Introduction
Werner syndrome (WS) is a rare autosomal recessive premature aging disorder that begins at a young age with graying and loss of hair and cataracts, followed by accelerated aging symptoms such as diabetes, atherosclerosis, and cancer [1–4]. The median life expectancy is in the mid-50s, and most deaths are due to arteriosclerosis and malignancy [5]. Owing to the founder mutation, a high incidence of WS is observed in Japan [6, 7].
The causative gene is WRN, which is located on chromosome 8 and is involved in DNA replication, DNA repair, and telomere maintenance [8]. WS fibroblasts with deficient or dysfunctional WRN proteins show premature cellular senescence in vitro [9]. This phenotype is largely dependent on telomere shortening and can be overcome by telomerase overexpression [10, 11].
WS mimics various symptoms of general aging. However, there are also phenotypes specific to WS, such as refractory skin ulcers with severe pain in the extremities, which affect over 80% of patients and may even result in limb amputation [12]. Common sites for ulceration are the heels, soles, toes, Achilles tendons, and elbows. Painful subcutaneous calcification has been reported to precede skin ulcers [13]. The atrophy of subcutaneous fat and muscle when present in the extremities resembles branches of dried trees, which is diagnosed as sarcopenia [14]. In contrast, there is an accumulation of subcutaneous fat in the trunk [15, 16]. While the skin of the extremities, frequently accompanied by ulcers, is atrophic and tight, that of the trunk retains its elasticity and does not develop ulcers. The underlying mechanism behind these differences remains unknown.
To clarify the relationship between the skin properties and the high prevalence of skin ulcers in the extremities, we established fibroblasts from the skin of the trunk and that of the foot from the vicinity of ulceration in two WS patients.
Results
Foot fibroblasts exhibited reduced proliferation compared to the trunk fibroblasts in a telomere-dependent manner
In WS, the skin in the extremities atrophies and hardens, and a skin ulcer develops, while there is no obvious abnormality in the trunk skin. Therefore, plastic surgery is often performed to graft skin from the trunk to the ulcer site in order to treat the skin ulcers. In this study, we established fibroblasts from the foot skin (normal skin adjacent to the ulcer) and trunk skin (graft) of two WS patients (WS1 and WS2) who were admitted to our hospital for plastic surgery. We hypothesized that the difference in skin symptoms between the limb and trunk was related to a reduced proliferative capacity of the limb fibroblasts compared to that of the trunk. As expected, the proliferation rate of foot skin-derived fibroblasts was lower than that of the trunk (Figure 1A). In previous reports, the cause of the reduced proliferative potential of WS fibroblasts was explained by shortened telomere length [10, 11]. Consistent with these reports, there was a significant difference in the telomere lengths between the foot and trunk fibroblasts (Figure 1B, 1C and Supplementary Figure 1). These results suggest that the difference in proliferation ability between skin fibroblasts of the foot and trunk is dependent on the telomere length.
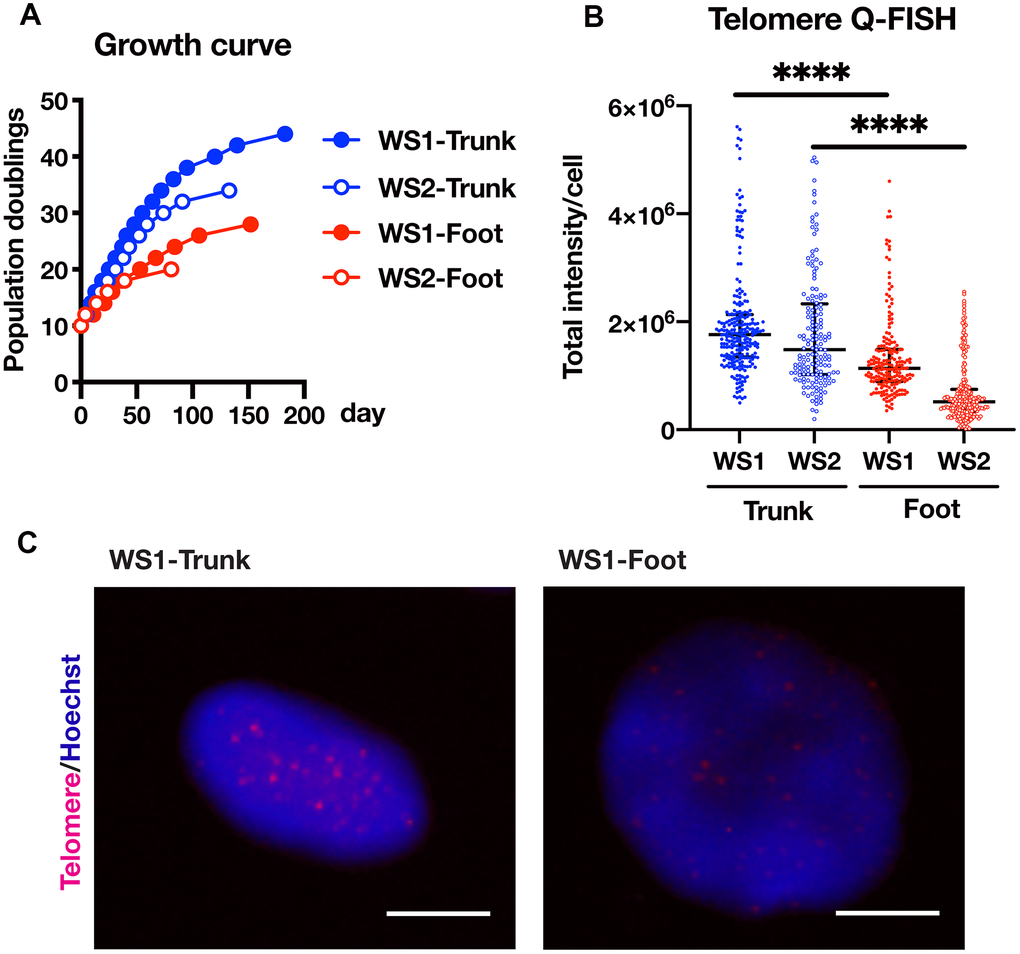
Figure 1. Foot fibroblasts exhibited reduced proliferative capacity compared to that from the trunk in a telomere length-dependent manner. (A) Growth curves of the fibroblasts from the trunk and foot in two WS patients. (B) Telomere length quantification through Q-FISH. Data are median values ± interquartile range of each cell line. More than 150 cells for each cell line were analyzed. For statistical analysis, Mann Whitney test was performed (****p<0.0001). (C) Representative image of telomere Q-FISH of WS1. Bar = 10 μm.
