Introduction
The gut microbiota has emerged as one of the most important factors that contribute to host health and aging [1]. The interplay between gut microbes and epithelial cells regulates various aspects of gut physiology, including proper epithelial and immune development, immune function and metabolism [2]. High-throughput screening of microbial pro-longevity factors from commensal Escherichia coli showed that 29 bacterial genes when deleted, increase the longevity of the host by interacting with several aging-regulatory pathways [3]. Also, the reported age-related changes in the gut microbiota composition, including a decrease in the Firmicutes phylum with perturbations in the Lactobacillus species diversity, suggest a bacterial impact on host aging [2, 4, 5]. Interestingly, diets based on probiotic lactobacilli supplementation have been shown to improve the state of dysbiosis observed in elderly people [6]. Reports show that beneficial lactobacilli exhibit the potential to stimulate aging-related genes capable of enhancing host longevity and the anti-stress responses [7, 8]. It is important to note that the activation of a specific longevity program is strain-dependent and strongly related to the applied Lactobacillus strain, which might interfere with different cellular targets [9].
The roundworm Caenorhabditis elegans, with its short and easily monitored lifespan, has been successfully used as a model system in the field of aging research. The worms are reared on a single bacterial strain of E. coli and this well-defined microbiota enables using C. elegans for studying the relationship between gut microbes and host aging [10]. Several self-protective stress-response regulators, including the forkhead transcription factor FOXO/DAF-16 [11], canonical p38 mitogen-activated protein kinase (MAPK)/PMK-1 [12] and Nrf2/SKN-1 [13], have been found to participate in the longevity assurance in C. elegans triggered by different probiotic bacteria.
Interestingly, it has been shown that the surface biomolecules [14] and metabolites [15] of lactobacilli could trigger autophagy, a commonly known pro-survival mechanism responsible for the recycling of long-lived proteins, molecules and organelles [16]. Currently, little is known on whether the activated autophagy could be the mechanism responsible for the pro-longevity effects of lactobacilli. In C. elegans, one of the major regulators of many autophagy-related genes is the basic helix-loop-helix 30 (HLH-30), a functional mammalian orthologue of the transcription factor EB (TFEB), exhibiting a key role in lifespan determination [17]. Besides autophagy, HLH-30 links lysosomal lipolysis and autophagy to control fat storage [18] and dictates host defense against infection [19]. By controlling the majority of host cytoprotective responses, the activation of HLH-30/TFEB distinguished itself as an attractive target for beneficial microbes to strengthen the host resistance.
Next to the traditional use of probiotics, the application of non-viable microbial cells, microbial fractions or cell lysates (postbiotics), which can mimic the physiological benefits of probiotics in a more controllable way than using live bacteria, started gaining attention for various purposes [20]. Therefore, we report here that the heat-inactivated human commensal Lactobacillus fermentum BGHV110 strain extends the lifespan of C. elegans in an HLH-30-dependent manner. We showed that increased transcription of the hlh-30 gene primarily results in the upregulation of autophagy, contributing to the pro-longevity effect of the BGHV110 postbiotic.
Results
Heat-inactivated Lb. fermentum BGHV110 extends lifespan in C. elegans
To investigate the effect of a postbiotic-based diet on organismal aging, we used C. elegans as a model system. Our previous study identified the prospective postbiotic potential of the strain Lb. fermentum BGHV110 in the modulation of autophagy in human hepatoma HepG2 cells [14]. Given the role of autophagy as a longevity-related mechanism [21, 22], we assumed that heat-inactivated BGHV110 could exhibit beneficial effects on the host lifespan. Initially, the effect of the heat-inactivated BGHV110 strain was assessed in a C. elegans N2 wild-type (WT) strain. In order to focus our research exclusively on the effect on aging, and not host development, feeding with BGHV110 was started from the first day of adulthood. Hence, age-synchronous worms were grown to the L4 stage on an E. coli OP50 strain and then transferred to plates containing heat-inactivated OP50 (control) or BGHV110 bacteria. Lifespan analysis revealed that feeding with BGHV110 significantly extended both median and maximal lifespan in worms compared to the OP50 fed control (Figure 1A).
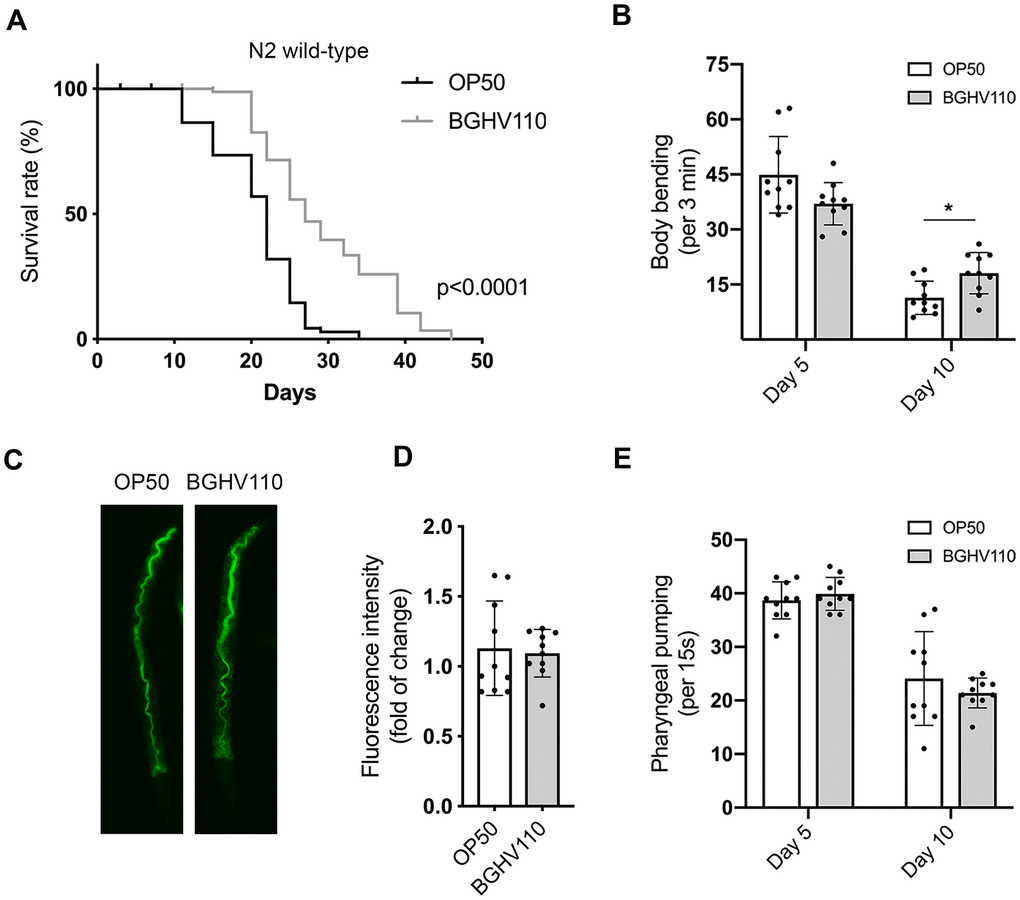
Figure 1. Heat-inactivated Lb. fermentum BGHV110 affects longevity and healthspan in C. elegans. (A) Lifespan curve of WT animals fed with heat-inactivated control (OP50) and BGHV110 bacteria from the L4 developmental stage maintained at 20° C (n=100 per group, results from one of two experiments with similar results are shown). (B) Body bending rates were measured in WT animals on day 5 and day 10 of adulthood (n=10 per group, results are representative of 3 independent assays). (C) Gut localization and (D) fluorescence intensity quantification of acridine orange-stained heat-inactivated OP50 and BGHV110 bacteria visualized by fluorescence microscopy in day 1 adult WT animals (n=10 per group, results are representative of 3 independent assays). (E) Pharyngeal pumping rates were measured in WT animals on day 5 and day 10 of adulthood (n=10 per group, results are representative of 3 independent assays). All values are presented as mean ± SD. Student’s t-test was used to compare the treated group relative to control (*p < 0.05). The log-rank (Mantel-Cox) test was used to assess the p-value in lifespan analysis.
