Introduction
Hepatocellular carcinoma (HCC) accounts for 70–85% of all primary liver cancer cases and is one of the most common malignant tumors in the world [1]. The 5-year recurrence rate is more than 60% and median 5-year survival rates are 35.2%, 48.3%, and 15.5% after repeat hepatectomy, ablation, and transarterial chemoembolization (TACE), respectively [2, 3]. The development of HCC involves several epigenetic and genetic changes; environmental factors such as cytotoxic and DNA-damaging chemicals, and chronic infections with hepatitis B virus (HBV) and hepatitis C virus (HCV) are common risk factors associated with HCC [4–6]. HCC is associated with aberrant activities of several signaling pathways such as Wnt/β-catenin, PI3K/AKT/GSK-3β, MAPK/ERK, TGFß/BMP, mTOR, and Stat3, as well as mutations in p53, K-Ras, c-Myc, HBx, p16INK4A, APC, BRCA2, CDK2, cyclin E1, cyclin D, cyclin A and p38γ genes [4–10]. However, several factors involved in the molecular pathogenesis and development of HCC are unknown and remain under extensive investigation.
Cripto-1 (CR-1) is a member of the epidermal growth factor (EGF)-Cripto-1/fibroblast growth factor related ligand (FRL1)/Criptic (EGF-CFC) protein family that coordinates primitive streak formation, mesoderm and endoderm specification, and orientation of anterior and posterior (A/P) axis during embryogenesis; it is a known marker of undifferentiated embryonic stem cells [11–16]. CR-1 is either undetected or expressed at very low levels in normal adult tissues, but is highly expressed in nasopharyngeal carcinoma and colon, lung, stomach, and breast cancers [11–16]. Aberrant activation of embryonic genes such as Oct-4 in adult tissues is frequently associated with several cancers [17–19]. Ectopic expression of the CR-1 transgene in mammary glands induced mammary hyperplasia and adenocarcinoma in the WAP-CR-1 or MMTV-CR-1 transgenic mice [15, 20] and leiomyosarcoma of the uterus in the MMTV-CR-1 transgenic mice [21]. CR-1 plays an oncogenic role in colorectal cancer [22], melanoma [23], and esophageal squamous cell carcinoma [24].
Several studies have reported that CR-1 is associated with HCC progression [25–29]. Wang et al. demonstrated that elevated expression of CR-1 in tumor tissues was associated with high aggressiveness and poor prognosis of HCC patients [26]. Zhang et al. reported that serum CR-1 levels were significantly higher in patients with chronic hepatitis, cirrhosis, and HCC compared to healthy individuals; moreover, serum CR-1 levels were significantly higher in HBV-related HCC compared to HCV-related HCC [27]. Lo et al. reported that CR-1 positively modulated growth, tumorigenicity, invasion and chemoresistance of HCC cells by promoting stemness through stabilization of Dishevelled-3 and activation of the Wnt/β-catenin signaling pathway [25]. These findings indirectly suggested that CR-1 regulated HCC. However, direct evidence of the oncogenic role and underlying mechanisms of CR-1 in hepatocarcinogenesis are not known. Therefore, in this study, we investigated the mechanistic role of CR-1 in HCC using transgenic mice expressing hepatocyte-specific CR-1.
Results
Truncated CR-1 mRNA is predominantly expressed in HCC cell lines and tissues
Previous studies reported that mammalian cells and tissues expressed both full length (FL-CR-1; 2.0 kb) and truncated (SF-CR-1; 1.7 kb) CR-1 mRNAs [30, 31]. Therefore, we performed RT-PCR analysis to quantify FL-CR-1 and SF-CR1 mRNA levels in HCC cell lines and tissues using primer sets that specifically detect FL-CR1 mRNA (UND/UNB) and total CR-1 mRNA (UNA/UNB) (Figure 1A). RT-PCR analysis showed that all six HCC cell lines (QGY7701, Hep3B, HepG2, SK-Hep-1, SNU-387 and SNU-182) expressed both FL-CR1and SF-CR1 mRNAs (Figure 1B). FL-CR-1 was highly expressed in the SK-Hep-1 cells (62%), but was lowly expressed (less than 16%) in the other HCC cell lines (QGY7701, HepG2, Hep3B, SNU-387 and SNU-182 cells) (Figure 1C). This demonstrated that SF-CR-1 was the predominant CR-1 transcript in the QGY7701, HepG2, Hep3B, SNU-182 and SNU-387 cell lines.
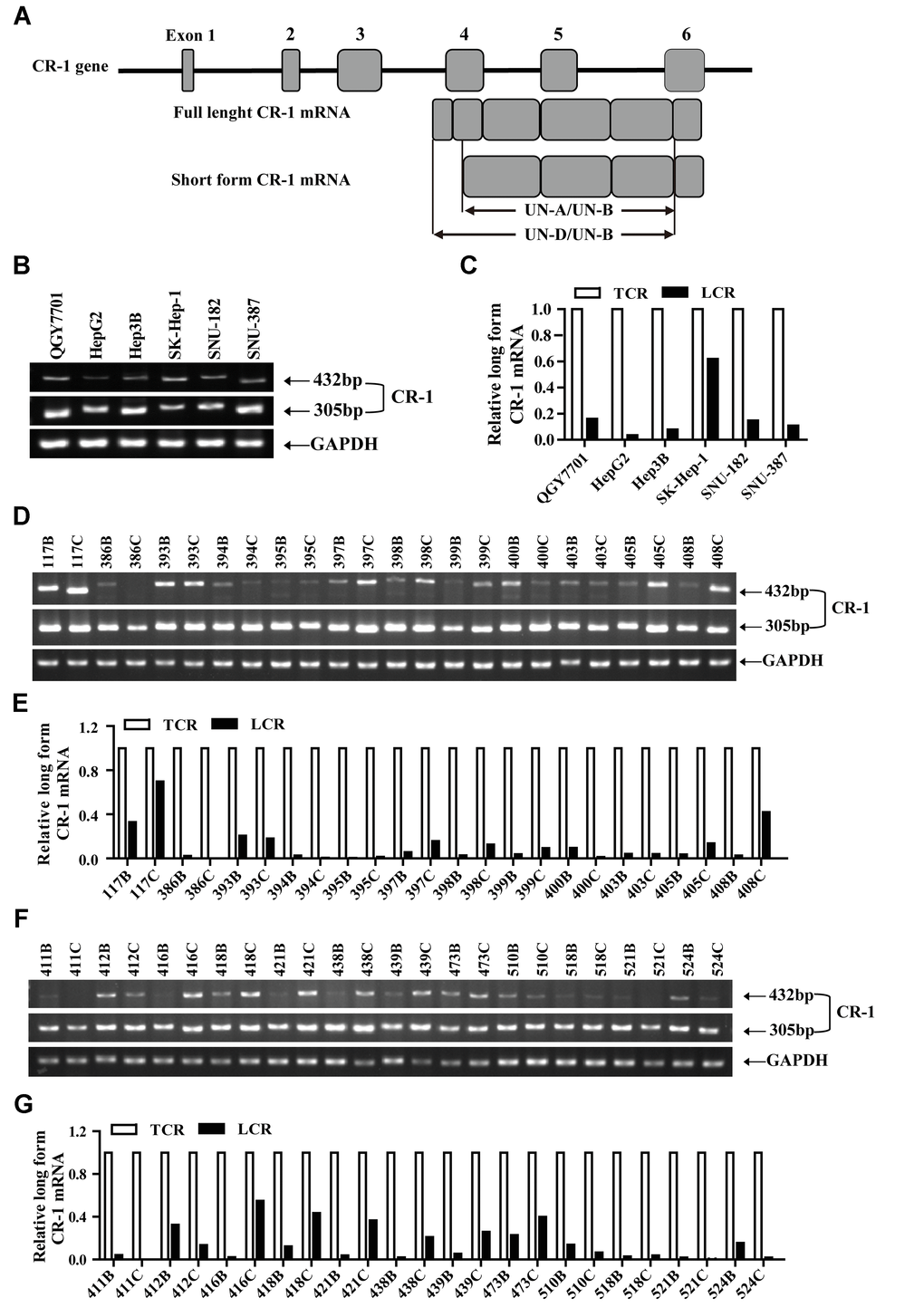
Figure 1. The short form Cripto-1 (CR-1) mRNA is predominantly expressed in HCC cell lines and tissues. (A) Diagrammatic representation shows structure of the CR-1 gene and mRNAs (full-length and short-length forms). UN-D/UN-B: full-length CR-1 (FL-CR1) transcript specific primer set yields 432 bp PCR product; UN-A/UN-B: total CR-1 transcript specific primer set yields 305 bp PCR product. (B–C) RT-PCR analysis shows levels of FL-CR-1 transcript relative to total CR-1 (TCR) transcript levels in human HCC cell lines, QGY7701, Hep3B, HepG2, SK-Hep-1, SNU-387 and SNU-182. GAPDH was used as internal control. (D–G) RT-PCR analysis shows levels of FL-CR-1 transcript relative to total CR-1 transcript in human HCC (T/C) and their corresponding adjacent non-tumor liver tissues (N/B).
