveals a wide gene dosage effect. Genes Chromosomes Cancer. 2009; 48:603–14. https://doi.org/10.1002/gcc.20668 [PubMed]
- 171. Zhan F, Barlogie B, Arzoumanian V, Huang Y, Williams DR, Hollmig K, Pineda-Roman M, Tricot G, van Rhee F, Zangari M, Dhodapkar M, Shaughnessy JD
Jr . Gene-expression signature of benign monoclonal gammopathy evident in multiple myeloma is linked to good prognosis. Blood. 2007; 109:1692–700. https://doi.org/10.1182/blood-2006-07-037077 [PubMed] - 172. Dickens NJ, Walker BA, Leone PE, Johnson DC, Brito JL, Zeisig A, Jenner MW, Boyd KD, Gonzalez D, Gregory WM, Ross FM, Davies FE, Morgan GJ. Homozygous deletion mapping in myeloma samples identifies genes and an expression signature relevant to pathogenesis and outcome. Clin Cancer Res. 2010; 16:1856–64. https://doi.org/10.1158/1078-0432.CCR-09-2831 [PubMed]
- 173. Chapman MA, Lawrence MS, Keats JJ, Cibulskis K, Sougnez C, Schinzel AC, Harview CL, Brunet JP, Ahmann GJ, Adli M, Anderson KC, Ardlie KG, Auclair D, et al. Initial genome sequencing and analysis of multiple myeloma. Nature. 2011; 471:467–72. https://doi.org/10.1038/nature09837 [PubMed]
- 174. Broyl A, Hose D, Lokhorst H, de Knegt Y, Peeters J, Jauch A, Bertsch U, Buijs A, Stevens-Kroef M, Beverloo HB, Vellenga E, Zweegman S, Kersten MJ, et al. Gene expression profiling for molecular classification of multiple myeloma in newly diagnosed patients. Blood. 2010; 116:2543–53. https://doi.org/10.1182/blood-2009-12-261032 [PubMed]
- 175. Zhou Y, Zhang Q, Stephens O, Heuck CJ, Tian E, Sawyer JR, Cartron-Mizeracki MA, Qu P, Keller J, Epstein J, Barlogie B, Shaughnessy JD
Jr . Prediction of cytogenetic abnormalities with gene expression profiles. Blood. 2012; 119:e148–50. https://doi.org/10.1182/blood-2011-10-388702 [PubMed] - 176. Carrasco DR, Tonon G, Huang Y, Zhang Y, Sinha R, Feng B, Stewart JP, Zhan F, Khatry D, Protopopova M, Protopopov A, Sukhdeo K, Hanamura I, et al. High-resolution genomic profiles define distinct clinico-pathogenetic subgroups of multiple myeloma patients. Cancer Cell. 2006; 9:313–25. https://doi.org/10.1016/j.ccr.2006.03.019 [PubMed]
- 177. Mulligan G, Mitsiades C, Bryant B, Zhan F, Chng WJ, Roels S, Koenig E, Fergus A, Huang Y, Richardson P, Trepicchio WL, Broyl A, Sonneveld P, et al. Gene expression profiling and correlation with outcome in clinical trials of the proteasome inhibitor bortezomib. Blood. 2007; 109:3177–88. https://doi.org/10.1182/blood-2006-09-044974 [PubMed]
- 178. Paganin M, Ferrando A. Molecular pathogenesis and targeted therapies for NOTCH1-induced T-cell acute lymphoblastic leukemia. Blood Rev. 2011; 25:83–90. https://doi.org/10.1016/j.blre.2010.09.004 [PubMed]
- 179. Palomero T, Dominguez M, Ferrando AA. The role of the PTEN/AKT pathway in NOTCH1-induced leukemia. Cell Cycle. 2008; 7:965–70. https://doi.org/10.4161/cc.7.8.5753 [PubMed]
- 180. Carbonneau M, M Gagné L, Lalonde ME, Germain MA, Motorina A, Guiot MC, Secco B, Vincent EE, Tumber A, Hulea L, Bergeman J, Oppermann U, Jones RG, et al. The oncometabolite 2-hydroxyglutarate activates the mTOR signalling pathway. Nat Commun. 2016; 7:12700. https://doi.org/10.1038/ncomms12700 [PubMed]