Since aging also affects different fitness parameters, we next assessed locomotor activity, shown in worms to decline with age. Indeed, in the control animals, we detected a steep decrease in the frequency of body bending with increasing age (Figure 1B). The decline was less prominent in the BGHV110 fed animals that exhibited a higher motility rate on day 10 of adulthood in comparison with the OP50 fed control (Figure 1B). To test if the longevity-promoting signal might be a consequence of the dietary restriction (DR) caused by decreased feeding on the Lactobacillus strain, we prestained the BGHV110 with acridine orange and visualized its presence in the worm intestinal lumen. However, no difference in the fluorescence intensity of the bacteria present in the gut lumen was detected between the two groups, suggesting that the increase in lifespan was not caused by the putative DR (Figure 1C, 1D).
We further measured the pharyngeal pumping rate that controls the food intake ability in C. elegans and also decreases with age. Although an age-associated decline in the pharyngeal muscle function was detected, no differences in bacterial uptake between the OP50 and the BGHV110 fed animals were observed, either on day 5 or day 10 of adulthood (Figure 1E).
Overall, these results indicate that the heat-inactivated BGHV110 postbiotics are able to delay aging and improve the motility of treated animals independently of the number of ingested bacteria.
Heat-inactivated Lb. fermentum BGHV110 upregulates autophagy
We next focused on the possible involvement of autophagy in the pro-longevity effect of BGHV110 feeding. We used a reporter DA2123 transgenic strain expressing a GFP-tagged version of LGG-1 under the lgg-1 promoter, an orthologue of the mammalian LC3 autophagy marker. Our results revealed a significant increase in GFP fluorescence intensity in the worms treated with the BGHV110 strain compared to control (Figure 2A, 2B). To confirm this result, we followed the expression levels of cleaved GFP, which appears upon the formation of the autolysosome, and the LGG-1-II::GFP expression as indicators of autophagy activation by using the Western blot analysis [23]. The intensity of the band corresponding to free GFP was significantly increased in the worms fed with BGHV110, as well as the LGG-1-II/HSC-70 ratio, compared to OP50 control, suggesting an increase in autophagy (Figure 2C, 2D). To gain further insights into the observed activation of autophagy, we analyzed the transcript levels of relevant genes. Using this analysis, we were able to show that the transcript levels of the unc-51, bec-1, lgg-1, atg-7 and atg-18, encoding proteins involved in the different steps of the autophagy process, were all significantly upregulated upon feeding the animals with BGHV110. Interestingly, an increased mRNA level was also observed for the hlh-30 gene (Figure 2E). However, in order to address the possibility that the activation of autophagy came as a DR-induced phenomenon, we performed an additional experiment where we evaluated the transcript levels of autophagy-related genes after feeding the animals with heat-inactivated OP50 containing heat-inactivated fractions of BGHV110 in a 1:2 ratio. We observed increased transcription of all tested genes in the worms fed with the heat-inactivated OP50 supplemented with BGHV110, which strengthened our conclusion that BGHV110 triggered autophagy independently of the changes in nutrient intake (Figure 2F). Collectively, these results imply that the effect of BGHV110 on the worms’ lifespan could be mediated through the activation of autophagy controlled by HLH-30.
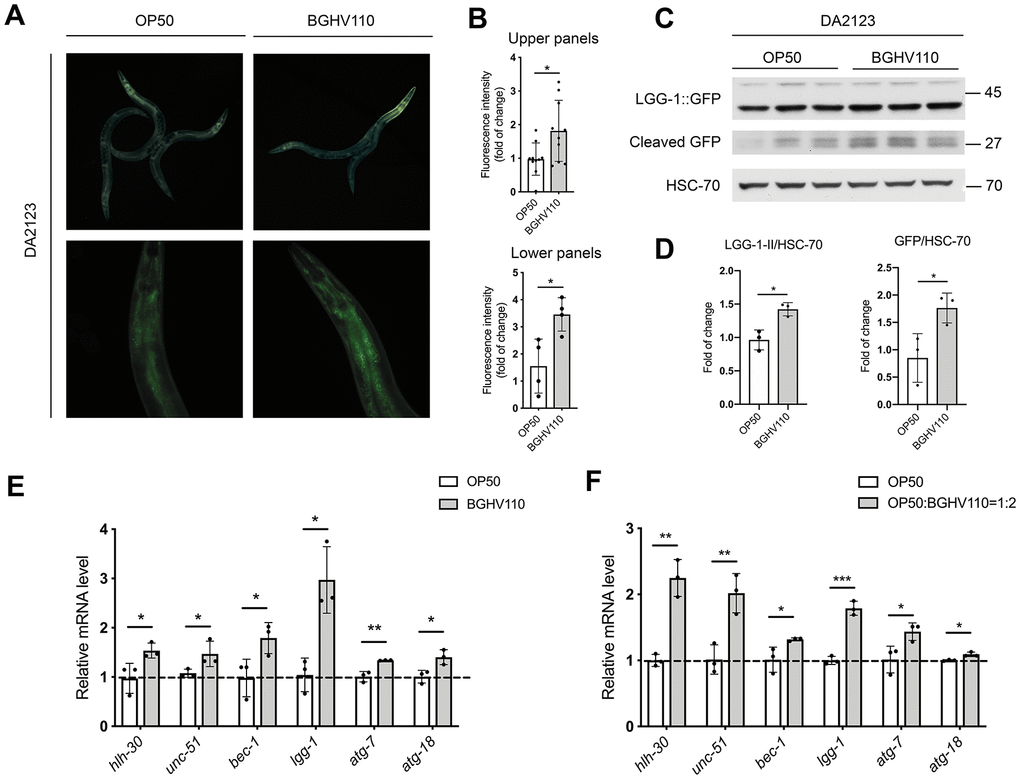
Figure 2. Heat-inactivated Lb. fermentum BGHV110 triggers autophagy in C. elegans. Representative fluorescence images (A) and quantifications (B) of day 1 adult DA2123 transgenic animals expressing GFP::LGG-1 under the lgg-1 promoter after overnight BGHV110 treatment (n=4–10, results are representative of 3 independent assays). Western blots (C) and densitometric analysis (D) showing the levels of GFP::LGG-1 and cleaved GFP proteins isolated from the DA2123 transgenic strain on day 1 of adulthood after overnight BGHV110 treatment. HSC-70 was used as a loading control (n=3, three independent experiments). Expression of hlh-30 and autophagy-related genes was measured by qRT-PCR in L4 stage WT animals after 6 h of treatment with (E) heat-inactivated BGHV110 and (F) heat-inactivated OP50 supplemented with heat-inactivated BGHV110 in a 1:2 ratio (n=3, three independent experiments). All values are presented as mean ± SD. Student’s t-test was used to compare the treated group relative to control (*p < 0.05, **p < 0.01, ***p < 0.001).
HLH-30 is necessary for the activation of the longevity program in C. elegans triggered by heat-inactivated BGHV110
To further elucidate the role of autophagy in the longevity assurance program activated by the BGHV110 postbiotic, we used the hlh-30 (tm1978) C. elegans mutant. The lifespan extension caused by BGHV110-feeding in WT worms was completely abrogated in the hlh-30 mutant (Figure 3A). As expected, feeding of the hlh-30 mutant with the BGHV110 strain also failed to improve the aging-related decrease of body bending rate at day 10 of adulthood (Figure 3B). To identify what happens with the autophagy markers, which were upregulated in WT animals upon BGHV110 treatment, we performed a qRT-PCR analysis in the hlh-30 mutant. Notably, most of the genes involved in the autophagy process showed no changes in transcription after the BGHV110 treatment of the hlh-30 mutant (Figure 3C). Moreover, the transcript level of the atg-18, which is required for autophagosome formation and responsible for the longevity phenotype of C. elegans [24, 25], was significantly decreased in the hlh-30 mutant after the BGHV110 treatment (Figure 3C). Surprisingly, feeding with BGHV110 still resulted in an elevated lgg-1 transcript level in the hlh-30 animals, demonstrating that a higher lgg-1 level alone is not sufficient to prolong lifespan. Collectively, these data strongly suggest that BGHV110 activates HLH-30-dependent mechanisms to extend the lifespan of C. elegans. These protective responses seem to highly depend on autophagy, as it was not activated in the hlh-30 mutant.