Foot and trunk fibroblasts in WS showed differential expression of genes, especially those involved in embryogenesis and differentiation
Next, transcriptome analysis was performed using RNA sequences to characterize the gene expression profiles of the foot and trunk fibroblasts. In the hierarchical clustering analysis, the WS foot and trunk fibroblasts were classified into different clusters beyond individual differences (Supplementary Figure 2A and Supplementary Table 1). The analysis of differentially expressed genes (DEGs) was performed to identify genes with more than 2-fold differences in expression, and a total of 140 up-regulated and 119 down-regulated genes were identified in the foot (Figure 2A and Supplementary Tables 2, 3). Enrichment analysis of DEGs revealed that their pathways are mainly involved in differentiation and embryogenesis (Figure 2B). Among them, Homeobox A13 (HOXA13) was explicitly expressed in the foot, while Homeobox B5 (HOXB5), Homeobox B6 (HOXB6), Homeobox B7 (HOXB7), and Homeobox D4 (HOXD4) expression were specific to the trunk (Figure 2C), which is consistent with the site-specific gene expression of fibroblasts from normal individuals in previous reports [17, 18]. Intriguingly, the foot fibroblasts showed an elevated expression of genes that relate to the promotion of osteogenesis and suppression of adipogenesis and chondrogenesis, including Stathmin 2 (STMN2), Copine 7 (CPNE7), Protein Tyrosine Phosphatase Receptor Type B (PTPRB), Solute Carrier Family 2 Member 5 (SLC2A5), and HOXA Distal Transcript Antisense RNA (HOTTIP) (Figure 2C) [19–23]. In contrast, in the fibroblasts of the trunk, the expression of the following genes were increased; Clusterin (CLU), Peroxisome Proliferator-Activated Receptor Gamma (PPARG), Insulin-Like Growth Factor 2 MRNA Binding Protein 3 (IGF2BP3), Cysteine Dioxygenase Type 1 (CDO1), and Zinc Finger Protein, FOG Family Member 2 (ZFPM2) (Figure 2C), which are associated with the promotion of adipogenesis or chondrogenesis and suppression of osteogenesis [24–29]. However, regarding senescence-associated genes, no significant differences were observed in the expression of Cyclin-Dependent Kinase Inhibitor 1A (CDKN1A, p21) and Cyclin-Dependent Kinase Inhibitor 2A (CDKN2A, p16) (Supplementary Figure 2B) [30, 31]. These results suggest that the foot and trunk fibroblasts in WS have distinct gene expressions that regulate mesenchymal-lineage differentiation, but these differences are independent of cellular senescence.
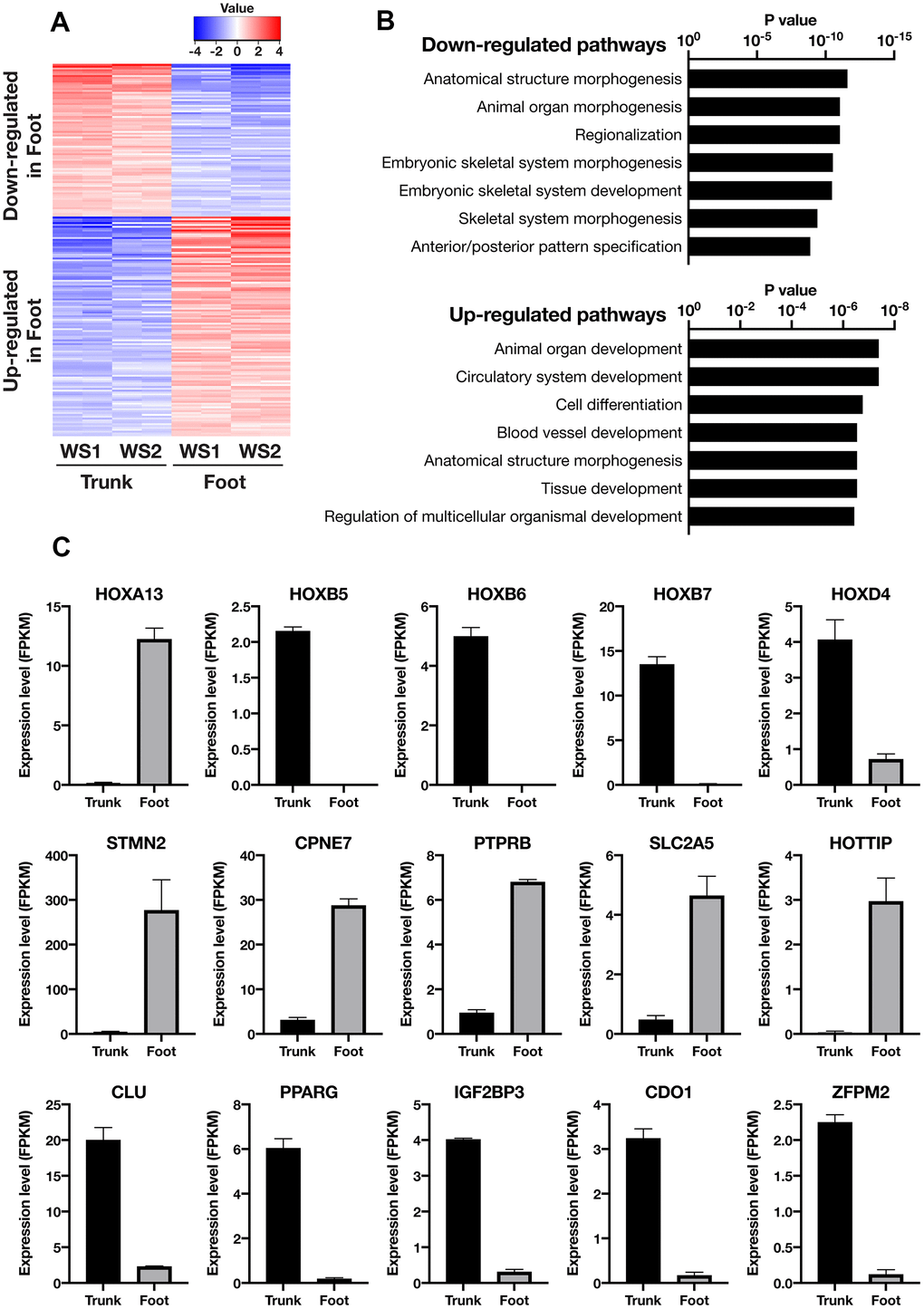
Figure 2. Transcriptome analysis showed distinct gene expression profiles between the trunk and foot fibroblasts. (A) Heatmap of differentially expressed genes between the trunk and foot. Cutoff: |log2(Foot/Trunk)| > 1 and FDR < 0.05. (B) List of the top seven Gene Ontology (GO: biological process) terms and corresponding p values related to Figure 2A. (C) Differentially expressed genes specifically involved in embryonic development and mesenchymal cell differentiation. Data are means ± SEM of two patients (technically n=2 in each sample).