- 181. Tsukada Y, Fang J, Erdjument-Bromage H, Warren ME, Borchers CH, Tempst P, Zhang Y. Histone demethylation by a family of JmjC domain-containing proteins. Nature. 2006; 439:811–6. https://doi.org/10.1038/nature04433 [PubMed]
- 182. Liu S, Wan J, Kong Y, Zhang Y, Wan L, Zhang Z. Inhibition of CRL-NEDD8 pathway as a new approach to enhance ATRA-induced differentiation of acute promyelocytic leukemia cells. Int J Med Sci. 2018; 15:674–81. https://doi.org/10.7150/ijms.23782 [PubMed]
- 183. Haferlach T, Kohlmann A, Wieczorek L, Basso G, Kronnie GT, Béné MC, De Vos J, Hernández JM, Hofmann WK, Mills KI, Gilkes A, Chiaretti S, Shurtleff SA, et al. Clinical utility of microarray-based gene expression profiling in the diagnosis and subclassification of leukemia: report from the international microarray innovations in leukemia study group. J Clin Oncol. 2010; 28:2529–37. https://doi.org/10.1200/JCO.2009.23.4732 [PubMed]
- 184. Andersson A, Ritz C, Lindgren D, Edén P, Lassen C, Heldrup J, Olofsson T, Råde J, Fontes M, Porwit-Macdonald A, Behrendtz M, Höglund M, Johansson B, Fioretos T. Microarray-based classification of a consecutive series of 121 childhood acute leukemias: prediction of leukemic and genetic subtype as well as of minimal residual disease status. Leukemia. 2007; 21:1198–203. https://doi.org/10.1038/sj.leu.2404688 [PubMed]
- 185. Valk PJ, Verhaak RG, Beijen MA, Erpelinck CA, Barjesteh van Waalwijk van Doorn-Khosrovani S, Boer JM, Beverloo HB, Moorhouse MJ, van der Spek PJ, Löwenberg B, Delwel R. Prognostically useful gene-expression profiles in acute myeloid leukemia. N Engl J Med. 2004; 350:1617–28. https://doi.org/10.1056/NEJMoa040465 [PubMed]
- 186. Gutiérrez NC, López-Pérez R, Hernández JM, Isidro I, González B, Delgado M, Fermiñán E, García JL, Vázquez L, González M, San Miguel JF. Gene expression profile reveals deregulation of genes with relevant functions in the different subclasses of acute myeloid leukemia. Leukemia. 2005; 19:402–09. https://doi.org/10.1038/sj.leu.2403625 [PubMed]
- 187. Metzeler KH, Hummel M, Bloomfield CD, Spiekermann K, Braess J, Sauerland MC, Heinecke A, Radmacher M, Marcucci G, Whitman SP, Maharry K, Paschka P, Larson RA, et al, Cancer and Leukemia Group B, and German AML Cooperative Group. An 86-probe-set gene-expression signature predicts survival in cytogenetically normal acute myeloid leukemia. Blood. 2008; 112:4193–201. https://doi.org/10.1182/blood-2008-02-134411 [PubMed]
- 188. Heuser M, Wingen LU, Steinemann D, Cario G, von Neuhoff N, Tauscher M, Bullinger L, Krauter J, Heil G, Döhner H, Schlegelberger B, Ganser A. Gene-expression profiles and their association with drug resistance in adult acute myeloid leukemia. Haematologica. 2005; 90:1484–92. [PubMed]
- 189. Raponi M, Harousseau JL, Lancet JE, Löwenberg B, Stone R, Zhang Y, Rackoff W, Wang Y, Atkins D. Identification of molecular predictors of response in a study of tipifarnib treatment in relapsed and refractory acute myelogenous leukemia. Clin Cancer Res. 2007; 13:2254–60. https://doi.org/10.1158/1078-0432.CCR-06-2609 [PubMed]