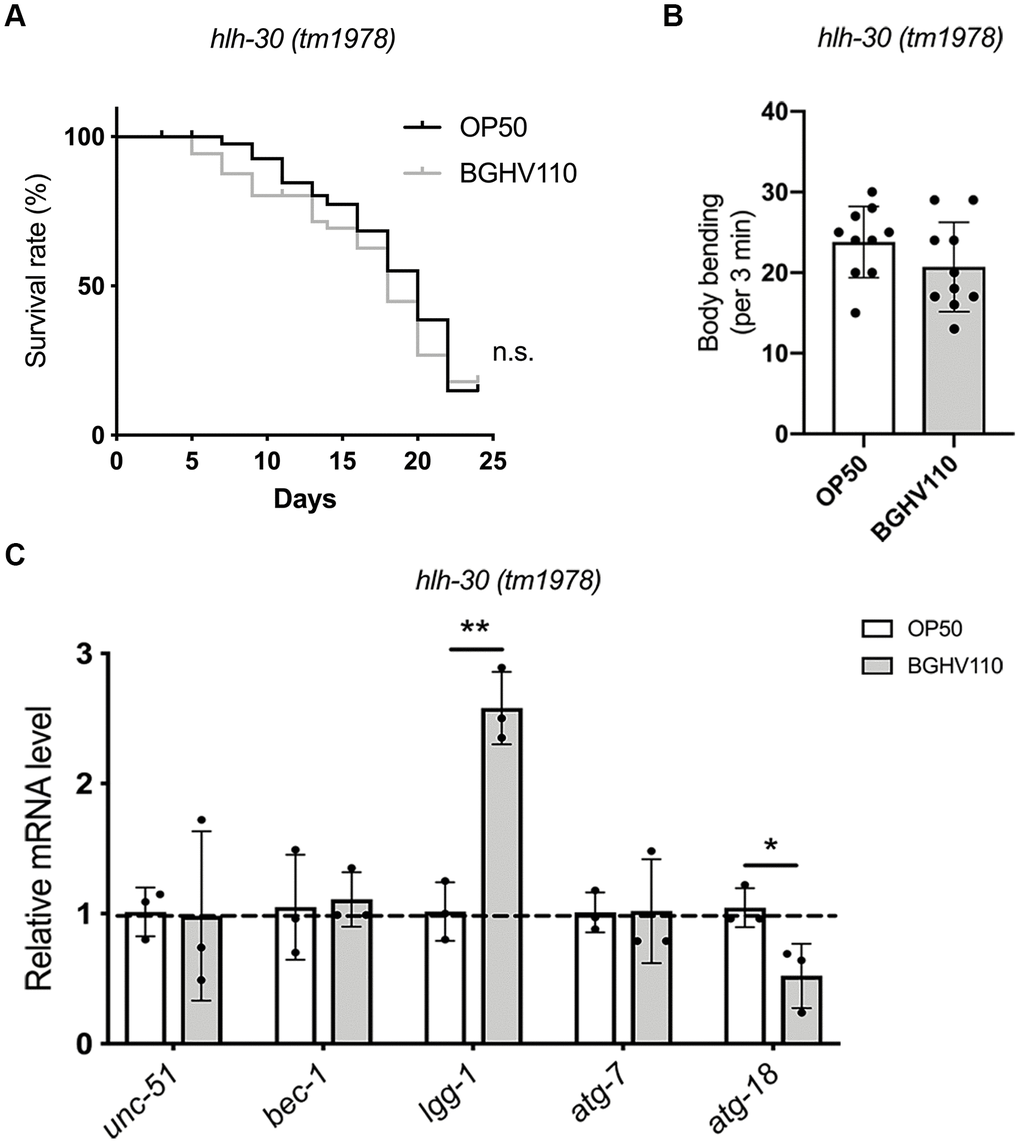
Figure 3. Heat-inactivated Lb. fermentum BGHV110 delays aging in C. elegans in an HLH-30-dependent manner. (A) Lifespan curve of the hlh-30 (tm1978) mutant fed with heat-inactivated control (OP50) and BGHV110 bacteria from the L4 developmental stage maintained at 20° C (n=100 per group, results from one of two experiments with similar results are shown). (B) Body bending rate was measured in the hlh-30 (tm1978) mutant on day 10 of adulthood (n=10 per group, results are representative of 3 independent assays). (C) Expression of autophagy-related genes was measured by qRT-PCR in the L4 stage of the hlh-30 (tm1978) mutant after 6 h of treatment with heat-inactivated BGHV110 (n=3, three independent experiments). All values are presented as mean ± SD. Student’s t-test was used to compare the treated group relative to control (*p < 0.05, **p < 0.01). The log-rank (Mantel-Cox) test was used to assess the p-value in lifespan analysis.
Heat-inactivated Lb. fermentum BGHV110 reduces lipid droplets
Having in mind that one of the hallmarks of aging is the accumulation of lipids and that HLH-30 could control fat storage in C. elegans through a process called lipophagy [18], we analyzed lipid accumulation in WT and the hlh-30 mutant on day 5 of adulthood. We found that animals fed with BGHV110 showed 2.2-fold lower lipid levels in comparison with the OP50 fed control (Figure 4A, 4B). Surprisingly, we also observed that BGHV110 caused a 2-fold decrease in lipid levels in the hlh-30 animals compared to OP50 control (Figure 4A, 4B). We further looked into the expression levels of lipl genes that encode lipases, the enzymes proposed to break down lipids in C. elegans [18]. Upon feeding with BGHV110, elevated transcript levels of the lipl-1 and lipl-3 genes were observed in WT animals, exclusively. In contrast, elevated transcripts of the lipl-4 and lipl-5 genes were detected only in the hlh-30 mutant fed with BGHV110. Finally, the BGHV110 diet caused upregulation of the lipl-2 gene in both C. elegans strains (Figure 4C). These results suggest that HLH-30 controls the expression of lipl-1 and lipl-3, which is in accordance with the literature data showing that LIPL-1 and LIPL-3 are the key enzymes breaking down lipids through lipophagy. The LIPL-1 and LIPL-3 are activated by HLH-30 in C. elegans [18] and together with LIPL-2 could be responsible for the lipolytic effect induced by BGHV110 mainly through the lipophagy process. Alternatively, activation of different lipases, such as LIPL-4 and LIPL-5, could be related to the reduced lipid content observed in the hlh-30 mutant treated with BGHV110 in a lipophagy-independent fashion. However, despite the differences in the expression profile of lipases-encoded genes, the reduction of lipid levels in both strains implies that BGHV110 could drive lipid degradation both in an HLH-30-independent and HLH-30-dependent manner. These data indicate that, while the BGHV110 diet controls lipid content in worms, this effect seems to be a consequence of the different lipl gene expression profile and not related to the observed increase in longevity.