Foot fibroblasts in WS were less capable of adipogenesis compared with the trunk
WS patients present with subcutaneous lipoatrophy in the extremities, while subcutaneous fat tends to accumulate in the trunk. Several reports have previously shown that human dermal fibroblasts can differentiate into mesodermal lineages in vitro, including adipocytes [32–34]. Taken together with the above results, we hypothesized that the adipogenesis potential is lower in foot fibroblasts than in the trunk. Thus, we investigated the adipogenic capacity of WS fibroblasts by culturing them in the adipogenic differentiation medium. After induction of adipogenesis, Oil red O staining results showed foot fibroblasts with significantly fewer oil droplets than the trunk (Figure 3A, 3B and Supplementary Figure 3). In gene expression analysis by qRT-PCR, adipocyte marker genes, PPARG, Fatty Acid Binding Protein 4 (FABP4), CCAAT Enhancer Binding Protein Alpha (CEBPA), and Leptin (LEP) were significantly decreased in the foot group compared with the trunk (Figure 3C) [35]. These results indicate that the trunk fibroblasts of WS maintain adipogenic capacity but the foot fibroblasts do not.
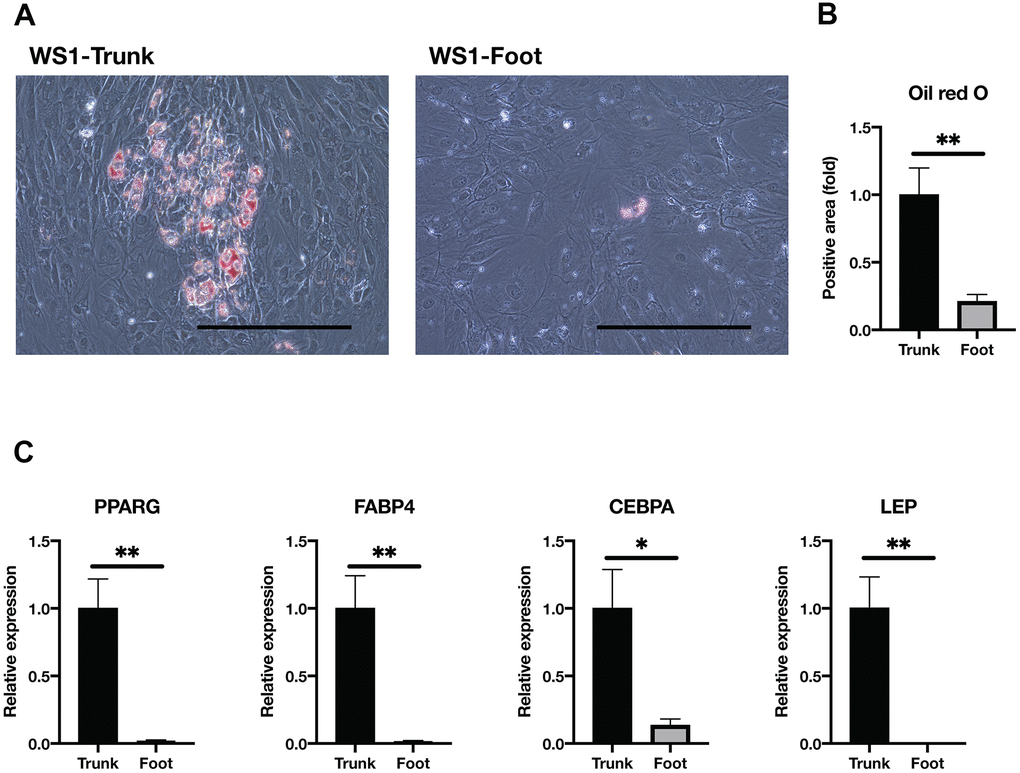
Figure 3. Adipogenesis was impaired in the foot fibroblasts. (A) Representative images of Oil red O staining after two weeks induction of adipogenesis in the trunk and foot fibroblasts of WS1. Bar = 300 μm. (B) Quantification of relative Oil red O staining area. Data are means ± SEM of two patients from four microscopic views. For statistical analysis, student t-test was performed (**p<0.01). (C) Relative gene expression analyzed by qRT-PCR. Data are means ± SEM of two patients with three technical replicates. For statistical analysis, student t-test was performed (*p<0.05; **p<0.01).
Foot fibroblasts in WS exhibited an attenuated capacity for chondrogenesis
Next, we performed chondrogenic differentiation to confirm a disparity in chondrogenesis between the trunk and foot fibroblasts. After the induction of chondrogenesis using the pellet method, the spheroid diameter was significantly smaller in the foot fibroblasts from WS1 than in the trunk group (Figure 4A, 4B). On the other hand, WS2 foot fibroblasts failed to maintain spheroid morphology (Supplementary Figure 4). The chondrogenesis differentiation marker, SRY-Box Transcription Factor 9 (SOX9), was significantly decreased in the foot group, and this tendency was also observed for Aggrecan (ACAN) (Figure 4C) [36]. These results suggest that WS foot fibroblasts tend to have a reduced capacity for chondrogenic differentiation compared with the trunk.