- 190. Raponi M, Lancet JE, Fan H, Dossey L, Lee G, Gojo I, Feldman EJ, Gotlib J, Morris LE, Greenberg PL, Wright JJ, Harousseau JL, Löwenberg B, et al. A 2-gene classifier for predicting response to the farnesyltransferase inhibitor tipifarnib in acute myeloid leukemia. Blood. 2008; 111:2589–96. https://doi.org/10.1182/blood-2007-09-112730 [PubMed]
- 191. Balgobind BV, Van den Heuvel-Eibrink MM, De Menezes RX, Reinhardt D, Hollink IH, Arentsen-Peters ST, van Wering ER, Kaspers GJ, Cloos J, de Bont ES, Cayuela JM, Baruchel A, Meyer C, et al. Evaluation of gene expression signatures predictive of cytogenetic and molecular subtypes of pediatric acute myeloid leukemia. Haematologica. 2011; 96:221–30. https://doi.org/10.3324/haematol.2010.029660 [PubMed]
- 192. Kühn MW, Radtke I, Bullinger L, Goorha S, Cheng J, Edelmann J, Gohlke J, Su X, Paschka P, Pounds S, Krauter J, Ganser A, Quessar A, et al. High-resolution genomic profiling of adult and pediatric core-binding factor acute myeloid leukemia reveals new recurrent genomic alterations. Blood. 2012; 119:e67–75. https://doi.org/10.1182/blood-2011-09-380444 [PubMed]
- 193. Döhner K, Paschka P, Döhner H. [Acute myeloid leukemia]. Internist (Berl). 2015; 56:354–63. https://doi.org/10.1007/s00108-014-3596-5 [PubMed]
- 194. Horowitz M, Schreiber H, Elder A, Heidenreich O, Vormoor J, Toffalori C, Vago L, Kröger N. Epidemiology and biology of relapse after stem cell transplantation. Bone Marrow Transplant. 2018; 53:1379–89. https://doi.org/10.1038/s41409-018-0171-z [PubMed]
- 195. Dik WA, Brahim W, Braun C, Asnafi V, Dastugue N, Bernard OA, van Dongen JJ, Langerak AW, Macintyre EA, Delabesse E. CALM-AF10+ T-ALL expression profiles are characterized by overexpression of HOXA and BMI1 oncogenes. Leukemia. 2005; 19:1948–57. https://doi.org/10.1038/sj.leu.2403891 [PubMed]
- 196. Coustan-Smith E, Song G, Clark C, Key L, Liu P, Mehrpooya M, Stow P, Su X, Shurtleff S, Pui CH, Downing JR, Campana D. New markers for minimal residual disease detection in acute lymphoblastic leukemia. Blood. 2011; 117:6267–76. https://doi.org/10.1182/blood-2010-12-324004 [PubMed]
- 197. Bhojwani D, Kang H, Moskowitz NP, Min DJ, Lee H, Potter JW, Davidson G, Willman CL, Borowitz MJ, Belitskaya-Levy I, Hunger SP, Raetz EA, Carroll WL. Biologic pathways associated with relapse in childhood acute lymphoblastic leukemia: a children’s oncology group study. Blood. 2006; 108:711–17. https://doi.org/10.1182/blood-2006-02-002824 [PubMed]
- 198. Peirs S, Van der Meulen J, Van de Walle I, Taghon T, Speleman F, Poppe B, Van Vlierberghe P. Epigenetics in T-cell acute lymphoblastic leukemia. Immunol Rev. 2015; 263:50–67. https://doi.org/10.1111/imr.12237 [PubMed]
- 199. Pui CH. Central nervous system disease in acute lymphoblastic leukemia: prophylaxis and treatment. Hematology Am Soc Hematol Educ Program. 2006:142–6. https://doi.org/10.1182/asheducation-2006.1.142 [PubMed]
- 200. Allen A, Sireci A, Colovai A, Pinkney K, Sulis M, Bhagat G, Alobeid B. Early T-cell precursor leukemia/lymphoma in adults and children. Leuk Res. 2013; 37:1027–34. https://doi.org/10.1016/j.leukres.2013.06.010 [PubMed]