RT-PCR analysis also demonstrated the presence of both FL-CR1 and SF-CR1 transcripts in 24 pairs of HCC and adjacent non-tumor liver tissue biopsies (Figure 1D, 1F). Furthermore, expression levels of FL-CR-1 transcripts were significantly lower (less than 20%) in majority of the HCC tissues (n = 18, 75%) and adjacent non-tumor liver tissues (n = 22, 92%) compared to SF-CR-1 levels (Figure 1E, 1G). Collectively, these data suggested that SF-CR-1 was the predominant CR-1 transcript in majority of the HCC cell lines, HCC and adjacent non-tumor liver tissues.
CR-1 is significantly upregulated in HCC tissues
We then assessed the potential role of CR-1 in HCC progression. RT-PCR analysis demonstrated that CR-1 transcripts were significantly up-regulated in HCC tissues compared to adjacent non-tumor liver tissues (Figure 2A, 2B). Furthermore, qRT-PCR analysis confirmed that relative CR-1 mRNA expression levels were significantly higher in HCC tissues compared to the adjacent non-tumor liver tissues (Figure 2C). Moreover, CR-1 mRNA levels were up-regulated in HCC tissues from the TCGA datasets compared to the non-tumor liver tissues (Figure 2D). These results demonstrated that CR-1 expression was significantly upregulated in HCC tissues.
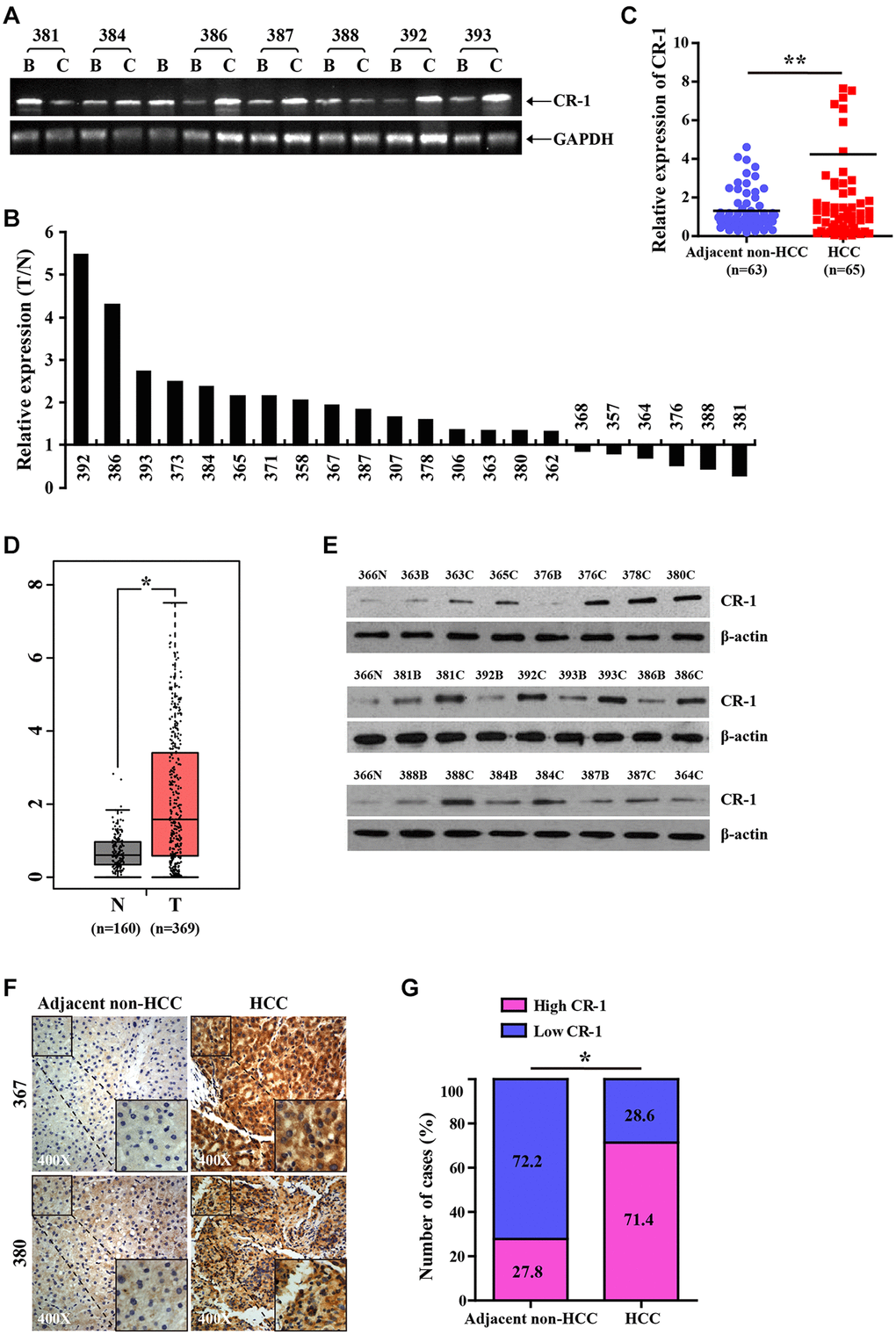
Figure 2. CR-1 expression is significantly upregulated in HCC tissues. (A) RT-PCR analysis of CR-1 transcript levels in HCC tissues (T) and matched adjacent non-tumor liver tissues (N). (B) The histogram plot shows ratio (T/N) of CR-1 mRNA levels in HCC (T) and matched non-tumor liver tissues (N) based on semi-quantitative analysis of RT-PCR data shown in Figure 2A. (C) qRT-PCR analysis shows relative levels of CR-1 transcript in 65 HCC (T) and 63 adjacent non-HCC liver tissue biopsies (N). (D) The expression levels of CR-1 transcript in HCC clinical specimens (T; n = 369) and non-cancerous liver tissue biopsies (N; n = 160) from TCGA datasets. (E) Representative western blot shows CR-1 protein expression levels in HCC (T) and adjacent non-HCC liver tissue biopsies (N). (F) Representative images show CR-1 protein expression in HCC and adjacent non-HCC liver tissue biopsies, as examined by IHC. (G) IHC assay analysis shows that CR-1 protein expression was significantly higher in majority of HCC tissues compared to the adjacent non-cancerous liver tissues (71.4% vs. 27.8%).
Western blotting analysis of 9 paired human HCC tissues, 4 unpaired human HCC tissues and 1 normal liver tissue confirmed that CR-1 protein levels were significantly upregulated in HCC tissues compared to normal liver tissues (Figure 2E). We then performed immunohistochemical staining of a tissue array with several pairs of HCC samples using a specific antibody against CR-1. The results showed that CR-1 protein levels were significantly upregulated in the HCC tissues compared to the corresponding noncancerous liver tissues (Figure 2F, 2G). These results demonstrated that CR-1 mRNA and protein levels were significantly upregulated in HCC tissues compared to the non-cancerous liver tissues.
Generation and analysis of RCLG transgenic mice
To understand the in vivo functional role of CR-1 in HCC, we generated CR-1 transgenic mice (RCLG mice) (Supplementary Figure 1) using protocols as described previously [16]. We obtained two founder mice (referred to as 190# and 220#) that strongly expressed mRFP (marker for transgenic CR-1 expression) and showed normal phenotype (Supplementary Figure 1B). In general, RCLG mice were viable and fertile, and did not manifest any gross behavioral or phenotypic abnormalities.
Next, we analyzed expression patterns of the mRFP transgene in various organs harvested from RCLG mice. MRFP expression (red fluorescence) was detected in the heart, liver, brain, lung, spleen, kidney, intestine, testis, thymus, and pancreas of the RCLG mice, but was not observed in any of the tissues from the control littermates (Supplementary Figure 2). Furthermore, mRFP expression was significantly higher in the heart, liver, kidney, lung, intestine, brain, testis, thymus, and pancreas, but was lowly expressed in the spleen (Supplementary Figure 2). Overall, mRFP was ubiquitously expressed in majority of the tissues from the RCLG mice.
Fluorescence microscopy analysis of tissue sections showed that most cells of brain, heart, kidney, lung, liver, intestine, stomach and testis from the RCLG mice were positive for mRFP expression (Supplementary Figure 3). However, some cells did not appear to be RFP positive. Therefore, we concluded that a RCLG derived mouse line was not a useful tool for investigating the role of CR-1 in HCC.
Hepatocyte-specific overexpression of CR-1 transgene in RCLG mice mediated by Cre/lox P system
To further determine whether CR-1 overexpression promoted liver oncogenesis, we generated RCLG/Alb-Cre mice by crossing RCLG mice with Alb-Cre mice, in which the Cre is under the control of the hepatocyte-specific albumin (Alb) promoter [32]. The RCLG/Alb-Cre mice showed expression of Luc and CR-1 transgenes in a liver-restricted manner (Supplementary Figure 4A). Whole-animal bioluminescence imaging demonstrated Luc activity in the liver of RCLG/Alb-Cre newborn offspring, which were also mRFP-positive (Supplementary Figure 4B, 4C). This was further confirmed by PCR-based genotyping (Supplementary Figure 4D) and whole-animal bioluminescence imaging in adult mice (Supplementary Figure 4E), thereby demonstrating that Luc expression was mediated by Alb-Cre. Organ-specific bioluminescence imaging demonstrated that Luc activity was present only in the liver of Luc-positive RCLG/Alb-Cre mice (Supplementary Figure 4E) and was absent in the other organs (Supplementary Figure 4F). Luc activity was absent in all organs from the Luc-negative mouse shown in Supplementary Figure 4E (data not shown). RT-PCR and western blotting analysis showed that CR-1 mRNA and CR-1 protein levels were significantly higher in the liver tissues derived from Luc-positive RCLG/Alb-Cre mice compared to Luc-negative mice (Supplementary Figure 4G–4H). These findings confirmed liver-specific overexpression of the CR-1 transgene in the RCLG mice.