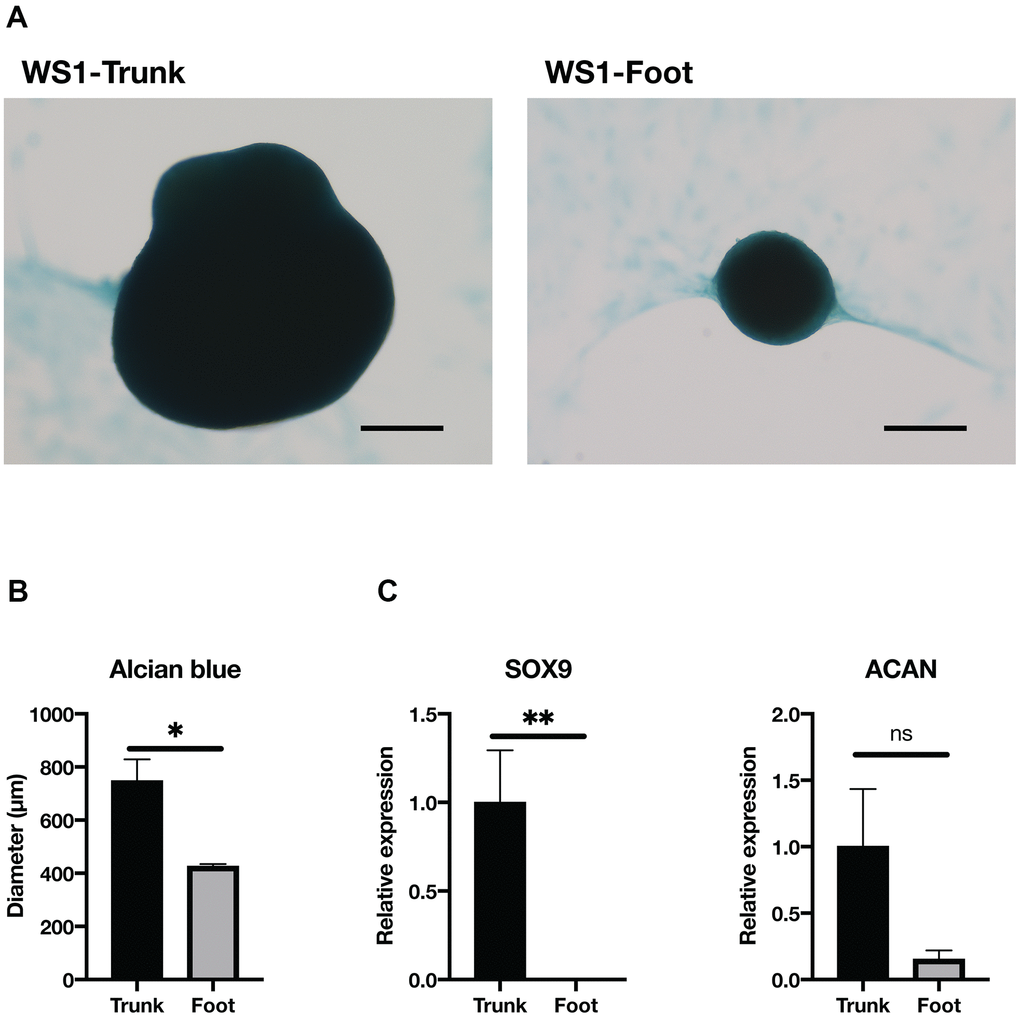
Figure 4. Chondrogenesis tended to be reduced in the foot fibroblasts. (A) Representative images of Alcian blue staining after two weeks induction of chondrogenesis in the trunk and foot fibroblasts of WS1. Bar = 300 μm. (B) Quantification of the diameter of chondrogenic spheroids. WS1-trunk, WS2-trunk, and WS1-foot were included. Data are means ± SEM. For statistical analysis, student t-test was performed (*p<0.05). (C) Relative gene expression analyzed by qRT-PCR. Data are means ± SEM of two patients with three technical replicates. For statistical analysis, student t-test was performed (ns, not significant; *p<0.05; **p<0.01).
Foot and trunk fibroblasts in WS were equally capable of osteogenesis
Next, the osteogenic differentiation ability was compared. After culturing in the osteogenesis medium, no clear difference was observed in the Alizarin red-stained area between the trunk and foot groups (Figure 5A, 5B and Supplementary Figure 5). The expression of Alkaline Phosphatase, Biomineralization Associated (ALPL), a marker of osteogenesis, was significantly elevated in the foot group (Figure 5C). On the other hand, RUNX Family Transcription Factor 2 (RUNX2) expression was significantly elevated in the trunk group, and there were no significant differences in other differentiation markers (Secreted Phosphoprotein 1, SPP1; Collagen Type I Alpha 1 Chain, COL1A1) (Figure 5C) [37]. These results suggest that foot fibroblasts in WS maintain the equivalent level of osteogenic differentiation capacity to the trunk.
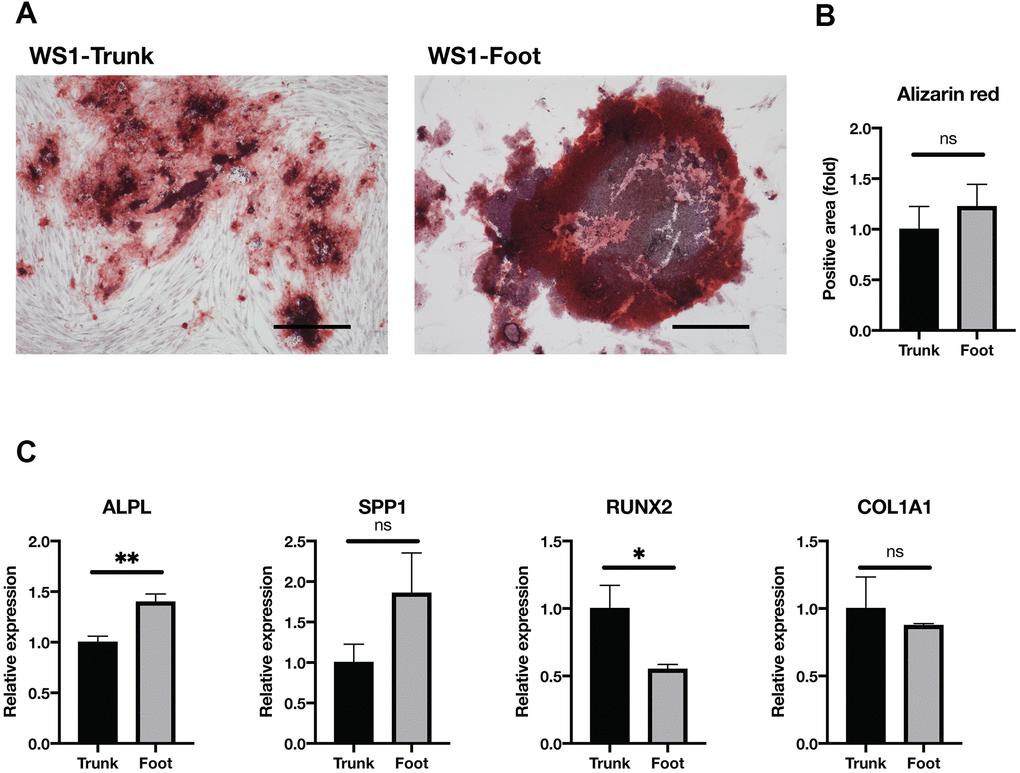
Figure 5. Osteogenesis was maintained in both groups. (A) Representative images of Alizarin red staining two weeks after induction of osteogenesis in the trunk and foot fibroblasts of WS1. Bar = 300 μm. (B) Quantification of relative stained area with Alizarin red. Data are means ± SEM of two patients with four microscopic views. For statistical analysis, student t-test was performed (ns, not significant). (C) Relative gene expression analyzed by qRT-PCR. Data are means ± SEM of two patients with three technical replicates. For statistical analysis, student t-test was performed (ns, not significant; *p<0.05; **p<0.01).