- 201. Dores GM, Devesa SS, Curtis RE, Linet MS, Morton LM. Acute leukemia incidence and patient survival among children and adults in the United States, 2001-2007. Blood. 2012; 119:34–43. https://doi.org/10.1182/blood-2011-04-347872 [PubMed]
- 202. Meani N, Minardi S, Licciulli S, Gelmetti V, Coco FL, Nervi C, Pelicci PG, Müller H, Alcalay M. Molecular signature of retinoic acid treatment in acute promyelocytic leukemia. Oncogene. 2005; 24:3358–68. https://doi.org/10.1038/sj.onc.1208498 [PubMed]
- 203. Chiorazzi N, Rai KR, Ferrarini M. Chronic lymphocytic leukemia. N Engl J Med. 2005; 352:804–15. https://doi.org/10.1056/NEJMra041720 [PubMed]
- 204. Kipps TJ, Stevenson FK, Wu CJ, Croce CM, Packham G, Wierda WG, O'Brien S, Gribben J, Rai K. Chronic lymphocytic leukaemia. Nat Rev Dis Primers. 2017; 3:16096. https://doi.org/10.1038/nrdp.2016.96 [PubMed]
- 205. Bosch F, Dalla-Favera R. Chronic lymphocytic leukaemia: from genetics to treatment. Nat Rev Clin Oncol. 2019; 16:684–701. https://doi.org/10.1038/s41571-019-0239-8 [PubMed]
- 206. Wang P, Zhang Q, Tan L, Xu Y, Xie X, Zhao Y. The regulatory effects of mTOR complexes in the differentiation and function of CD4+ T cell subsets. J Immunol Res. 2020; 2020:3406032. https://doi.org/10.1155/2020/3406032 [PubMed]
- 207. Wedel J, Nakayama H, Kochupurakkal NM, Koch J, Klagsbrun M, Bielenberg DR, Briscoe DM. The intragraft microenvironment as a central determinant of chronic rejection or local immunoregulation/tolerance. Curr Opin Organ Transplant. 2017; 22:55–63. https://doi.org/10.1097/MOT.0000000000000373 [PubMed]
- 208. Compagno M, Lim WK, Grunn A, Nandula SV, Brahmachary M, Shen Q, Bertoni F, Ponzoni M, Scandurra M, Califano A, Bhagat G, Chadburn A, Dalla-Favera R, Pasqualucci L. Mutations of multiple genes cause deregulation of NF-kappaB in diffuse large B-cell lymphoma. Nature. 2009; 459:717–21. https://doi.org/10.1038/nature07968 [PubMed]
- 209. Hummel M, Bentink S, Berger H, Klapper W, Wessendorf S, Barth TF, Bernd HW, Cogliatti SB, Dierlamm J, Feller AC, Hansmann ML, Haralambieva E, Harder L, et al, and Molecular Mechanisms in Malignant Lymphomas Network Project of the Deutsche Krebshilfe. A biologic definition of Burkitt’s lymphoma from transcriptional and genomic profiling. N Engl J Med. 2006; 354:2419–30. https://doi.org/10.1056/NEJMoa055351 [PubMed]
- 210. Jais JP, Haioun C, Molina TJ, Rickman DS, de Reynies A, Berger F, Gisselbrecht C, Brière J, Reyes F, Gaulard P, Feugier P, Labouyrie E, Tilly H, et al, and Groupe d’Etude des Lymphomes de l’Adulte. The expression of 16 genes related to the cell of origin and immune response predicts survival in elderly patients with diffuse large B-cell lymphoma treated with CHOP and rituximab. Leukemia. 2008; 22:1917–24. https://doi.org/10.1038/leu.2008.188 [PubMed]
- 211. Brune V, Tiacci E, Pfeil I, Döring C, Eckerle S, van Noesel CJ, Klapper W, Falini B, von Heydebreck A, Metzler D, Bräuninger A, Hansmann ML, Küppers R. Origin and pathogenesis of nodular lymphocyte-predominant hodgkin lymphoma as revealed by global gene expression analysis. J Exp Med. 2008; 205:2251–68. https://doi.org/10.1084/jem.20080809 [PubMed]