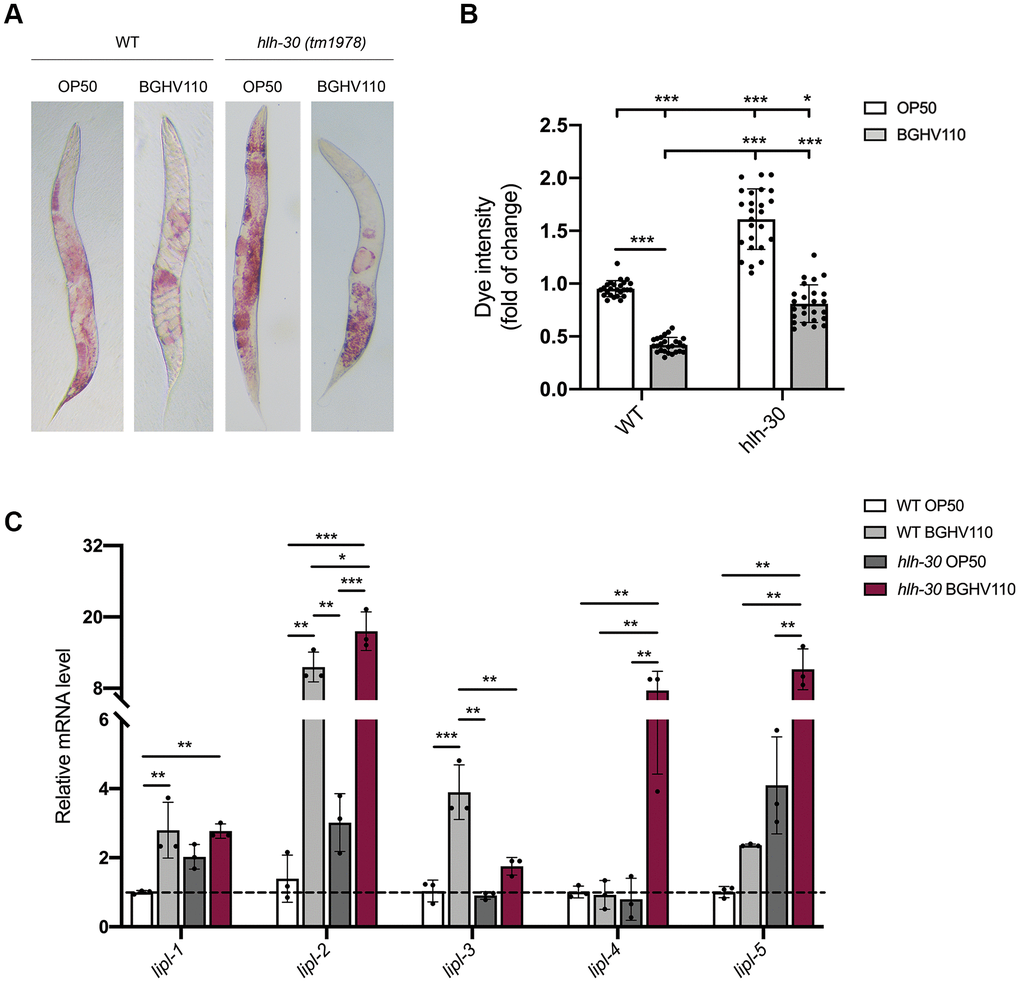
Figure 4. Heat-inactivated Lb. fermentum BGHV110 affects lipid metabolism. (A) Oil Red O staining of the WT and hlh-30 mutant animals fed with heat-inactivated control (OP50) and BGHV110 bacteria analyzed on day 5 of adulthood. (B) Dye intensity quantification of the WT and hlh-30 mutant animals compared to OP50 control (n=25 per group, results are representative of 3 independent assays). (C) The expression of lipl-1, 2, 3, 4, 5 genes was measured by qRT-PCR in the L4 stage WT and hlh-30 mutant after 6 h of treatment with heat-inactivated BGHV110 (n=3, three independent experiments). All values are presented as mean ± SD. One-way ANOVA followed by Tukey post hoc test for multiple comparisons was used (*p < 0.05, **p < 0.01, ***p < 0.001).
Heat-inactivated Lb. fermentum BGHV110 does not act through the SKN-1 pathway
One of the transcription factors that is commonly involved in the regulation of longevity through multiple pathways is SKN-1, mainly acting as a regulator of oxidative stress and xenobiotic responses [13]. We hypothesize that SKN-1 could also be involved in the regulation of the longevity phenotype triggered by inactivated lactobacilli intake. We used a reporter strain harboring GFP expressed under the gst-4 (glutathione-S-transferases-4) promoter, which is primary regulated by SKN-1. Remarkably, we did not observe any differences in signal intensity between the OP50 and BGHV110 fed animals (Figure 5A). Moreover, the skn-1 and gst-4 transcript levels were unchanged upon feeding with BGHV110 (Figure 5B). In agreement with these results, feeding with the BGHV110 strain also failed to ameliorate oxidative stress damage in the functional H2O2 assay, suggesting that BGHV110 does not influence the general antioxidant activity, nor does it change redox homeostasis to promote lifespan extension in C. elegans (Figure 5C).
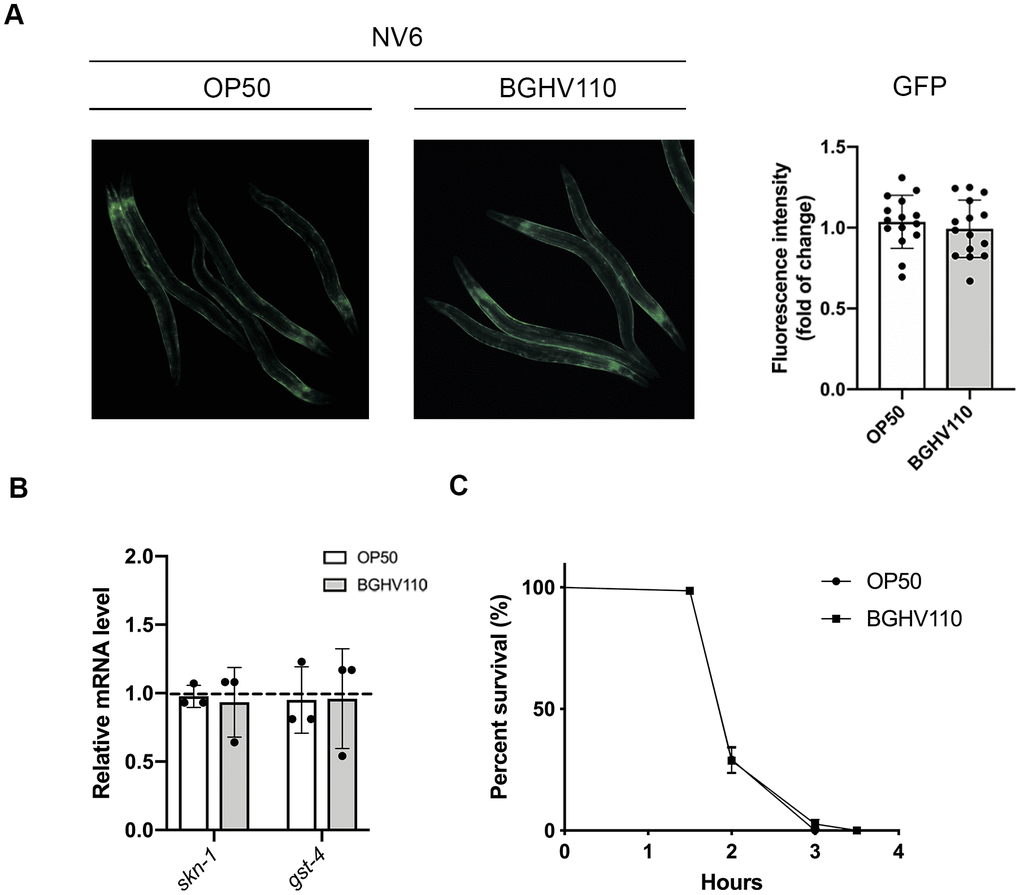
Figure 5. Heat-inactivated Lb. fermentum BGHV110 effects on the SKN-1 detoxification pathway. GFP fluorescence intensity was visualized by fluorescence microscopy (A, left panel) and intensity was quantified (A, right panel) in day 1 adult NV6 transgenic animals expressing GFP under the gst-4 promoter after overnight BGHV110 treatment (n=10, results are representative of 3 independent assays). (B) Expression of skn-1 and gst-4 genes was measured by qRT-PCR in L4 stage WT animals after 6 h of treatment with BGHV110 (n=3, three independent experiments). (C) Overnight OP50 and BGHV110 treated WT animals were exposed to 20 mM H2O2 on day 1 of adulthood and assayed for survival 4 h later. All values are presented as mean ± SD. Student’s t-test was used to compare the treated group relative to control. The log-rank (Mantel-Cox) test was used to assess the p-value in the H2O2 assay.