Discussion
This is the first report comparing the phenotypes of dermal fibroblasts taken from different parts of the body of the same WS patient. The WS foot fibroblasts showed a reduced proliferative capacity with shorter telomeres, in comparison to the trunk fibroblasts. Transcriptome analysis showed increased gene expression related to osteogenic differentiation in the foot group and that of adipogenic and chondrogenic differentiation in the trunk group. Indeed, in vitro induction of adipogenesis and chondrogenesis of the foot fibroblasts showed significantly reduced differentiation capacity compared with the trunk. However, there was no difference in osteogenic capacity between the trunk and foot fibroblasts.
Previously, Rinn et al. conducted transcriptome analyses among normal human fibroblasts taken from different sites in the body [17, 18]. Among the DEGs between the extremities and trunk, the expression distribution of HOX genes is consistent with cell migration during human development [38]. Rinn et al. identified that HOXA13 was explicitly up-regulated in foot-derived fibroblasts, while the HOXB gene cluster was trunk-specific [17, 18]; these data are consistent with our results.
However, most of the genes we extracted through the DEG analysis in this study did not show site-specific changes in the previous reports [17, 18]. PPARG, which we found to be up-regulated in the trunk fibroblasts of the WS patients, is a master regulator of adipogenesis, and its overexpression promotes adipose differentiation [39]. In this study, the WS trunk-derived fibroblasts showed higher PPARG expression than the foot-derived ones, and there was a clear difference in the in vitro adipose differentiation ability. These apparent discrepancies are reminiscent of WS phenotypes, namely the trunk with relatively abundant subcutaneous fat and the extremities with lipoatrophy [15, 16]. In addition, although the anatomic origin of the fibroblast is unclear, WRN-depleted fibroblasts exhibit upregulation of PPARG [40]. Taken together with our findings, these results suggest that the regulation of PPARG gene expression in WRN-depleted cells might be context-dependent and that they can be down-regulated in the fibroblasts in the disease site. Further research is needed to understand the mechanism of downregulation of PPARG in WS foot fibroblasts compared to the WS trunk-derived fibroblasts. In addition, STMN2, a marker of osteogenesis, which is up-regulated during the osteogenic induction of mesenchymal stem cells [19], is the gene with the most distinct regulation in this study: the expression was overwhelmingly increased in the foot fibroblasts compared with the trunk.
Honjo et al. reported that painful subcutaneous calcification precedes skin ulcers in WS patients [13]. Patients with WS frequently suffer painful clavus and callus on the feet, which leads to the development of intractable skin ulcers [1, 12]. Ectopic soft tissue calcification also occurs in the limbs of WS patients [41]. Considering our findings that WS foot fibroblasts have a diminished ability to differentiate into adipocytes and chondrocytes while their osteogenic differentiation capacity remains fully preserved, the ossification of fibroblasts in the dermal and subcutaneous layers of the skin may result in these symptoms. In addition, reversible direct conversion of subcutaneous adipocytes into fibroblasts, in vivo, has been reported [42–44]. Therefore, our results suggest the possibility that subcutaneous lipoatrophy in WS extremities might attribute to the inability of adipogenic differentiation in fibroblasts in the disease site.
In this study, we revealed the distinct gene expression profiles and phenotypes in WS dermal fibroblasts derived from the foot and trunk. This study highlights the relationship between fibroblast phenotypes and WS-specific symptoms, refractory skin ulcers and subcutaneous lipoatrophy in extremities. These results could lead to a further understanding of the disease’s mechanism and development of a new therapeutic strategy in the future.
Materials and Methods
Establishment of fibroblasts and cell culture
WS dermal fibroblasts were established from two WS patients (WS1 and WS2, Supplementary Table 4). Both were hospitalized for treatment of their foot skin ulcers, and the skin graft was taken from the trunk (groin). The healthy skin neighboring the ulcer and the skin partly taken from the graft were explanted into a dish as previously described [45]. Cell culture was performed with DMEM (043-30085, Wako, Osaka, Japan), supplemented with 10% FBS (10270106, Gibco, Waltham, MA, U.S.A), and antibiotic-antimycotic (15240062, Gibco) in humidified 5% CO2 air. The medium was changed every two days. When reaching sub-confluency, cells were passaged at a 1:4 split ratio until growth arrest and population doublings were calculated.
Telomere quantitative fluorescence in situ hybridization (Q-FISH)
Fibroblasts at PD10 were treated with the colcemid kit (Chromocenter, Tottori, Japan) and fixed in Carnoy’s solution following the manufacturer’s protocol. The fixed cells on coverslips were treated with ribonuclease (312-01931, Nippon Gene, Tokyo, Japan) and 0.005 % pepsin (V1959, Promega, WI, U.S.), hybridized with peptide nucleic acid oligonucleotide probes (F1002, Panagene, Daejeon, South Korea), and immuno-stained with Hoechst 33342 (H342, Dojindo, Kumamoto, Japan), according to the manufacturer’s protocols. Images were recorded using a BZ-X700 microscope (Keyence, Osaka, Japan). Quantification was performed using the Telometer (https://demarzolab.pathology.jhmi.edu/telometer/), as previously described [46, 47].