- 212. Storz MN, van de Rijn M, Kim YH, Mraz-Gernhard S, Hoppe RT, Kohler S. Gene expression profiles of cutaneous B cell lymphoma. J Invest Dermatol. 2003; 120:865–70. https://doi.org/10.1046/j.1523-1747.2003.12142.x [PubMed]
- 213. Barretina J, Caponigro G, Stransky N, Venkatesan K, Margolin AA, Kim S, Wilson CJ, Lehár J, Kryukov GV, Sonkin D, Reddy A, Liu M, Murray L, et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature. 2012; 483:603–7. https://doi.org/10.1038/nature11003 [PubMed]
- 214. A clinical evaluation of the international lymphoma study group classification of non-Hodgkin’s lymphoma. The non-Hodgkin’s lymphoma classification project. Blood. 1997; 89:3909–18. [PubMed]
- 215. Alizadeh AA, Eisen MB, Davis RE, Ma C, Lossos IS, Rosenwald A, Boldrick JC, Sabet H, Tran T, Yu X, Powell JI, Yang L, Marti GE, et al. Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature. 2000; 403:503–11. https://doi.org/10.1038/35000501 [PubMed]
- 216. Vitolo U, Trněný M, Belada D, Burke JM, Carella AM, Chua N, Abrisqueta P, Demeter J, Flinn I, Hong X, Kim WS, Pinto A, Shi YK, et al. Obinutuzumab or rituximab plus cyclophosphamide, doxorubicin, vincristine, and prednisone in previously untreated diffuse large B-cell lymphoma. J Clin Oncol. 2017; 35:3529–37. https://doi.org/10.1200/JCO.2017.73.3402 [PubMed]
- 217. Lenz G, Wright G, Dave SS, Xiao W, Powell J, Zhao H, Xu W, Tan B, Goldschmidt N, Iqbal J, Vose J, Bast M, Fu K, et al, and Lymphoma/Leukemia Molecular Profiling Project. Stromal gene signatures in large-B-cell lymphomas. N Engl J Med. 2008; 359:2313–23. https://doi.org/10.1056/NEJMoa0802885 [PubMed]
- 218. Shaknovich R, Geng H, Johnson NA, Tsikitas L, Cerchietti L, Greally JM, Gascoyne RD, Elemento O, Melnick A. DNA methylation signatures define molecular subtypes of diffuse large B-cell lymphoma. Blood. 2010; 116:e81–89. https://doi.org/10.1182/blood-2010-05-285320 [PubMed]
- 219. Luminari S, Bellei M, Biasoli I, Federico M. Follicular lymphoma - treatment and prognostic factors. Rev Bras Hematol Hemoter. 2012; 34:54–9. https://doi.org/10.5581/1516-8484.20120015 [PubMed]
- 220. Dave SS, Wright G, Tan B, Rosenwald A, Gascoyne RD, Chan WC, Fisher RI, Braziel RM, Rimsza LM, Grogan TM, Miller TP, LeBlanc M, Greiner TC, et al. Prediction of survival in follicular lymphoma based on molecular features of tumor-infiltrating immune cells. N Engl J Med. 2004; 351:2159–69. https://doi.org/10.1056/NEJMoa041869 [PubMed]
- 221. Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011; 144:646–74. https://doi.org/10.1016/j.cell.2011.02.013 [PubMed]
- 222. Masqué-Soler N, Szczepanowski M, Kohler CW, Aukema SM, Nagel I, Richter J, Siebert R, Spang R, Burkhardt B, Klapper W. Clinical and pathological features of Burkitt lymphoma showing expression of BCL2—an analysis including gene expression in formalin-fixed paraffin-embedded tissue. Br J Haematol. 2015; 171:501–08. https://doi.org/10.1111/bjh.13624 [PubMed]
- 223. Klanova M, Klener P. BCL-2 proteins in pathogenesis and therapy of B-cell non-hodgkin lymphomas. Cancers (Basel). 2020; 12:938. https://doi.org/10.3390/cancers12040938 [PubMed]