Heat-inactivated Lb. fermentum BGHV110 slightly affects mitochondrial dynamics without triggering the UPRmt response
A recent study showed that five mutants of E. coli with different changes in bacterial metabolism increase longevity in C. elegans, mainly by regulating mitochondrial dynamics and the mitochondrial unfolded protein response (UPRmt) [3]. Therefore, we evaluated the expression levels of heat shock protein 60 (HSP-60), as a commonly used marker of UPRmt activation. However, no differences in HSP-60 expression were observed between the OP50 and the BGHV110 fed worms (Figure 6A, 6B). Assessment of mitochondrial homeostasis was performed by analyzing steady-state levels of NADH dehydrogenase [ubiquinone] iron-sulfur protein 3 (NDUFS3) and ATP synthase subunit 5 (ATP5A), subunits of the mitochondrial respiratory (OXPHOS) complexes I and V, respectively. The obtained results indicated that feeding with BGHV110 increased the amount of the OXPHOS subunits, ATP5A significantly (Figure 6A, 6B). To further address mitochondrial involvement in the health-promoting potential of BGHV110, we analyzed the transcription of the mtDNA encoded subunit I (cox-1) and the genes involved in mtDNA replication (mtss-1 and polg-1). We detected elevated transcript levels of all tested genes after 1 day and 5 days of treatment with BGHV110, suggesting higher mitochondrial activity in the worms treated with BGHV110 (Figure 6C, 6D). This result was in accordance with the elevated protein level of complex V, supporting the conclusion that BGHV110 does not activate the UPRmt protective response, but affects mitochondrial homeostasis, most likely through the modulation of mitochondrial biogenesis.
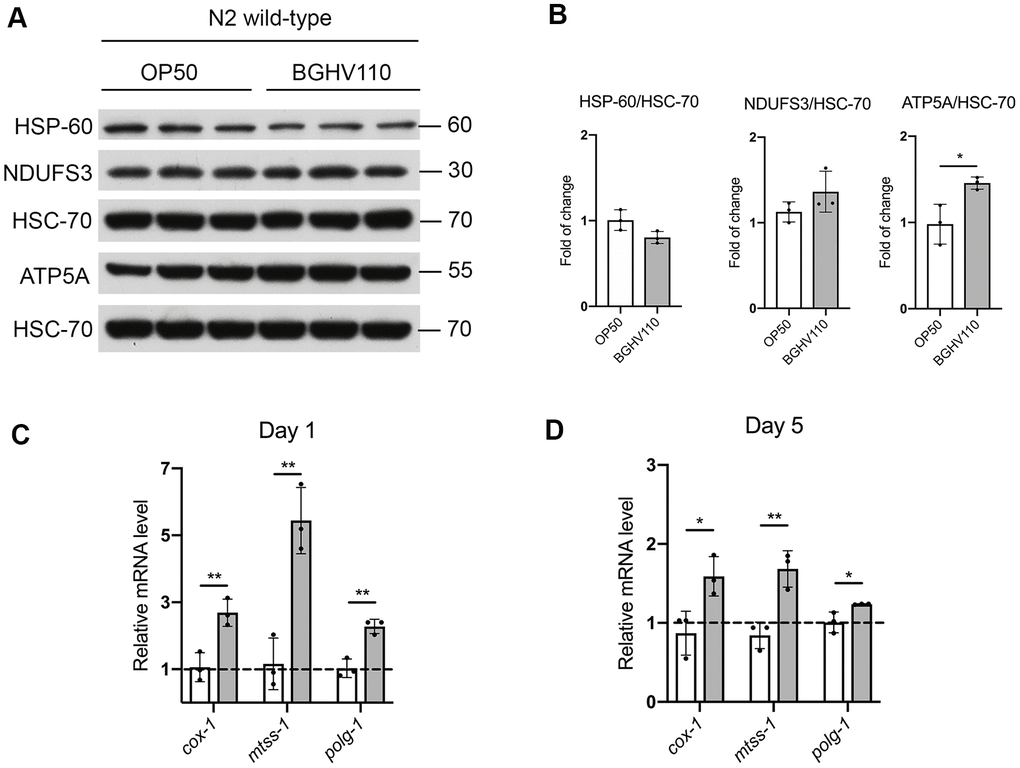
Figure 6. Mitochondrial homeostasis and the UPRmt assessment. (A) Western blots and (B) densitometric analysis showing levels of HSP-60, ATP5A and NDUFS3 proteins isolated from WT animals on day 1 of adulthood after overnight treatment with heat-inactivated BGHV110. HSC-70 was used as a loading control (n=3, three independent experiments). Expression of cox-1, mtss-1 and polg-1 genes was measured by qRT-PCR after (C) 1 day and (D) 5 days of treatment with heat-inactivated BGHV110 (n=3, three independent experiments). All values are presented as mean ± SD. Student’s t-test was used to compare the treated group relative to control (*p < 0.05; **p < 0.01).
Discussion
Our data provide strong evidence that feeding with the human commensal Lb. fermentum BGHV110 strain, as a postbiotic, extends the lifespan and fitness of C. elegans. Further, we identified the HLH-30/TFEB transcription factor as the main regulator of the longevity program initiated by feeding with the BGHV110 commensal bacteria, while disputing the roles of oxidative stress, mitochondrial dysfunction and lipid metabolism in this process.
In the general sense, the probiotic–host cross-talk is reflected in the interaction between bacterial cell surface and secreted macromolecules [26, 27], as well as metabolites [15] that can interact with the host receptors. Indeed, we have previously demonstrated the cytoprotective effect of cell surface macromolecules of the BGHV110 postbiotic in a mammalian cell culture [14]. Here, we focused instead on their potential anti-aging effect on the whole organism by using C. elegans as an in vivo model. Previous studies have shown that feeding C. elegans with heat-, UV- or antibiotic-treated E. coli could extend the lifespan by preventing bacterial proliferation and decreasing bacterial metabolism in the gut [28, 29]. This so-called postbiotic concept, which includes replacing live bacteria with dead ones or with defined bacterial metabolites, lowers the risks of high immune stimulating potential. Furthermore, the administration of postbiotics ensures stable biological effects that do not depend on live bacteria or changes in bacterial metabolism [30, 31]. Taking into account that, in this study, postbiotics obtained from heat-inactivated cells of BGHV110 were used, it seems that the observed prolongation of C. elegans lifespan could be a consequence of the longevity signals coming from bacterial surface molecules or some heat-resistant microbial metabolites with retained bioactivity. This conclusion is consistent with a recent study showing that feeding with both live and dead Lactobacillus gasseri SBT2055 increased the lifespan of C. elegans, indicating that the functional molecule is most likely a surface biomolecule or some metabolite with stable bioactivity [13].
HLH-30/TFEB is the master regulator of lysosomal biogenesis that controls the expression of lysosomal hydrolases, membrane proteins and other genes involved in autophagy [32]. The tendency of HLH-30/TFEB to respond to bacterial infection during an innate immune response to Staphylococcus aureus infection in C. elegans has been previously described [19]. Also, it has been shown that E. coli activates TFEB and enhances lysosomal function via TLR4 in mouse macrophages [33], confirming that TFEB could be activated as a result of bacterial–host interaction. Consistently, an interesting hypothesis has proposed that cells use the TFEB pathway as an additional mechanism to activate innate immunity and autophagy in response to bacteria [34]. Our data collected from WT animals and the hlh-30 mutant indicate that BGHV110 activates HLH-30/TFEB and its downstream targets to promote lifespan and maintain locomotor function in C. elegans. Our results pointed to autophagy as a key pathway responsible for the longevity program initiated in C. elegans upon feeding with BGHV110. These findings are completely comparable with the upregulation of autophagy and prolonged lifespan in C. elegans induced by the anti-aging drug verapamil [35]. Given the crucial role of nutrient deprivation in the induction of autophagy and lifespan determination [36], it was important to distinguish between the upregulation of autophagy due to possible lower intake of BGHV110 and the autophagy promoted by bacterial biomolecules. Although it is very challenging to separate the nutritional from the signaling contribution of BGHV110, the elevated transcript levels of autophagy-related genes induced by the heat-inactivated fractions of BGHV110 and the comparable levels of pharyngeal pumping rate and bacteria intestinal load between control and treatment groups pointed that DR was not responsible for the autophagy induction in C. elegans. Moreover, DR-related effects were not observed in the hlh-30 mutant, which is most likely a consequence of the similar caloric value of Escherichia and Lactobacillus species [37] and not the lack of nutrients. However, as changes in amino acid levels represent one of the multiple stressors that can activate TFEB [38, 39], it is possible to expect that the difference in amino acid content between E. coli and Lb. fermentum species could be a contributing factor that stimulated the HLH-30/TFEB activation.