Quantitative polymerase chain reaction (qPCR)
RNA was extracted and reverse-transcribed, as previously described [48]. TaqMan Gene Expression Assays for PPARG (Hs01115513_m1), FABP4 (Hs01086177_m1), CEBPA (Hs00269972_s1), LEP (Hs00174877_m1), SOX9 (Hs00165814_m1), ACAN (Hs00153936_m1), ALPL (Hs01029144_m1), SPP1 (Hs00959010_m1), RUNX2 (Hs01047973_m1), COL1A1 (Hs00164004_m1), and B2M (Hs00187842_m1) were purchased from Applied Biosystems (Waltham, MA, U.S.). Quantification was performed with the
Tri-lineage differentiation
In vitro differentiation potentials of the fibroblasts into three mesenchymal lineages were evaluated by using adipogenesis, chondrogenesis, and osteogenesis differentiation kits (A1007001, A1007101, and A1007201, respectively; Gibco) according to manufacturer's protocols. Cells at PD 9 to 10 were used. After two weeks of differentiation, cells were sampled and stained. For each staining assay, Oil red O, Alcian blue, and Alizarin red staining (Sigma-Aldrich, St. Louis, MO, U.S.A) were used, respectively. Quantification of the stains was performed using a BZ-X700 microscope (Keyence, Osaka, Japan).
Transcriptome analysis
mRNA was extracted from fibroblasts at PD 10 and the cDNA library was synthesized using the NEBNext Ultra RNA Library Prep Kit (E7370S, New England BioLabs, Beverly, MA, U.S.A). Sequencing was carried out (technically n=2 in each sample) by HiSeq1500 (Illumina, San Diego, CA, U.S.A) with 60bp single-reads. The reference genome mapping (UCSC/hg19) was performed using TopHat (version 2.0.13; with default parameters) with annotation data from iGenomes (Illumina). Cuffdiff (Cufflinks version 2.2.1; with default parameters) was used to quantify the gene expression levels. FPKM data were analyzed by iDEP (http://bioinformatics.sdstate.edu/idep/) as described by the authors [49].
Study approval
All experiments were approved by the institutional review boards at the Chiba University Graduate School of Medicine (Chiba, Japan). Written informed consent was obtained from study participants before the commencement of this research.
Supplementary Materials
Author Contributions
H.Kato, Y.M., and K.Y. designed the study, analyzed the data, and wrote the manuscript; H.Kato carried out the experiments and made the figures; N.T., M.O., and A.I. conducted transcriptome analysis; Y.O., M.K., and K.E. discussed the data; H.Kaneko, D.K., and A.T.W. carried out the experiments; H.O., Y.K., and N.M. conducted sampling from patients; all authors approved the final version of the manuscript.
Acknowledgments
We want to thank Ms. Naoko Tomoda (Department of Endocrinology, Hematology and Gerontology, Chiba University Graduate School of Medicine, Chiba, Japan) for supporting the funding management.
Conflicts of Interest
The authors declare that they have no conflicts of interest.
Funding
This work was supported by Japan Society for the Promotion of Science (JSPS) KAKENHI under Grant Numbers JP19K23939 (H.Kato), JP20H00524 (K.Y.), JP20K16542 (H.Kato); Japan Agency for Medical Research and Development (AMED) under Grant Numbers JP20bm0804016 (K.Y.), JP20ek0109353 (K.Y.), JP20gm5010002 (K.Y.); Ministry of Health, Labour and Welfare (MHLW) of Japan under Grant Number H30-nanchi-ippan-009 (K.Y.).
References
- 1. Oshima J, Sidorova JM, Monnat RJ
Jr . Werner syndrome: clinical features, pathogenesis and potential therapeutic interventions. Ageing Res Rev. 2017; 33:105–14. https://doi.org/10.1016/j.arr.2016.03.002 [PubMed] - 2. Muftuoglu M, Oshima J, von Kobbe C, Cheng WH, Leistritz DF, Bohr VA. The clinical characteristics of Werner syndrome: molecular and biochemical diagnosis. Hum Genet. 2008; 124:369–77. https://doi.org/10.1007/s00439-008-0562-0 [PubMed]
- 3. Hisama FM, Oshima J, Martin GM. How research on human progeroid and antigeroid syndromes can contribute to the longevity dividend initiative. Cold Spring Harb Perspect Med. 2016; 6:a025882. https://doi.org/10.1101/cshperspect.a025882 [PubMed]
- 4. Monnat RJ
Jr . ”...Rewritten in the skin“: clues to skin biology and aging from inherited disease. J Invest Dermatol. 2015; 135:1484–90. https://doi.org/10.1038/jid.2015.88 [PubMed] - 5. Huang S, Lee L, Hanson NB, Lenaerts C, Hoehn H, Poot M, Rubin CD, Chen DF, Yang CC, Juch H, Dorn T, Spiegel R, Oral EA, et al. The spectrum of WRN mutations in Werner syndrome patients. Hum Mutat. 2006; 27:558–67. https://doi.org/10.1002/humu.20337 [PubMed]
- 6. Yokote K, Chanprasert S, Lee L, Eirich K, Takemoto M, Watanabe A, Koizumi N, Lessel D, Mori T, Hisama FM, Ladd PD, Angle B, Baris H, et al. WRN mutation update: mutation spectrum, patient registries, and translational prospects. Hum Mutat. 2017; 38:7–15. https://doi.org/10.1002/humu.23128 [PubMed]
- 7. Satoh M, Imai M, Sugimoto M, Goto M, Furuichi Y. Prevalence of Werner’s syndrome heterozygotes in Japan. Lancet. 1999; 353:1766. https://doi.org/10.1016/S0140-6736(98)05869-3 [PubMed]
- 8. Rossi ML, Ghosh AK, Bohr VA. Roles of Werner syndrome protein in protection of genome integrity. DNA Repair (Amst). 2010; 9:331–44. https://doi.org/10.1016/j.dnarep.2009.12.011 [PubMed]
- 9. Faragher RG, Kill IR, Hunter JA, Pope FM, Tannock C, Shall S. The gene responsible for Werner syndrome may be a cell division ”counting“ gene. Proc Natl Acad Sci USA. 1993; 90:12030–34. https://doi.org/10.1073/pnas.90.24.12030 [PubMed]