Characterization of gross morphology and histopathology of RCLG/Alb-Cre transgenic mice
Next, we analyzed the morphological and histological differences between RCLG/Alb-Cre mice and Alb-Cre control mice to determine the functional relevance of liver-specific CR-1 overexpression. The body weights (Figure 3A, 3B) and liver weights (Figure 3C) of RCLG/Alb-Cre and control mice at 3, 6 and 8 months were similar. Moreover, we did not observe any significant differences in the gross whole body morphology of mice (Figure 3A) and livers (Figure 3D) of control and RCLG/Alb-Cre mice at 3, 6 and 8 months, respectively. H&E stained liver sections from 3, 6, and 8 month old RCLG/Alb-Cre mice did not show any abnormal liver structure, increased number of mitotic hepatocytes, liver cell dysplasia or malignant liver cells (Figure 3E). We observed similar percentages of Ki67-positive nuclei in the hepatocytes from 3, 6, and 8 month old control and RCLG/Alb-Cre mice (Figure 3F). These results demonstrated that the hepatocyte-specific expression of CR-1 transgene did not cause any liver pathology such as hepatocyte hyperplasia or hepatocellular carcinogenesis.
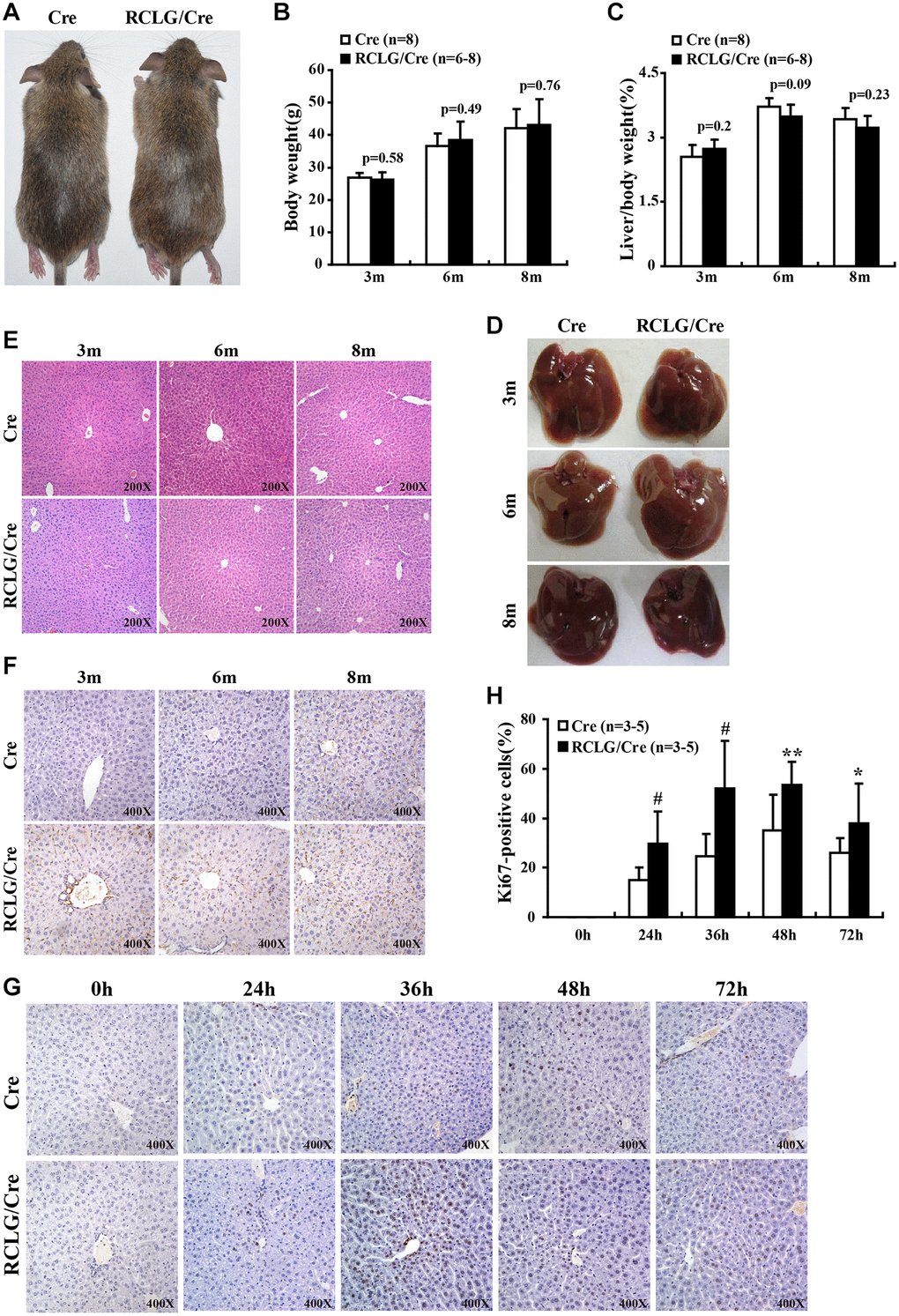
Figure 3. CR-1 overexpression enhances hepatocyte proliferation in the liver after 2/3 partial hepatectomy (PHx). (A) Representative images show 5-month old RCLG/Alb-Cre (right) and Alb-Cre (left) mice fed a normal diet. (B) Body weights of 3, 6, and 8 month-old RCLG/Alb-Cre and Alb-Cre mice. (C) Relative liver weights of 3, 6, and 8 month-old RCLG/Alb-Cre and Alb-Cre mice. (D) Representative images show gross morphology of livers from 3, 6, and 8 month-old RCLG/Alb-Cre (right) and Alb-Cre (left) mice. (E) Representative images of H&E-stained liver sections from 3, 6, and 8 month-old Alb-Cre and RCLG/Alb-Cre mice. (F) Representative Ki-67 immunostaining images show proliferation status of hepatocytes from 3, 6, and 8 month-old Alb-Cre and RCLG/Alb-Cre mice. (G) Representative Ki-67 immunostaining images show proliferation of hepatocytes from RCLG/Alb-Cre and Alb-Cre mice at 0, 24, 36, 48, and 72 h after PHx. (H) Quantitative analysis of hepatocyte proliferation based on Ki-67 immunostaining at 0, 24, 36, 48, and 72 h after PHx in RCLG/Alb-Cre (n = 3–5) and Alb-Cre (n = 3–5) mice.
In vivo hepatocyte proliferation is increased in RCLG/Alb-Cre mice after 2/3 PHx
Next, we performed 2/3 partial hepatectomy (2/3 PHx) in RCLG/Alb-Cre mice and littermate controls to determine if hepatocyte-specific CR-1 overexpression enhanced liver regeneration. The liver tissue samples were collected on days 1, 1.5, 2 and 3 after 2/3 PHx and liver regeneration status was analyzed by determining the percentage of Ki67-positive hepatocytes. Ki67 immunohistochemistry results demonstrated that hepatocyte proliferation was significantly increased in the RCLG/Alb-Cre mice compared to their littermate controls on days 1, 1.5, 2 and 3 after 2/3 PHx (Figure 3G, 3H). These data demonstrated that CR-1 overexpression in the mouse liver enhanced hepatocyte proliferation after hepatectomy.
CR-1 positively regulates in vitro HCC cell growth, migration and invasiveness
Since CR-1 was significantly up-regulated in HCC tissues (Figure 2), we performed loss-of-function and gain-of-function in vitro experiments to determine the role of CR-1 on HCC cell proliferation, migration and invasion using colony formation, transwell migration and Boyden chamber invasion assays, respectively. Western blot analysis confirmed that CR-1 was significantly overexpressed in LV-CR-1-transduced BEL-7402 and HepG2 cells compared to the corresponding controls (Figure 4A). Furthermore, western blot analysis confirmed that CR-1 protein was significantly knocked down in LV-SHCR-1-transduced BEL-7402 and HepG2 cells compared to the corresponding controls (Figure 4B). Colony formation assay results demonstrated that CR-1 overexpression significantly increased proliferation of BEL-7402 and HepG2 cells, whereas, CR-1 silencing significantly reduced proliferation of BEL-7402 and HepG2 cells (Figure 4C and Supplementary Figure 5). Transwell migration and Boyden chamber invasion assays demonstrated that CR-1 overexpression significantly increased migration and invasion of BEL-7402 and HepG2 cells, whereas, CR-1 silencing significantly reduced migration and invasion of BEL-7402 and HepG2 cells (Figure 4D, 4E and Supplementary Figure 6). These results demonstrated that CR-1 modulated in vitro HCC cell proliferation, migration and invasion.