The HLH-30/TFEB complex signaling network might affect lipid metabolism by regulating lipophagy [18, 40]. A recent study monitored lipid accumulation as another feature of aging and showed that lactobacilli could decrease the amount of lipid droplets [41]. This observation is in accordance with our results, showing that feeding with BGHV110 decreases lipid levels, as well as the activation of lipases. However, the changes in lipid metabolism were uncoupled from the increased longevity. We propose that differences in the macronutrients of bacterial strains, as well as the different lipl gene expression signature, may be responsible for the difference in fat levels observed in WT and the hlh-30 mutant animals fed on OP50 and BGHV110 bacterial strains. This is in agreement with the study that showed that feeding on different E. coli strains regulates fat storage levels in worms, but does not correlate with changes in lifespan [42].
Although the result regarding the potential of lactobacilli to activate the SKN-1 singling pathway in C. elegans is negative, the present study demonstrates that the HLH-30-controlled cytoprotective genes potentially overlap with changes in mitochondrial dynamics, which correlates with the growing body of evidence reporting on the emerging role of TFEB in mitochondrial quality control [43].
Overall, our findings uncover an additional mechanism of the host protective program initiated in C. elegans by the heat-inactivated Lb. fermentum BGHV110, which is important for the selection of probiotics. In view of this study, exploitation of commensal-induced HLH-30/TFEB-mediated autophagy could be further investigated in order to better understand the potential of using probiotics to modulate HLH-30/TFEB activity and promote host lifespan.
Materials and Methods
C. elegans maintenance and strains
The following strains were used in the study: the wild-type N2 (Bristol), DA2123 adIs2122 [plgg-1::gfp::lgg-1], JIN1375 hlh-30 (tm1978), NV6 [pgst-4::nls-gfp]. All nematodes were maintained at 20° C using standard growing protocols [44]. Age-synchronous worms were grown to the L4 stage on nematode growth medium (NGM) plates seeded with a live E. coli OP50 strain followed by transfer to NGM plates containing heat-inactivated OP50, heat-inactivated Lb. fermentum BGHV110 or heat-inactivated OP50 supplemented with heat-inactivated Lb. fermentum BGHV110 in a 1:2 ratio for different treatments.
Bacteria preparation and treatment
Lb. fermentum BGHV110 strain from the collection of the Laboratory of Molecular Microbiology, Institute of Molecular Genetics and Genetic Engineering, University of Belgrade, was used in this study. The strain was grown overnight at 37° C in deMan-Rogosa-Sharpe (MRS) broth (Sigma-Aldrich) under anaerobic conditions using Anaerocult A (Merck, Darmstadt, Germany). The E. coli strain OP50 was cultivated overnight in LB medium at 37° C with shaking/aerobic conditions. For the treatments, BGHV110 cells were pelleted by centrifugation at 5000 × g for 10 min at room temperature and washed twice in phosphate-buffered saline (PBS). The washed BGHV110 cells were resuspended in the same volume of LB medium as OP50 and heat-inactivated at 70° C for 70 min. Heat-inactivation was performed for OP50 under the same conditions. Efficacy of heat inactivation was checked by inoculation of bacterial suspension (10 μl) on MRS/LB agar plates and incubation overnight at 37° C. The heat-inactivated bacterial suspensions, alone or in mixture, were spread on appropriate NGM plates and dried at room temperature.
Acridine orange staining
Heat-inactivated bacterial suspensions were centrifuged (10000 × g for 5 min) and resuspended in PBS containing acridine orange dye (Sigma-Aldrich) in a final concentration of 10 μM. After 15 min, the suspensions were centrifuged to remove dye and washed three times in PBS. Bacterial pellets were resuspended in LB medium, spread on NGM plates and dried at room temperature. After overnight treatment, fluorescence was examined microscopically.
Lifespan analysis
For the lifespan analysis, 25 worms per 3.5 cm plate in the L4 developmental stage were transferred to OP50 or BGHV110 NGM plates containing 20 μM of 5-Fluorodeoxyuridine (FudR, Sigma-Aldrich) to avoid progeny hatching. In total, 100 worms (4 plates) were used per condition. First day of adulthood was defined as day 1 in lifespan analysis. Animals were examined every second day by prodding with a silver wire and live worms were transferred to fresh plates. The worms that escaped, or died due to internal hatching or protrusions, were censored.
Movement
Body bending analysis was performed on day 5 and day 10 of adulthood by transferring worms to non-seeded 9 cm NGM plates. After a short incubation period, the body bending of worms was assessed as the number of full sinusoidal curves the worms made by moving forward or backward. The results are presented as the number of body bends counted over a period of 3 min.
Pharyngeal pumping assay
Pharyngeal pumping analysis was performed on day 5 and day 10 of adulthood by counting the number of the pharynx rear bulb movements. During the experiment, the animals were placed in bacterial lawns using silver wire. The results are presented as the number of pumping rates in a period of 15 s.
RNA isolation and quantitative real-time PCR (qRT-PCR)
Total RNA was isolated from approximately 200 worms using a Trizol reagent (Invitrogen). DNase I treatment was performed using an Ambion DNA-free™ Kit (Thermo Fisher Scientific). Reversed transcription was done using 1 μg of isolated RNA as a template, according to the manufacturer′s protocol (Thermo Fisher Scientific). Random hexamers (Applied Biosystems) and RiboLock RNase inhibitor (Thermo Fisher Scientific) were used in the reactions. Synthesized cDNA was further amplified in a 7500 real-time PCR system (Applied Biosystems) using SYBR™ Green PCR Master Mix (Applied Biosystems) under the following conditions: 10 min at 95° C activation, 40 cycles of 15 s at 95° C and 60 s at 60° C. The results were normalized against the act-1 gene [13] and expressed as relative target abundance using the 2-ΔΔCt method [45]. Primers used in the study are presented in Supplementary Table 1. All primers were purchased from Thermo Fisher Scientific. For each treatment, three independent replicates were used.
Western blotting
For protein extraction, approximately 200 worms were collected from 9 cm plates using M9 buffer and washed three times to remove the remaining bacteria. Protein isolation was performed as described previously by Herholz et al. [46]. Protein concentration was measured with the Bradford assay (Thermo Fisher Scientific). The extracted proteins (30 μg) were separated on 12% SDS–PAGE and transferred to a 0.2 mm nitrocellulose membrane (GE Healthcare). Western blotting was performed overnight at 4° C with antibodies against: GFP (1:2000, kindly provided by Jan Riemer), HSC-70 (1:2000, Santa Cruz, #sc-7298), HSP-60 (1:2000, BD Transduction Laboratories, #611562), NDUFS3 (1:1000, Mitosciences, MS112) and ATP5A (1:1000, Mitosciences, MS507). The intensity of the bands was quantified using ImageJ (National Institutes of Health) software. For each treatment, three independent replicates were used.
Microscopy
Worms were immobilized on 2% agarose pads using 5 mM levamisole buffer and observed under an AxioImager Z.1 epifluorescence microscope. The images were collected with a Hammamatsu camera (OrcaR2) and AxioVision software 4.8. Images were analyzed using ImageJ software and normalized to OP50 control.
Oil-red O staining
For lipid staining, worms were collected from 9 cm plates using M9 buffer after 5 days of treatment. Worms were washed three times with PBS and fixed in MRWB buffer (1% paraformaldehyde, 80 mM KCl, 20 mM NaCl, 7 mM EGTA, 0.5 mM spermidine-HCl, 0.2 mM spermine, 15 mM PIPES, 0.1% 2-mercaptoethanol) for 1 h. Then, the worms were dehydrated in 60% isopropanol for 15 min and stained with filtered Oil Red O solution (5 mg/ml Oil Red O in 60% isopropanol) for 3 h. After the incubation, the dye was removed and the worms were resuspended in PBS containing 0.01% Triton X-100 and visualized using a DM IL LED inverted microscope (Leica). Relative dye intensity was quantified using ImageJ software.