- 224. Hartmann EM, Ott G, Rosenwald A. Molecular outcome prediction in mantle cell lymphoma. Future Oncol. 2009; 5:63–73. https://doi.org/10.2217/14796694.5.1.63 [PubMed]
- 225. Eyre TA, Khan D, Hall GW, Collins GP. Anaplastic lymphoma kinase-positive anaplastic large cell lymphoma: current and future perspectives in adult and paediatric disease. Eur J Haematol. 2014; 93:455–68. https://doi.org/10.1111/ejh.12360 [PubMed]
- 226. Leventaki V, Bhattacharyya S, Lim MS. Pathology and genetics of anaplastic large cell lymphoma. Semin Diagn Pathol. 2020; 37:57–71. https://doi.org/10.1053/j.semdp.2019.12.002 [PubMed]
- 227. Eckerle S, Brune V, Döring C, Tiacci E, Bohle V, Sundström C, Kodet R, Paulli M, Falini B, Klapper W, Chaubert AB, Willenbrock K, Metzler D, et al. Gene expression profiling of isolated tumour cells from anaplastic large cell lymphomas: insights into its cellular origin, pathogenesis and relation to Hodgkin lymphoma. Leukemia. 2009; 23:2129–38. https://doi.org/10.1038/leu.2009.161 [PubMed]
- 228. Iqbal J, Weisenburger DD, Greiner TC, Vose JM, McKeithan T, Kucuk C, Geng H, Deffenbacher K, Smith L, Dybkaer K, Nakamura S, Seto M, Delabie J, et al, and International Peripheral T-Cell Lymphoma Project. Molecular signatures to improve diagnosis in peripheral T-cell lymphoma and prognostication in angioimmunoblastic T-cell lymphoma. Blood. 2010; 115:1026–36. https://doi.org/10.1182/blood-2009-06-227579 [PubMed]
- 229. Shimoyama M. Diagnostic criteria and classification of clinical subtypes of adult T-cell leukaemia-lymphoma. A report from the lymphoma study group (1984-87). Br J Haematol. 1991; 79:428–37. https://doi.org/10.1111/j.1365-2141.1991.tb08051.x [PubMed]
- 230. Ishitsuka K, Tamura K. Human T-cell leukaemia virus type I and adult T-cell leukaemia-lymphoma. Lancet Oncol. 2014; 15:e517–26. https://doi.org/10.1016/S1470-2045(14)70202-5 [PubMed]
- 231. Timmins MA, Wagner SD, Ahearne MJ. The new biology of PTCL-NOS and AITL: current status and future clinical impact. Br J Haematol. 2020; 189:54–66. https://doi.org/10.1111/bjh.16428 [PubMed]
- 232. Wang HW, Balakrishna JP, Pittaluga S, Jaffe ES. Diagnosis of hodgkin lymphoma in the modern era. Br J Haematol. 2019; 184:45–59. https://doi.org/10.1111/bjh.15614 [PubMed]
- 233. Shanbhag S, Ambinder RF. Hodgkin lymphoma: a review and update on recent progress. CA Cancer J Clin. 2018; 68:116–32. https://doi.org/10.3322/caac.21438 [PubMed]
- 234. Steidl C, Lee T, Shah SP, Farinha P, Han G, Nayar T, Delaney A, Jones SJ, Iqbal J, Weisenburger DD, Bast MA, Rosenwald A, Muller-Hermelink HK, et al. Tumor-associated macrophages and survival in classic Hodgkin's lymphoma. N Engl J Med. 2010; 362:875–85. https://doi.org/10.1056/NEJMoa0905680 [PubMed]
- 235. Chetaille B, Bertucci F, Finetti P, Esterni B, Stamatoullas A, Picquenot JM, Copin MC, Morschhauser F, Casasnovas O, Petrella T, Molina T, Vekhoff A, Feugier P, et al. Molecular profiling of classical Hodgkin lymphoma tissues uncovers variations in the tumor microenvironment and correlations with EBV infection and outcome. Blood. 2009; 113:2765–3775. https://doi.org/10.1182/blood-2008-07-168096 [PubMed]