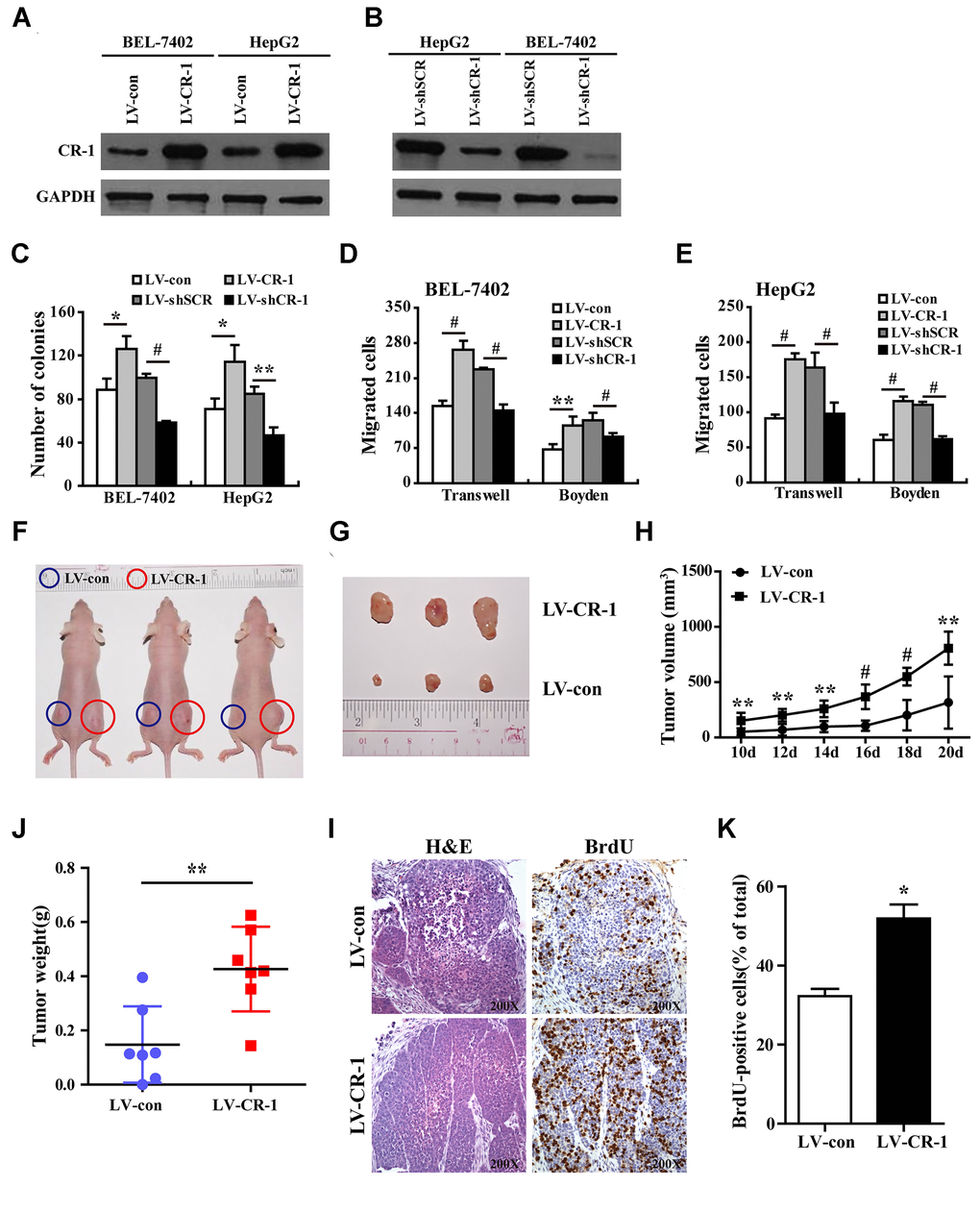
Figure 4. CR-1 overexpression promotes in vitro proliferation, migration and invasion of HCC cells and in vivo xenograft HCC growth in nude mice. (A) Western blot analysis shows CR-1 protein expression in control (LV-con) and CR-1-overexpressing (LV-CR-1) BEL-7402 and HepG2 cells. (B) Western blot analysis shows CR-1 protein expression in scrambled control (LV-shSCR) and CR-1-silenced (LV-shCR-1) BEL-7402 and HepG2 cells. (C) Colony formation assay results show proliferation ability of CR-1-overexpressing and CR-1-knockdown HCC cells with their corresponding controls. The representative images of this assay are shown in Supplementary Figure 5. (D–E) Transwell migration and Boyden invasion assay results show the migration and invasion abilities of CR-1-overexpressing, CR-1-knockdown, and control BEL-7402 (D) and HepG2 (E) cells, respectively. (F) Representative pictures of nude mice bearing subcutaneous xenografts from LV-con (blue circle) or LV-CR-1 (red circle) transduced BEL-7402 cells. (G) Representative images of subcutaneous xenograft tumors formed from LV-con or LV-CR-1 transduced BEL-7402 cells. (H) Growth curve of xenograft tumor volumes derived from LV-con or LV-CR-1 transduced BEL-7402 cells. (I) The weights of xenograft tumors derived from LV-con or LV-CR-1 transduced BEL-7402 cells. (J) Representative images of H&E-stained and BrdU-stained sections of xenograft tumors derived from LV-con or LV-CR-1 transduced BEL-7402 cells. (K) The percentages of BrdU-positive cancer cells in xenograft tumors derived from LV-con or LV-CR-1 transduced BEL-7402 cells, as calculated by total number of BrdU-positive cells relative to total number of cancer cells.
CR-1 overexpression promotes in vivo growth of HCC cell-derived xenograft tumors
To determine the in vivo role of CR-1 in HCC, we performed xenograft experiments in nude mice with subcutaneous injections of control and CR-1 overexpressing BEL-7402 cells. The subcutaneous xenograft tumors derived from CR-1 overexpressing BEL-7402 cells were significantly larger compared to those derived from control vector-transduced BEL-7402 cells (Figure 4F). The mice injected with CR-1 overexpressing BEL-7402 cells showed significantly larger tumor volumes (Figure 4H), tumor sizes (Figure 4G), and tumor weights (Figure 4I) compared to those injected with control vector-transduced BEL-7402 cells. Immunohistochemical analysis demonstrated that the percentages of BrdU-positive cells were significantly higher in tumors derived from CR-1 overexpressing BEL-7402 cells compared to those derived from control vector-transduced BEL-7402 cells (Figure 4J, 4K). Taken together, these data demonstrated that CR-1 overexpression promoted in vivo tumorigenicity of HCC cells.
Molecular signaling pathways are significantly altered in the livers of RCLG/Alb-Cre mice
We then comprehensively analyzed the status of several molecular signaling pathways in CR-1-overexpressing and CR-1-silenced HCC cells as well as control and CR-1-overexpressing mouse hepatocytes. CR-1-overexpressing BEL-7402 and HepG2 cells showed significantly higher levels of p-AKT, p-GSK-3β, p-JNK and p-Stat3 compared to the corresponding controls (Figure 5A). This suggested that AKT, JNK and Stat3 signaling pathways were activated in CR-1-overexpressing HCC cells. Conversely, p-AKT, p-GSK-3β, p-Stat3 and p-JNK levels were significantly downregulated in CR-1-silenced BEL-7402 and HepG2 cells compared to their corresponding controls (Figure 5A). We also observed elevated levels of p-AKT, p-GSK-3β, p-Stat3, p-ERK and p-JNK in the liver tissues of RCLG/Alb-Cre transgenic mice compared to those from littermate controls (Figure 5B). β-catenin levels were significantly increased in the CR-1-overexpressing HCC cells (Figure 5A) and the liver tissues of RCLG/Alb-Cre mice (Figure 5B), but were significantly reduced in CR-1-silenced HCC cells (Figure 5A). qRT-PCR assay results demonstrated that relative levels of IL-6, IL-1, Notch1, and TGF-β1 mRNAs were significantly higher in the liver tissues of RCLG/Alb-Cre mice compared to the control mice (Figure 5C). Taken together, our results demonstrated that CR-1 overexpression in the mouse liver tissues and HCC cells significantly activated AKT, Stat3, ERK, and JNK pathways, which are closely associated with hepatocyte proliferation, liver regeneration and hepatocellular carcinogenesis.