H2O2 resistance assay
Following overnight treatments, worms were picked into 96-well plates filled with M9 buffer containing 20 mM H2O2 in a final concentration. For both conditions, 12 wells of 6 worms were scored for survival every hour. Results are presented as percentage of survival during 4 h of H2O2 treatment.
Statistical analysis
All data are presented as mean values ± standard deviation (SD). The differences between control and experimental groups were compared using Student’s t-test. One-way ANOVA followed by Tukey post hoc test for multiple comparisons was used in the lipid staining analysis. The differences between survival curves in lifespan and H2O2 analysis were assessed using the log-rank (Mantel-Cox) test. A p value less than 0.05 was considered statistically significant. The statistical analysis was performed and graphs were prepared using GraphPad Prism 8 software.
Supplementary Materials
Author Contributions
MD, MH and AT designed the study. MD, MH, UK, DR, KN and JĐ performed the experiments. MD, MH, JĐ, AT, NG interpreted findings. MD and DR prepared figures. MD wrote the manuscript. AT and NG supervised the work. All authors read and approved the manuscript.
Acknowledgments
The authors would like to thank Maja Vojvodić Jovanović for reviewing the language in the manuscript.
Conflicts of Interest
The authors declare that they have no conflicts of interest.
Funding
This work was supported by the Ministry of Education, Science and Technological Development of the Republic of Serbia (Grant No. 451-03-9/2021-14/200042). The study was further supported by A. Trifunovic’s grant of the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation - SFB 1218 – Projektnummer 269925409). Miroslav Dinić received a Research and Training grant (FEMS-GO-2017-016) from the Federation of European Microbiological Societies (FEMS).
References
- 1. Biagi E, Candela M, Turroni S, Garagnani P, Franceschi C, Brigidi P. Ageing and gut microbes: perspectives for health maintenance and longevity. Pharmacol Res. 2013; 69:11–20. https://doi.org/10.1016/j.phrs.2012.10.005 [PubMed]
- 2. Maynard C, Weinkove D. The gut microbiota and ageing. Subcell Biochem. 2018; 90:351–71. https://doi.org/10.1007/978-981-13-2835-0_12 [PubMed]
- 3. Han B, Sivaramakrishnan P, Lin CJ, Neve IA, He J, Tay LW, Sowa JN, Sizovs A, Du G, Wang J, Herman C, Wang MC. Microbial genetic composition tunes host longevity. Cell. 2017; 169:1249–62.e13. https://doi.org/10.1016/j.cell.2017.05.036 [PubMed]
- 4. Le Roy CI, Štšepetova J, Sepp E, Songisepp E, Claus SP, Mikelsaar M. New insights into the impact of Lactobacillus population on host-bacteria metabolic interplay. Oncotarget. 2015; 6:30545–56. https://doi.org/10.18632/oncotarget.5906 [PubMed]
- 5. Kong F, Deng F, Li Y, Zhao J. Identification of gut microbiome signatures associated with longevity provides a promising modulation target for healthy aging. Gut Microbes. 2019; 10:210–15. https://doi.org/10.1080/19490976.2018.1494102 [PubMed]
- 6. Bischoff SC. Microbiota and aging. Curr Opin Clin Nutr Metab Care. 2016; 19:26–30. https://doi.org/10.1097/MCO.0000000000000242 [PubMed]
- 7. Grompone G, Martorell P, Llopis S, González N, Genovés S, Mulet AP, Fernández-Calero T, Tiscornia I, Bollati-Fogolín M, Chambaud I, Foligné B, Montserrat A, Ramón D. Anti-inflammatory Lactobacillus rhamnosus CNCM I-3690 strain protects against oxidative stress and increases lifespan in Caenorhabditis elegans. PLoS One. 2012; 7:e52493. https://doi.org/10.1371/journal.pone.0052493 [PubMed]
- 8. Park MR, Ryu S, Maburutse BE, Oh NS, Kim SH, Oh S, Jeong SY, Jeong DY, Oh S, Kim Y. Probiotic Lactobacillus fermentum strain JDFM216 stimulates the longevity and immune response of Caenorhabditis elegans through a nuclear hormone receptor. Sci Rep. 2018; 8:7441. https://doi.org/10.1038/s41598-018-25333-8 [PubMed]
- 9. Schifano E, Zinno P, Guantario B, Roselli M, Marcoccia S, Devirgiliis C, Uccelletti D. The foodborne strain Lactobacillus fermentum MBC2 triggers pept-1-dependent pro-longevity effects in Caenorhabditis elegans. Microorganisms. 2019; 7:45. https://doi.org/10.3390/microorganisms7020045 [PubMed]
- 10. Heintz C, Mair W. You are what you host: microbiome modulation of the aging process. Cell. 2014; 156:408–11. https://doi.org/10.1016/j.cell.2014.01.025 [PubMed]
- 11. Sugawara T, Sakamoto K. Killed Bifidobacterium longum enhanced stress tolerance and prolonged life span of Caenorhabditis elegans via DAF-16. Br J Nutr. 2018; 120:872–80. https://doi.org/10.1017/S0007114518001563 [PubMed]
- 12. Kim Y, Mylonakis E. Caenorhabditis elegans immune conditioning with the probiotic bacterium Lactobacillus acidophilus strain NCFM enhances gram-positive immune responses. Infect Immun. 2012; 80:2500–08. https://doi.org/10.1128/IAI.06350-11 [PubMed]
- 13. Nakagawa H, Shiozaki T, Kobatake E, Hosoya T, Moriya T, Sakai F, Taru H, Miyazaki T. Effects and mechanisms of prolongevity induced by Lactobacillus gasseri SBT2055 in Caenorhabditis elegans. Aging Cell. 2016; 15:227–36. https://doi.org/10.1111/acel.12431 [PubMed]
- 14. Dinić M, Lukić J, Djokić J, Milenković M, Strahinić I, Golić N, Begović J. Lactobacillus fermentum postbiotic-induced autophagy as potential approach for treatment of acetaminophen hepatotoxicity. Front Microbiol. 2017; 8:594. https://doi.org/10.3389/fmicb.2017.00594 [PubMed]
- 15. Bajić SS, Đokić J, Dinić M, Tomić S, Popović N, Brdarić E, Golić N, Tolinački M. GABA potentiate the immunoregulatory effects of Lactobacillus brevis BGZLS10-17 via ATG5-dependent autophagy in vitro. Sci Rep. 2020; 10:1347. https://doi.org/10.1038/s41598-020-58177-2 [PubMed]
- 16. Megalou EV, Tavernarakis N. Autophagy in Caenorhabditis elegans. Biochim Biophys Acta. 2009; 1793:1444–51. https://doi.org/10.1016/j.bbamcr.2008.12.010 [PubMed]
- 17. Lapierre LR, De Magalhaes Filho CD, McQuary PR, Chu CC, Visvikis O, Chang JT, Gelino S, Ong B, Davis AE, Irazoqui JE, Dillin A, Hansen M. The TFEB orthologue HLH-30 regulates autophagy and modulates longevity in Caenorhabditis elegans. Nat Commun. 2013; 4:2267. https://doi.org/10.1038/ncomms3267 [PubMed]
- 18. O’Rourke EJ, Ruvkun G. MXL-3 and HLH-30 transcriptionally link lipolysis and autophagy to nutrient availability. Nat Cell Biol. 2013; 15:668–76. https://doi.org/10.1038/ncb2741 [PubMed]