- 10. Wyllie FS, Jones CJ, Skinner JW, Haughton MF, Wallis C, Wynford-Thomas D, Faragher RG, Kipling D. Telomerase prevents the accelerated cell ageing of Werner syndrome fibroblasts. Nat Genet. 2000; 24:16–17. https://doi.org/10.1038/71630 [PubMed]
- 11. Shimamoto A, Kagawa H, Zensho K, Sera Y, Kazuki Y, Osaki M, Oshimura M, Ishigaki Y, Hamasaki K, Kodama Y, Yuasa S, Fukuda K, Hirashima K, et al. Reprogramming suppresses premature senescence phenotypes of Werner syndrome cells and maintains chromosomal stability over long-term culture. PLoS One. 2014; 9:e112900. https://doi.org/10.1371/journal.pone.0112900 [PubMed]
- 12. Takemoto M, Mori S, Kuzuya M, Yoshimoto S, Shimamoto A, Igarashi M, Tanaka Y, Miki T, Yokote K. Diagnostic criteria for Werner syndrome based on Japanese nationwide epidemiological survey. Geriatr Gerontol Int. 2013; 13:475–81. https://doi.org/10.1111/j.1447-0594.2012.00913.x [PubMed]
- 13. Honjo S, Yokote K, Fujimoto M, Takemoto M, Kobayashi K, Maezawa Y, Shimoyama T, Satoh S, Koshizaka M, Takada A, Irisuna H, Saito Y. Clinical outcome and mechanism of soft tissue calcification in Werner syndrome. Rejuvenation Res. 2008; 11:809–19. https://doi.org/10.1089/rej.2007.0649 [PubMed]
- 14. Yamaga M, Takemoto M, Shoji M, Sakamoto K, Yamamoto M, Ishikawa T, Koshizaka M, Maezawa Y, Kobayashi K, Yokote K. Werner syndrome: a model for sarcopenia due to accelerated aging. Aging (Albany NY). 2017; 9:1738–44. https://doi.org/10.18632/aging.101265 [PubMed]
- 15. Mori S, Murano S, Yokote K, Takemoto M, Asaumi S, Take A, Saito Y. Enhanced intra-abdominal visceral fat accumulation in patients with werner’s syndrome. Int J Obes Relat Metab Disord. 2001; 25:292–95. https://doi.org/10.1038/sj.ijo.0801529 [PubMed]
- 16. Donadille B, D’Anella P, Auclair M, Uhrhammer N, Sorel M, Grigorescu R, Ouzounian S, Cambonie G, Boulot P, Laforêt P, Carbonne B, Christin-Maitre S, Bignon YJ, Vigouroux C. Partial lipodystrophy with severe insulin resistance and adult progeria Werner syndrome. Orphanet J Rare Dis. 2013; 8:106. https://doi.org/10.1186/1750-1172-8-106 [PubMed]
- 17. Rinn JL, Bondre C, Gladstone HB, Brown PO, Chang HY. Anatomic demarcation by positional variation in fibroblast gene expression programs. PLoS Genet. 2006; 2:e119. https://doi.org/10.1371/journal.pgen.0020119 [PubMed]
- 18. Rinn JL, Wang JK, Liu H, Montgomery K, van de Rijn M, Chang HY. A systems biology approach to anatomic diversity of skin. J Invest Dermatol. 2008; 128:776–82. https://doi.org/10.1038/sj.jid.5700986 [PubMed]
- 19. Chiellini C, Grenningloh G, Cochet O, Scheideler M, Trajanoski Z, Ailhaud G, Dani C, Amri EZ. Stathmin-like 2, a developmentally-associated neuronal marker, is expressed and modulated during osteogenesis of human mesenchymal stem cells. Biochem Biophys Res Commun. 2008; 374:64–68. https://doi.org/10.1016/j.bbrc.2008.06.121 [PubMed]
- 20. Choung HW, Lee DS, Lee JH, Shon WJ, Lee JH, Ku Y, Park JC. Tertiary dentin formation after indirect pulp capping using protein CPNE7. J Dent Res. 2016; 95:906–12. https://doi.org/10.1177/0022034516639919 [PubMed]
- 21. Kim JS, Kim WK, Oh KJ, Lee EW, Han BS, Lee SC, Bae KH. Protein tyrosine phosphatase, receptor type B (PTPRB) inhibits brown adipocyte differentiation through regulation of VEGFR2 phosphorylation. J Microbiol Biotechnol. 2019; 29:645–50. https://doi.org/10.4014/jmb.1810.10033 [PubMed]
- 22. Ambrosi TH, Scialdone A, Graja A, Gohlke S, Jank AM, Bocian C, Woelk L, Fan H, Logan DW, Schürmann A, Saraiva LR, Schulz TJ. Adipocyte accumulation in the bone marrow during obesity and aging impairs stem cell-based hematopoietic and bone regeneration. Cell Stem Cell. 2017; 20:771–84.e6. https://doi.org/10.1016/j.stem.2017.02.009 [PubMed]
- 23. Mao G, Kang Y, Lin R, Hu S, Zhang Z, Li H, Liao W, Zhang Z. Long non-coding RNA HOTTIP promotes CCL3 expression and induces cartilage degradation by sponging miR-455-3p. Front Cell Dev Biol. 2019; 7:161. https://doi.org/10.3389/fcell.2019.00161 [PubMed]
- 24. Lee EJ, Jan AT, Baig MH, Ahmad K, Malik A, Rabbani G, Kim T, Lee IK, Lee YH, Park SY, Choi I. Fibromodulin and regulation of the intricate balance between myoblast differentiation to myocytes or adipocyte-like cells. FASEB J. 2018; 32:768–81. https://doi.org/10.1096/fj.201700665R [PubMed]
- 25. Akune T, Ohba S, Kamekura S, Yamaguchi M, Chung UI, Kubota N, Terauchi Y, Harada Y, Azuma Y, Nakamura K, Kadowaki T, Kawaguchi H. PPARgamma insufficiency enhances osteogenesis through osteoblast formation from bone marrow progenitors. J Clin Invest. 2004; 113:846–55. https://doi.org/10.1172/JCI19900 [PubMed]
- 26. Wang Y, Liu Y, Fan Z, Liu D, Wang F, Zhou Y. IGFBP2 enhances adipogenic differentiation potentials of mesenchymal stem cells from wharton’s jelly of the umbilical cord via JNK and Akt signaling pathways. PLoS One. 2017; 12:e0184182. https://doi.org/10.1371/journal.pone.0184182 [PubMed]
- 27. Deng P, Chen Y, Ji N, Lin Y, Yuan Q, Ye L, Chen Q. Cysteine dioxygenase type 1 promotes adipogenesis via interaction with peroxisome proliferator-activated receptor gamma. Biochem Biophys Res Commun. 2015; 458:123–27. https://doi.org/10.1016/j.bbrc.2015.01.080 [PubMed]