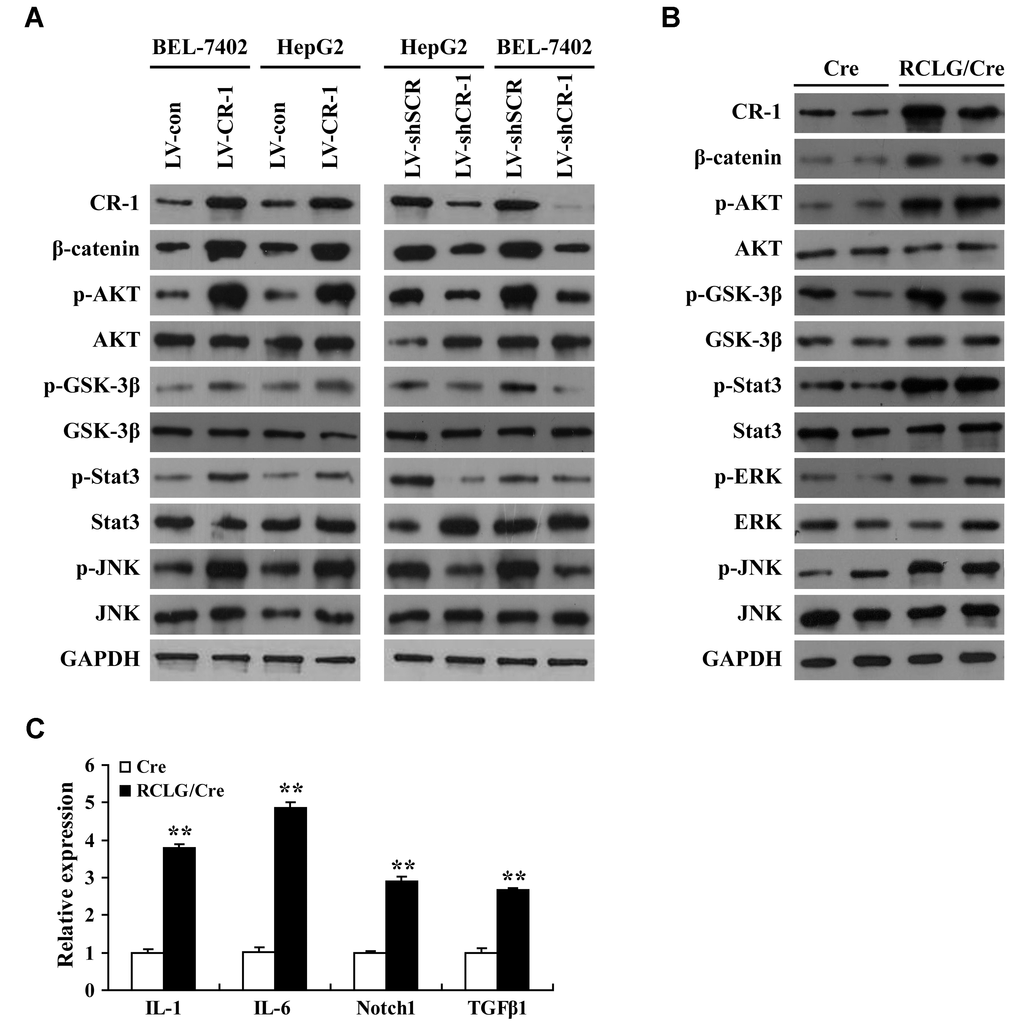
Figure 5. Liver-specific overexpression of CR-1 gene in transgenic mice activates HCC-related signaling pathways. (A) Representative western blot images show expression levels of the indicated proteins in CR-1-overexpressing, CR-1-knockdown, and corresponding control BEL-7402 and HepG2 cells. (B) Representative immunoblotting images show levels of the indicated proteins in the liver tissues of Alb-Cre and RCLG/Alb-Cre mice. (C) QRT-PCR analysis shows relative mRNA levels of IL-1, IL-6, Notch1, and TGF-β1 in the liver tissues of Alb-Cre and RCLG/Alb-Cre mice.
Discussion
Previous studies showed expression of both full-length (2-kb; FL-CR-1) and truncated (1.7-kb; SF-CR1) mRNA from the human CR-1 gene in human tissues and cells [30, 31]. SF-CR-1 mRNA was only present in tissues such as the pancreas, heart, stomach, mammary gland, liver and placenta in humans [30]. Our results demonstrated that both the longer 2-kb and the shorter 1.7-kb CR-1 transcripts were expressed in the human lung, kidney, brain, testis, skeletal muscle, ovary and spleen. Baldassarre et al. showed that the shorter CR-1 transcript was predominant in majority of primary human colorectal carcinomas (7 out of 9 analyzed) and liver metastases tissues from primary colon tumors (13 out of 14 analyzed) [30]. Our results also demonstrated that the truncated 1.7-kb CR-1 transcript was predominantly expressed in the adjacent non-tumor liver and primary HCC tissues (Figure 1). Previous studies showed that NT2/D1 cells and four different human embryonal carcinoma cell lines expressed only the FL-CR-1 transcript, whereas, human colon carcinoma and hepatocarcinoma cell lines (SW480, SW620, LS174T, GEO, CBS, HepG2 and Hep3B) expressed only the truncated form of CR-1 mRNA [30, 31]. In this study, we demonstrated that both FL-CR-1 and SF-CR-1 transcripts were expressed in QGY7701, HepG2, Hep3B, SNU-182 and SNU-387 cells. However, SF-CR-1 transcript was predominantly expressed in these cells.
Baldassarre et al. reported that the full-length and short-length CR-1 mRNAs in the NT2/D1, GEO and HepG2 cell lines were not due to alterations in the genomic DNA [30]. Hamada et al. demonstrated that the expression of the SF-CR-1 transcript in HepG2 and SW620 cells was modulated by the canonical Wnt/β-catenin/TCF signaling pathway through a intronic-exonic enhancer element with three tandem TCF/LEF binding sites in the CR-1 gene; moreover, SF-CR-1 mRNA was predominantly expressed in Wnt-active cell lines [31].
CR-1 is a multifunctional gene that plays an essential role in embryogenesis as well as cancer growth and progression [11–16]. CR-1 is highly expressed in various human cancers [13]. The truncated form of CR-1 mRNA lacks exons 1 and 2, but the putative open reading frame still contains EGF-like module, cysteine-rich domain and the carboxy-terminal linkage sequence [30, 31]. The truncated form of CR-1 is implicated in human colon carcinomas and hepatic metastases of colon carcinoma, but its function in HCC has not yet been elucidated. Since the short form of CR-1 is the major transcript in HCC and colon cancer cells [30, 31], we hypothesize that it may be involved in cancer progression. However, this hypothesis needs to be investigated in greater detail.
Embryonic genes such as Oct-4 [17–19] and CR-1 [15, 20, 21] are either undetected or expressed at very low levels in normal adult tissues [11–16]. However, CR-1 is overexpressed in several solid tumors [11–16]. Aberrant activation of embryonic genes such as Oct-4 [17–19] and CR-1 [15, 20, 21] is associated with oncogenesis. For example, overexpression of Cripto-1 transgene in the mammary glands of MMTV-CR-1 or WAP-CR-1 transgenic mice promoted mammary epithelial hyperplasia and adenocarcinoma [15, 20] and leiomyosarcoma of the uterus in the MMTV-CR-1 transgenic mice [21]. As reported previously [25–27], our study clearly showed that CR-1 overexpression promoted HCC progression. We demonstrated that hepatocyte-specific overexpression of transgenic CR-1 aberrantly activated several HCC-regulatory signaling pathways. Moreover, we observed deregulated expression of several genes that are key modulators of cellular growth, inflammation, malignant transformation, hepatocellular carcinogenesis, cancer progression, and poor prognosis. However, we did not observe any histological signs of hepatocyte dysplasia or HCC formation in the liver tissues from 3-, 6- or 8-month-old RCLG/Alb-Cre mice. This indicated that constitutive expression of CR-1 transgene alone was not sufficient to promote hepatocarcinogenesis. Our study demonstrated that CR-1 overexpression in the mouse liver and HCC cells significantly activated AKT, Stat3, ERK and JNK pathways, which are closely associated with hepatocyte proliferation, liver regeneration, and hepatocellular carcinogenesis. This suggested that CR-1 overexpression in the mouse liver partly contributed to hepatocyte proliferation and hepatocellular carcinogenesis by modulating the activation of these critical signaling pathways.
CR-1 regulates signaling pathways such as Wnt/β-catenin, AKT, Stat3, MAPK/ERK, TGF-β and Notch [12–15, 22, 25, 55, 56], all of which are implicated in hepatocarcinogenesis, HCC progression, and poor prognosis [9, 10, 57–59]. High levels of phosphorylated AKT and β-catenin were reported in mammary gland tumor tissues from MMTV-CR-1 or WAP-CR-1 transgenic mice [15, 20] and leiomyosarcoma tissues of the uterus in the MMTV-CR-1 transgenic mice [21]. In the present study, CR-1 overexpression in RCLG/Alb-Cre transgenic mouse liver and HCC cells significantly activated Wnt/β-catenin, AKT, MAPK/ERK, Stat3, TGF-β, JNK and Notch pathways. Therefore, our results suggested that hepatocyte-specific overexpression of CR-1 promoted aberrant activation of several signaling pathways involved in premalignant alterations during hepatocarcinogenesis.