- 19. Visvikis O, Ihuegbu N, Labed SA, Luhachack LG, Alves AF, Wollenberg AC, Stuart LM, Stormo GD, Irazoqui JE. Innate host defense requires TFEB-mediated transcription of cytoprotective and antimicrobial genes. Immunity. 2014; 40:896–909. https://doi.org/10.1016/j.immuni.2014.05.002 [PubMed]
- 20. Aguilar-Toalá JE, Garcia-Varela R, Garcia HS, Mata-Haro V, González-Córdova AF, Vallejo-Cordoba B, Hernández-Mendoza A. Postbiotics: An evolving term within the functional foods field. Trends Food Sci Technol. 2018; 75:105–14. https://doi.org/10.1016/j.tifs.2018.03.009
- 21. Nakamura S, Yoshimori T. Autophagy and longevity. Mol Cells. 2018; 41:65–72. https://doi.org/10.14348/molcells.2018.2333 [PubMed]
- 22. Gelino S, Chang JT, Kumsta C, She X, Davis A, Nguyen C, Panowski S, Hansen M. Intestinal Autophagy Improves Healthspan and Longevity in C. elegans during Dietary Restriction. PLoS Genet. 2016; 12:e1006135. https://doi.org/10.1371/journal.pgen.1006135 [PubMed]
- 23. Chen Y, Scarcelli V, Legouis R. Approaches for studying autophagy in Caenorhabditis elegans. Cells. 2017; 6:27. https://doi.org/10.3390/cells6030027 [PubMed]
- 24. Lapierre LR, Gelino S, Meléndez A, Hansen M. Autophagy and lipid metabolism coordinately modulate life span in germline-less C. Elegans. Curr Biol. 2011; 21:1507–14. https://doi.org/10.1016/j.cub.2011.07.042 [PubMed]
- 25. Lu Q, Yang P, Huang X, Hu W, Guo B, Wu F, Lin L, Kovács AL, Yu L, Zhang H. The WD40 repeat PtdIns(3)P-binding protein EPG-6 regulates progression of omegasomes to autophagosomes. Dev Cell. 2011; 21:343–57. https://doi.org/10.1016/j.devcel.2011.06.024 [PubMed]
- 26. Dinić M, Pecikoza U, Djokić J, Stepanović-Petrović R, Milenković M, Stevanović M, Filipović N, Begović J, Golić N, Lukić J. Exopolysaccharide produced by probiotic strain Lactobacillus paraplantarum BGCG11 reduces inflammatory hyperalgesia in rats. Front Pharmacol. 2018; 9:1. https://doi.org/10.3389/fphar.2018.00001 [PubMed]
- 27. Popović N, Djokić J, Brdarić E, Dinić M, Terzić-Vidojević A, Golić N, Veljović K. The influence of heat-killed Enterococcus faecium BGPAS1-3 on the tight junction protein expression and immune function in differentiated Caco-2 cells infected with Listeria monocytogenes ATCC 19111. Front Microbiol. 2019; 10:412. https://doi.org/10.3389/fmicb.2019.00412 [PubMed]
- 28. Garigan D, Hsu AL, Fraser AG, Kamath RS, Ahringer J, Kenyon C. Genetic analysis of tissue aging in Caenorhabditis elegans: a role for heat-shock factor and bacterial proliferation. Genetics. 2002; 161:1101–12. [PubMed]
- 29. Lee J, Choe J, Kim J, Oh S, Park S, Kim S, Kim Y. Heat-killed Lactobacillus spp. Cells enhance survivals of Caenorhabditis elegans against Salmonella and Yersinia infections. Lett Appl Microbiol. 2015; 61:523–30. https://doi.org/10.1111/lam.12478 [PubMed]
- 30. Tsilingiri K, Rescigno M. Postbiotics: what else? Benef Microbes. 2013; 4:101–07. https://doi.org/10.3920/BM2012.0046 [PubMed]
- 31. Tsilingiri K, Barbosa T, Penna G, Caprioli F, Sonzogni A, Viale G, Rescigno M. Probiotic and postbiotic activity in health and disease: comparison on a novel polarised ex-vivo organ culture model. Gut. 2012; 61:1007–15. https://doi.org/10.1136/gutjnl-2011-300971 [PubMed]
- 32. Napolitano G, Ballabio A. TFEB at a glance. J Cell Sci. 2016; 129:2475–81. https://doi.org/10.1242/jcs.146365 [PubMed]
- 33. Gray MA, Choy CH, Dayam RM, Ospina-Escobar E, Somerville A, Xiao X, Ferguson SM, Botelho RJ. Phagocytosis enhances lysosomal and bactericidal properties by activating the transcription factor TFEB. Curr Biol. 2016; 26:1955–64. https://doi.org/10.1016/j.cub.2016.05.070 [PubMed]
- 34. Pastore N, Brady OA, Diab HI, Martina JA, Sun L, Huynh T, Lim JA, Zare H, Raben N, Ballabio A, Puertollano R. TFEB and TFE3 cooperate in the regulation of the innate immune response in activated macrophages. Autophagy. 2016; 12:1240–58. https://doi.org/10.1080/15548627.2016.1179405 [PubMed]
- 35. Liu W, Lin H, Mao Z, Zhang L, Bao K, Jiang B, Xia C, Li W, Hu Z, Li J. Verapamil extends lifespan in Caenorhabditis elegans by inhibiting calcineurin activity and promoting autophagy. Aging (Albany NY). 2020; 12:5300–17. https://doi.org/10.18632/aging.102951 [PubMed]
- 36. Hansen M, Chandra A, Mitic LL, Onken B, Driscoll M, Kenyon C. A role for autophagy in the extension of lifespan by dietary restriction in C. Elegans. PLoS Genet. 2008; 4:e24. https://doi.org/10.1371/journal.pgen.0040024 [PubMed]
- 37. Prochazka GJ, Payne WJ, Mayberry WR. Calorific content of certain bacteria and fungi. J Bacteriol. 1970; 104:646–49. https://doi.org/10.1128/JB.104.2.646-649.1970 [PubMed]
- 38. Ren W, Rajendran R, Zhao Y, Tan B, Wu G, Bazer FW, Zhu G, Peng Y, Huang X, Deng J, Yin Y. Amino acids as mediators of metabolic cross talk between host and pathogen. Front Immunol. 2018; 9:319. https://doi.org/10.3389/fimmu.2018.00319 [PubMed]
- 39. Tattoli I, Sorbara MT, Vuckovic D, Ling A, Soares F, Carneiro LA, Yang C, Emili A, Philpott DJ, Girardin SE. Amino acid starvation induced by invasive bacterial pathogens triggers an innate host defense program. Cell Host Microbe. 2012; 11:563–75. https://doi.org/10.1016/j.chom.2012.04.012 [PubMed]
- 40. Settembre C, Ballabio A. Lysosome: regulator of lipid degradation pathways. Trends Cell Biol. 2014; 24:743–50. https://doi.org/10.1016/j.tcb.2014.06.006 [PubMed]
- 41. Johnson AA, Stolzing A. The role of lipid metabolism in aging, lifespan regulation, and age-related disease. Aging Cell. 2019; 18:e13048. https://doi.org/10.1111/acel.13048 [PubMed]
- 42. Brooks KK, Liang B, Watts JL. The influence of bacterial diet on fat storage in C. Elegans. PLoS One. 2009; 4:e7545. https://doi.org/10.1371/journal.pone.0007545 [PubMed]
- 43. Wang S, Chen Y, Li X, Zhang W, Liu Z, Wu M, Pan Q, Liu H. Emerging role of transcription factor EB in mitochondrial quality control. Biomed Pharmacother. 2020; 128:110272. https://doi.org/10.1016/j.biopha.2020.110272 [PubMed]
- 44. Brenner S. The genetics of Caenorhabditis elegans. Genetics. 1974; 77:71–94. https://doi.org/10.1093/genetics/77.1.71 [PubMed]
- 45. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods. 2001; 25:402–08. https://doi.org/10.1006/meth.2001.1262 [PubMed]
- 46. Herholz M, Cepeda E, Baumann L, Kukat A, Hermeling J, Maciej S, Szczepanowska K, Pavlenko V, Frommolt P, Trifunovic A. KLF-1 orchestrates a xenobiotic detoxification program essential for longevity of mitochondrial mutants. Nat Commun. 2019; 10:3323. https://doi.org/10.1038/s41467-019-11275-w [PubMed]