- 28. Zhao X, Deng P, Feng J, Wang Z, Xiang Z, Han X, Bai D, Pae EK. Cysteine dioxygenase type 1 inhibits osteogenesis by regulating Wnt signaling in primary mouse bone marrow stromal cells. Sci Rep. 2016; 6:19296. https://doi.org/10.1038/srep19296 [PubMed]
- 29. Huang J, Peng J, Cao G, Lu S, Liu L, Li Z, Zhou M, Feng M, Shen H. Hypoxia-induced MicroRNA-429 promotes differentiation of MC3T3-E1 osteoblastic cells by mediating ZFPM2 expression. Cell Physiol Biochem. 2016; 39:1177–86. https://doi.org/10.1159/000447824 [PubMed]
- 30. López-Otín C, Blasco MA, Partridge L, Serrano M, Kroemer G. The hallmarks of aging. Cell. 2013; 153:1194–217. https://doi.org/10.1016/j.cell.2013.05.039 [PubMed]
- 31. Hernandez-Segura A, Nehme J, Demaria M. Hallmarks of cellular senescence. Trends Cell Biol. 2018; 28:436–53. https://doi.org/10.1016/j.tcb.2018.02.001 [PubMed]
- 32. Blasi A, Martino C, Balducci L, Saldarelli M, Soleti A, Navone SE, Canzi L, Cristini S, Invernici G, Parati EA, Alessandri G. Dermal fibroblasts display similar phenotypic and differentiation capacity to fat-derived mesenchymal stem cells, but differ in anti-inflammatory and angiogenic potential. Vasc Cell. 2011; 3:5. https://doi.org/10.1186/2045-824X-3-5 [PubMed]
- 33. Junker JP, Sommar P, Skog M, Johnson H, Kratz G. Adipogenic, chondrogenic and osteogenic differentiation of clonally derived human dermal fibroblasts. Cells Tissues Organs. 2010; 191:105–18. https://doi.org/10.1159/000232157 [PubMed]
- 34. Lorenz K, Sicker M, Schmelzer E, Rupf T, Salvetter J, Schulz-Siegmund M, Bader A. Multilineage differentiation potential of human dermal skin-derived fibroblasts. Exp Dermatol. 2008; 17:925–32. https://doi.org/10.1111/j.1600-0625.2008.00724.x [PubMed]
- 35. Rosen ED, Walkey CJ, Puigserver P, Spiegelman BM. Transcriptional regulation of adipogenesis. Genes Dev. 2000; 14:1293–307. [PubMed]
- 36. Goldring MB, Tsuchimochi K, Ijiri K. The control of chondrogenesis. J Cell Biochem. 2006; 97:33–44. https://doi.org/10.1002/jcb.20652 [PubMed]
- 37. Komori T. Regulation of bone development and extracellular matrix protein genes by RUNX2. Cell Tissue Res. 2010; 339:189–95. https://doi.org/10.1007/s00441-009-0832-8 [PubMed]
- 38. Krumlauf R. Hox genes in vertebrate development. Cell. 1994; 78:191–201. https://doi.org/10.1016/0092-8674(94)90290-9 [PubMed]
- 39. Lehrke M, Lazar MA. The many faces of PPARgamma. Cell. 2005; 123:993–99. https://doi.org/10.1016/j.cell.2005.11.026 [PubMed]
- 40. Tang W, Robles AI, Beyer RP, Gray LT, Nguyen GH, Oshima J, Maizels N, Harris CC, Monnat RJ
Jr . The Werner syndrome RECQ helicase targets G4 DNA in human cells to modulate transcription. Hum Mol Genet. 2016; 25:2060–69. https://doi.org/10.1093/hmg/ddw079 [PubMed] - 41. Leone A, Costantini AM, Brigida R, Antoniol OM, Antonelli-Incalzi R, Bonomo L. Soft-tissue mineralization in Werner syndrome. Skeletal Radiol. 2005; 34:47–51. https://doi.org/10.1007/s00256-004-0792-8 [PubMed]
- 42. Chen SX, Zhang LJ, Gallo RL. Dermal white adipose tissue: a newly recognized layer of skin innate defense. J Invest Dermatol. 2019; 139:1002–09. https://doi.org/10.1016/j.jid.2018.12.031 [PubMed]
- 43. Plikus MV, Guerrero-Juarez CF, Ito M, Li YR, Dedhia PH, Zheng Y, Shao M, Gay DL, Ramos R, Hsi TC, Oh JW, Wang X, Ramirez A, et al. Regeneration of fat cells from myofibroblasts during wound healing. Science. 2017; 355:748–52. https://doi.org/10.1126/science.aai8792 [PubMed]
- 44. Zhang Z, Shao M, Hepler C, Zi Z, Zhao S, An YA, Zhu Y, Ghaben AL, Wang MY, Li N, Onodera T, Joffin N, Crewe C, et al. Dermal adipose tissue has high plasticity and undergoes reversible dedifferentiation in mice. J Clin Invest. 2019; 129:5327–42. https://doi.org/10.1172/JCI130239 [PubMed]
- 45. Toda T, Satoh M, Sugimoto M, Goto M, Furuichi Y, Kimura N. A comparative analysis of the proteins between the fibroblasts from Werner’s syndrome patients and age-matched normal individuals using two-dimensional gel electrophoresis. Mech Ageing Dev. 1998; 100:133–43. https://doi.org/10.1016/s0047-6374(97)00131-0 [PubMed]
- 46. Meeker AK, Gage WR, Hicks JL, Simon I, Coffman JR, Platz EA, March GE, De Marzo AM. Telomere length assessment in human archival tissues: combined telomere fluorescence in situ hybridization and immunostaining. Am J Pathol. 2002; 160:1259–68. https://doi.org/10.1016/S0002-9440(10)62553-9 [PubMed]
- 47. Mourkioti F, Kustan J, Kraft P, Day JW, Zhao MM, Kost-Alimova M, Protopopov A, DePinho RA, Bernstein D, Meeker AK, Blau HM. Role of telomere dysfunction in cardiac failure in duchenne muscular dystrophy. Nat Cell Biol. 2013; 15:895–904. https://doi.org/10.1038/ncb2790 [PubMed]
- 48. Ide S, Finer G, Maezawa Y, Onay T, Souma T, Scott R, Ide K, Akimoto Y, Li C, Ye M, Zhao X, Baba Y, Minamizuka T, et al. Transcription factor 21 is required for branching morphogenesis and regulates the gdnf-axis in kidney development. J Am Soc Nephrol. 2018; 29:2795–808. https://doi.org/10.1681/ASN.2017121278 [PubMed]
- 49. Ge SX, Son EW, Yao R. iDEP: an integrated web application for differential expression and pathway analysis of RNA-seq data. BMC Bioinformatics. 2018; 19:534. https://doi.org/10.1186/s12859-018-2486-6 [PubMed]