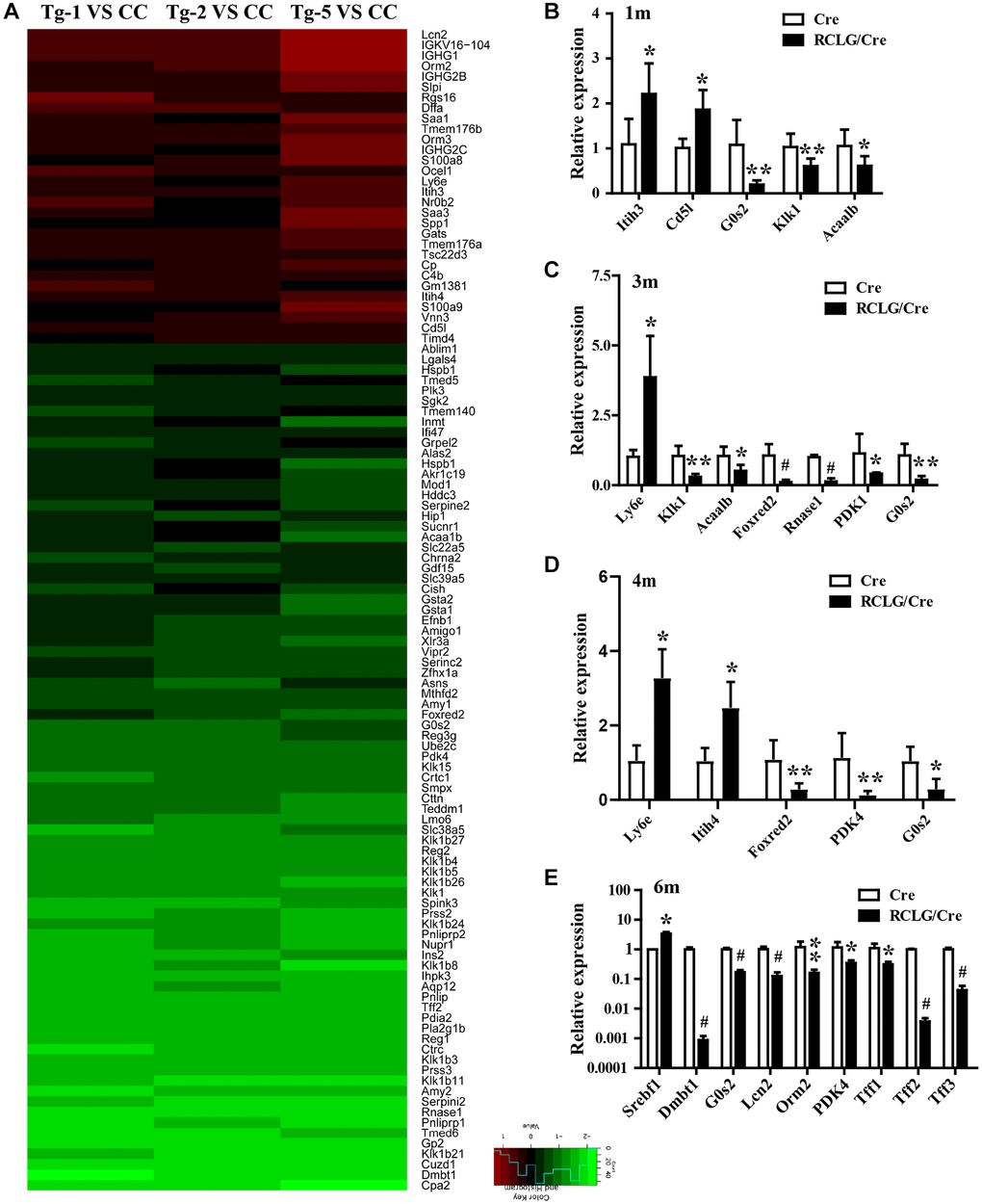
Microarray analysis showed that most of the 113 differentially expressed genes found in the liver tissues of RCLG/Alb-Cre mice were related to cellular proliferation, cell cycle, apoptosis, liver regeneration, DNA damage, stress responses, inflammation response, immune escape, defense response, acute-phase response, cellular malignant transformation, oncogenesis or cancer malignant progression, and poor prognosis. These processes are implicated in various liver pathologies including HCC [60–63].
Several genes that were differentially up-regulated (Orm2, Orm3, S100A8 and S100A9) or down-regulated (i.e., DMBT1, Gsta2 and G0S2) in the liver tissues of RCLG/Alb-Cre transgenic mice have been previously associated with hepatocarcinogenesis [37, 38, 64–66]. Transgenic mice with hepatocyte-specific overexpression of DNA binding protein A (dbpA) spontaneously developed HCC at about 1.5 years, but morphological changes in the liver were not observed at 30–40 weeks [37]. Moreover, 31–32-week-old male dbpA transgenic mouse livers showed upregulation of Orm2 (Orosomucoid 2) and Orm3 (Orosomucoid 3), as well as downregulation of G0S2 (G0/G1 switch gene 2) [37]. This was in accordance with our findings in HCC tissues (Figure 7). Gsta2 (glutathione-S-transferase, alpha type 2) mRNA was significantly downregulated in the liver tissues from fenofibrate-induced hepatocarcinogenesis model rats [66]. Deleted in malignant brain tumors 1 (DMBT1) gene was downregulated in RCLG/Alb-Cre transgenic livers (Figure 6 and Supplementary Table 2) and HCC tissues [67]. Deng et al. demonstrated that DMBT1 knockdown enhanced proliferation and malignant transformation of hepatic progenitor cells (HPCs) [38], thereby suggesting an important role for DMB1 in hepatocarcinogenesis. The damage-associated molecular pattern proteins, S100A8 and S100A9, enhanced growth and invasiveness of various cancer cell lines [68] and are implicated in liver, brain, breast, colon, thymus and thyroid cancers [64]. The upregulation of S100A8 and S100A9 correlates with poor differentiation of human liver cancer cells and tissues [64, 65, 69]. The expression levels of S100A8 and S100A9 were elevated in serum and tumor tissue samples from HCC patients [65, 69]. Moreover, higher S100A9 expression was reported in blood serum and tissue samples from HBV-positive patients with HCC compared to those from HBV-negative patients with HCC [69]. Overexpression of S100A8/A9 increased in vitro HCC cell survival, growth and invasiveness as well as xenograft tumor size in the mouse model [64, 69]. In the diethylnitrosamine-induced HCC model, S100A8/A9 ablation impaired liver cancer progression by decreasing cancer cell proliferation [64]. S100A8 and S100A9 are NF-μB target genes that synergistically enhance reactive oxygen species (ROS) and survival of HCC cells [64]. Previous studies show that S100A8 and S100A9 proteins form a heterodimer called calprotectin that promoted HCC development and progression [64, 65]. We postulate that increased expression of S100A9 and S100A8 in the liver tissues of RCLG/Alb-Cre transgenic mice promoted hepatocyte transformation and survival against injury.
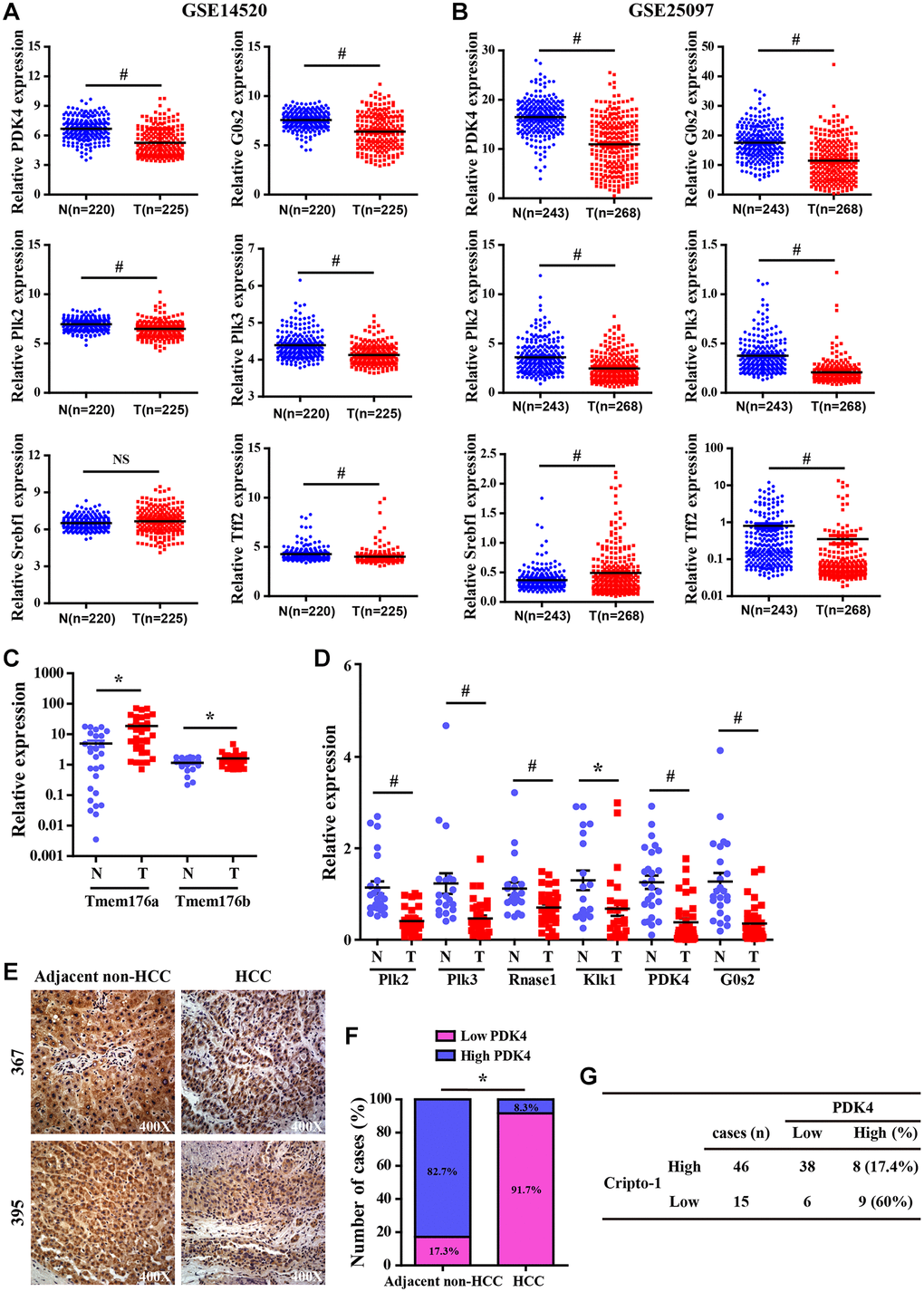
Pyruvate dehydrogenase complex (PDC) is a key modulator of tricarboxylic acid (TCA) cycle flux in the mitochondria by catalyzing the oxidation of pyruvate into acetyl CoA and NADH, which is required for the TCA cycle and mitochondrial respiration; phosphorylation of PDC is catalyzed in humans by four isozymes of pyruvate dehydrogenase kinase (PDK1, PDK2, PDK3 and PDK4) with about 70% homology [70–75]. PDKs 1–3 are closely associated with metabolism of cancer cells because they inactivate PDC via phosphorylation [73]. PDK4 acts as a potential tumor suppressor because its expression is significantly reduced in lung, ovarian, colon and breast cancers [76–78]. In this study, we demonstrated significantly decreased expression of PDK4 in the RCLG/Alb-Cre transgenic livers (Figure 6 and Supplementary Table 2) and human HCC tissue samples (Figure 7). Moreover, PDK4 silencing significantly increased proliferation, tumorigenicity and invasion of HCC cells (unpublished data). Furthermore, PDK4-/- livers showed enhanced hepatocyte growth, but these effects were inhibited by arsenic treatment [45]. Overall, our data suggested that PDK4 was a potential suppressor of hepatocarcinogenesis in mice and humans.
Several studies have shown that oxidative stress promotes hepatocarcinogenesis [79–81]. In this study, two genes encoding antioxidant proteins, glutathione S-transferase, alpha 1 (Gsta1) and glutathione S-transferase, alpha 2 (Gsta2), were down-regulated and a pro-oxidant protein, vanin 3 (Vnn3), was up-regulated in the RCLG/Alb-Cre transgenic livers (Figure 6 and Supplementary Table 2). Vanin/pantetheinase is highly expressed in several organs such as intestine, liver and kidney; moreover, vanin 1 (Vnn1), vanin 2 (Vnn2) and vanin 3 (Vnn3) induce oxidative stress [82–85]. Down-regulation of oxidant defense genes such as Gsta1 and Gsta2 in the liver was associated with fenofibrate-induced hepatocarcinogenesis in rats [66]. Therefore, we postulate that CR-1 overexpression in the hepatocytes of RCLG/Alb-Cre mice may induce oxidative stress by increasing the expression of Vnn3 and reducing the expression of antioxidant genes, Gsta1 and Gsta2, in RCLG/Alb-Cre liver tissues, thereby enhancing their survival and favoring the onset of hepatocarcinogenesis.
DNA damage plays an integral role in hepatocarcinogenesis [86, 87]. In this study, we observed significant down-regulation in the expression levels of DNA damage response factors such as polo-like kinase 2 (Plk2) and polo-like kinase 3 (Plk3) in the RCLG/Alb-Cre transgenic mouse livers (Figure 6 and Supplementary Table 2) and human HCC tissue samples (Figure 7). Plk2 and Plk3 act as tumor suppressors through their functions in the p53 signaling network and guard cells against various stress signals [88–92]. Plk3 promotes DNA damage-induced cell cycle arrest via the ATM/p53 pathway and Plk3-deficient mice develop tumors [88–92]. Plk2 and Plk3 are involved in checkpoint-mediated cell cycle arrest to protect against accumulation of genetic defects [88–92]. Therefore, the results of our study imply that decreased expression of Plk2 and Plk3 in RCLG/Alb-Cre transgenic liver tissues may promote liver carcinogenesis through increased oxidative stress-related damage and accumulation of genetic mutations in the hepatocytes and their subsequent proliferation.
Chronic liver inflammation promotes HCC because of increased oxidative/nitrosative stress and lipid peroxidation, which oxidatively mutates the genomic DNA [80, 93–96]. In this study, we documented up-regulation of several genes involved in liver inflammation (CD5L, S100A8, S100A9, Timd4 and Rgs16), immune escape (Orm2, Orm3, IGHG1, IGHG2B, IGHG2C and IGKV16-104), acute-phase response (Orm2, Orm3, Saa1, Saa3, Itih3 and Itih4), and immune defense response (Ly6e) (Figure 6 and Supplementary Table 2) in the RCLG/Alb-Cre transgenic livers.
Guerra et al. showed that Api6/AIM/Spa/CD5L was upregulated in HCC, and promoted HCC cell proliferation and survival by binding to HSPA5 (GRP78) [50]. Barcena et al. demonstrated that CD5L regulated liver damage, fibrosis and immune cell content [49]. CD5L overexpression in the alveolar type II epithelial cells (AT II cells) of transgenic mice induced malignant transformation and spontaneous lung adenocarcinoma by inhibiting apoptosis of lung epithelial cells and promoting immune escape [97]. Moreover, CD5L overexpression in AT II cells increased the levels of pro-inflammatory cytokines/chemokines in the bronchoalveolar lavage fluid and serum, promoted expansion of myeloid-derived suppressor cells (MDSC) in lungs and blood; lung MDSCs suppressed in vitro T-cell proliferation and activity and reduced in vivo levels of T cells in vivo following doxycycline-induced CD5L transgene activation [97]. IGHG1 promotes pancreatic cancer cell proliferation and immune evasion [98].
Tumor immune-evasion refers to the ability of cancer cells to circumvent host immune systems and utilize inflammatory factors for tumor growth and progression [99]. T-cell immunoglobulin mucin (TIM) gene family members maintain immune homeostasis by regulating multiple phases of the immune response [99]. Cancer cells evade immunosurveillance via TIM gene family members, which also inhibit inflammation-related tumor progression [99]. Timd4 (T-cell immunoglobulin domain and mucin domain 4; also known as TIM4) plays a critical role in regulating tumor immunosurveillance and anti-tumor immunity [99]. Timd4 overexpression is also associated with increased lung cancer cell proliferation [100].
Acute-phase response (APR) proteins such as Orm2 and Orm3 play an important role in anti-inflammatory and immunomodulatory responses that are initiated against infections, physical trauma, or malignancies [36]. Orm2 is frequently down-regulated in HCC tissues and negatively correlates with tumor progression and intrahepatic metastasis [36]. The serum levels of another APR protein, ITIH4, are elevated in HCC patients during acute phase; moreover, ITIH4 is up-regulated by interleukin-6 in HepG2 cells [101]. Collectively, our findings suggest that CR-1 regulates several inflammatory and host immunity factors that are involved in HCC growth and progression.
In conclusion, our study demonstrated that SF-CR-1 mRNA was predominantly expressed in most HCC tissues and cell lines. However, the biological function of SF-CR-1 mRNA in HCC progression requires further investigation. We also demonstrated that hepatocyte-specific overexpression of CR-1 in transgenic mice deregulated several signaling pathways and genes involved in hepatocarcinogenesis. However, hepatocyte-specific CR-1 overexpression alone was not sufficient to initiate hepatocarcinogenesis in mice. Therefore, the exact role of CR-1 in HCC remains to be further explored, plausibly using the carcinogen (DEN: diethylnitrosamine)-induced HCC model.
Materials and Methods
Cell lines and cell culture
HEK293T cells were purchased from the American Type Culture Collection (ATCC). The human HCC cell lines (HepG2 and BEL-7402) were obtained from the Cell Bank of the Chinese Academy of Sciences (Shanghai, China). These cell lines were cultured in Dulbecco’s modified Eagle’s medium (DMEM) (high glucose) supplemented with 10% fetal bovine serum (FBS) in a humidified incubator maintained at 5% CO2 and 37°C.
Clinical specimens
Fresh primary HCC specimens and tumor-adjacent noncancerous tissues were collected from HCC patients that underwent surgery at the Sun Yat-sen Memorial Hospital, Sun Yat-sen University (Guangzhou, China). We obtained informed consent from all patients. The inclusion criteria were: (1) a definite pathological diagnosis of HCC and (2) no other anti-cancer treatment before surgery. We extracted total RNA and protein from the fresh-frozen specimens for qRT-PCR and western blot, respectively. We then prepared formalin-fixed paraffin-embedded blocks from the fresh-frozen HCC and adjacent non-tumor liver specimens. This study was approved by the Medical Ethics Committee of Southern Medical University.
RT-PCR and quantitative real-time PCR (qRT-PCR)
We isolated total RNA and performed reverse transcription, RT-PCR and qRT-PCR using protocols as previously described [3, 16, 102–105]. For RT-PCR, we amplified two distinct regions of the CR-1 mRNA based on the previously published CR-1 cDNA sequence [30, 31] using CR-1-specific primers. The CR-1 specific primers were: UN-A forward primer (nucleotides 375–398): 5′-ACCTGGCCTTCAGAGATGACAGCA-3′, UN-B reverse primer (nucleotides 656–680): 5′-ATGCCTGAGGAAAGCAGCGGAGCT-3′ and UN-D forward primer (nucleotides 248–266): 5′-AAAGCTATGGACTGCAGGA-3′ (Figure 1A). UN-A/UN-B and UN-D/UN-B primer sets yielded 305 bp and 432 bp sized PCR products, respectively (Figure 1A). GAPDH was used as internal control. The primer pairs for amplification of GAPDH are listed in Supplementary Table 3. The primer pairs used in qRT-PCR are listed in Supplementary Tables 4 and 5. GAPDH was used as internal control. All samples were normalized to internal controls. The relative fold changes in specific mRNA levels were calculated using the 2-ΔΔCt method.
Western blotting
Western blot analysis was performed as previously described [3, 16, 102–105]. GAPDH or β-actin were used as loading controls. The primary antibodies used in this study are listed in Supplementary Table 6.
Histological analysis and immunohistochemistry (IHC)
Histological analysis and immunohistochemical staining were performed as previously described [3, 16, 106–109]. The antibodies used in the study and the experimental conditions are summarized in the Supplementary Table 6.
Lentivirus production and transduction
The human CR-1 lentiviral shRNA vector (pLV-shCR-1), empty lentiviral vector expressing scrambled shRNA (pLV-shSCR), lentiviral CR-1 expression vector (pLV-CR-1), and empty lentiviral vector (pLV-con) were generously provided by Prof. Peter C. Gray (The Salk Institute for Biological Studies, USA) [110]. The lentiviral packaging plasmids, psPAX2 and pMD2.G, were kindly provided by Dr. Didier Trono (University of Geneva, Geneva, Switzerland). We infected BEL-7402 and HepG2 cells with pLV-con, pLV-CR-1, pLV-shSCR, and pLV-shCR-1, respectively, in combination with lentiviral packaging plasmids (psPAX2 and pMD2.G) to generate stable cell lines, and then isolated recombinant lentiviruses, namely LV-con, LV-CR-1, LV-shSCR and LV-shCR-1 as previously described [105, 111].
Transwell migration and Boyden invasion assays
Transwell migration and Boyden invasion assays were performed as described previously [3, 104, 109].
Tumor xenografts in nude mice
Three- to four-week old female BALB/c nude mice were obtained from the Medical Laboratory Animal Center of Guangdong Province. We subcutaneously injected CR-1-overexpressing or vector-expressing BEL-7402 cells (1 × 106 cells) into the right or left dorsal thigh of mice (n = 7), respectively. The tumor size parameters were measured every 2 days using a caliper slide rule and tumor volumes were calculated using the following formula: Tumor volume = (D × d2)/2, where ‘D’ is the longest diameter and ‘d’ is the shortest diameter. The mice were sacrificed after 20 days. The subcutaneous tumors were harvested, weighed, fixed overnight in 4% paraformaldehyde, dehydrated, paraffin-embedded, and sectioned. The animal experiments were carried out strictly according to the recommendations in the Guide for the Care and Use of Laboratory Animals of the Southern Medical University. The protocols for animal experiments were approved by the Committee on Ethics of Animal Experiments of the Southern Medical University. All surgeries were performed under sodium pentobarbital anesthesia. All efforts were made to minimize the suffering of animals.
Generation of RCLG transgenic mice
The pCI-CR-1 vector [112] was generously provided by Dr. David S. Salomon (Center for Cancer Research, National Cancer Institute, USA). The pCAG-RLG vector [113, 114] was generously provided by Prof. Manuela Martins-Green (University of California, USA). We PCR amplified a 600 bp fragment of CR-1 cDNA from the pCI-CR-1 as previously described [112] and cloned it into the Sma I site of the pCAG-RLG plasmid [113, 114]. The RCLG transgenic construct was confirmed by PCR, DNA sequencing and enzyme digestion analysis (data not shown) and designated as pCAG-RCLG (Supplementary Figure 1A).
RCLG transgenic mice were generated by microinjecting DNA into the pronuclei of fertilized embryos as previously described [115]. Three days after birth, the off-springs were screened to identify potential RCLG transgenic founders using mRFP assay with the IVIS Lumina II imaging system (Xenogen Corp., Alameda, CA, USA) and subsequently confirmed by PCR-based genotyping.
Whole-animal (in vivo) and organ (ex vivo) fluorescence imaging
Fluorescence imaging of mRFP (monomeric red fluorescent protein) in the whole-animal and organs was performed using stereo fluorescence microscope (Nikon, AZ100) or the Xenogen IVIS Lumina II Imaging System according to the protocols described previously [16, 102, 104, 116–118].
In vivo and ex vivo optical imaging of firefly luciferase (Luc) activity
We purchased homozygous Alb-Cre mice (B6. Cg-Tg (Alb-cre) 21Mgn/J) from the Model Animal Research Center of Nanjing University and crossed them with the RCLG mice to obtain RCLG/Alb-Cre double transgenic mice. Firefly luciferase (Luc) expression was observed specifically in the liver of the RCLG/Alb-Cre double transgenic mice using the non-invasive in vivo bioluminescence imaging and quantified using the IVIS Lumina II imaging system. The protocols for bioluminescence imaging to detect Luc activity in the whole-animal (in vivo) and the dissected organs (ex vivo) using the IVIS system were as previously described [3, 16, 102, 104, 116–118].
PCR genotyping
We performed PCR analysis using the tail genomic DNA from the mice to further confirm the RCLG/Alb-Cre double transgenic mice. The specific forward primer (FP) and reverse primer (RP) sequences for the Luc and Cre genes are shown in Supplementary Table 7.
Microarray analysis
We used 32K mouse genome microarray (Beijing Capital biology company, China) to perform mRNA expression microarray analysis as previously described [28]. Total RNA was extracted from the livers of control and RCLG/Alb-Cre transgenic mice. The sequence hybridization and data analysis were performed by Capital Bio Corp. (Bejing, China).
Statistical analysis
The data are presented as means ± SEM. Statistical analysis was performed using the SPSS 13.0 software package and Graphpad 5.0 software. Two-tailed Student’s t test was used to compare data between two independent groups. Statistical significance was set at *P < 0.05, **P < 0.01 and #P < 0.001.
Author Contributions
Xiao-Lin Lin, Dong Xiao and Wen-Tao Zhao conceived the project idea, designed the experiments, and wrote the paper; Yu Liu, Yan-Qing Li, Shi-Hao Huang, Yong-Long Li, Jun-Shuang Jia Guan-Qi Dai, Yu-Cai Wang, Fang Wei, and Xiao-Yan Li performed experiments and analyzed the data; Jia-Wei Xia performed few experiments and performed statistical analyses of the data; Liu-Xin Han, Xiao-Ling Zhang and Xu-Dong Xiang helped with the analyses of results and revised the figures; Jia-Hong Wang assisted with the analyses of results and helped revise the manuscript. All authors approved the final manuscript.
Acknowledgments and Funding
We thank Prof. Peter C. Gray (The Salk Institute for Biological Studies, USA), Prof. David S. Salomon (Center for Cancer Research, National Cancer Institute, USA) and Prof. Manuela Martins-Green (University of California, USA) for generously providing plasmids.
Conflicts of Interest
The authors declare no conflicts of interest related to this study.
Funding
This work was financially supported by the National Natural Science Foundation of China (Grant Nos. 81872209, 81672689, 81372896 and 81172587 to D. Xiao; Grant Nos. 81600488 and 81870602 to X.L. Lin; Grant No. 81702778 to J.S. Jia and Grant No. 82060425 to W.T. Zhao), the Natural Science Foundation of Guangdong Province of China (Grant No. 2014A030313294 to D. Xiao), the Science and Technology Planning Project of Guangdong Province of China (Grant Nos. 2009B060300008, 2013B060300013, and 2017A010105017 to D. Xiao; Grant No. 2017A030303018 to J.S. Jia and Grant No. 2015A030302024, to X.L. Lin), the China Postdoctoral Science Foundation (Grant Nos. 2015M572338, 2016T90792, 2017M622740 and 2018T110884 to X.L. Lin and Grant No. 2020M673595XB, to W.T. Zhao), the Medical Scientific Research Foundation of Guangdong Province of China (Grant No. A2017420 to J.S. Jia), the Basic Research Foundation of Yunnan Province Local Universities (Grant No. 202001BA070001-063 to J.W. Xia and Grant No. 202001BA070001-043 to L.X. Han), Guangxi Natural Science Foundation Program (Grant No. 2020GXNSFAA159131 to X.L. Zhang), The Applied Basic Research Foundation of Yunnan Province Science and Technology Department and Kunming Medical University of China (Grant No. 2018FE001(-249) to W.T. Zhao and Grant No. 2018FE001(-243) to X.D. Xiang) and the Science and Technology Planning Project of Kunming City of China (Grant No. 2019-1-S-25318000001329 to J.W. Xia).
References
- 1. Akinyemiju T, Abera S, Ahmed M, Alam N, Alemayohu MA, Allen C, Al-Raddadi R, Alvis-Guzman N, Amoako Y, Artaman A, Ayele TA, Barac A, Bensenor I, et al, and Global Burden of Disease Liver Cancer Collaboration. The Burden of Primary Liver Cancer and Underlying Etiologies From 1990 to 2015 at the Global, Regional, and National Level: Results From the Global Burden of Disease Study 2015. JAMA Oncol. 2017; 3:1683–91. https://doi.org/10.1001/jamaoncol.2017.3055 [PubMed]
- 2. Erridge S, Pucher PH, Markar SR, Malietzis G, Athanasiou T, Darzi A, Sodergren MH, Jiao LR. Meta-analysis of determinants of survival following treatment of recurrent hepatocellular carcinoma. Br J Surg. 2017; 104:1433–42. https://doi.org/10.1002/bjs.10597 [PubMed]
- 3. Zhao WT, Lin XL, Liu Y, Han LX, Li J, Lin TY, Shi JW, Wang SC, Lian M, Chen HW, Sun Y, Xu K, Jia JS, et al. miR-26a promotes hepatocellular carcinoma invasion and metastasis by inhibiting PTEN and inhibits cell growth by repressing EZH2. Lab Invest. 2019; 99:1484–500. https://doi.org/10.1038/s41374-019-0270-5 [PubMed]
- 4. Anstee QM, Reeves HL, Kotsiliti E, Govaere O, Heikenwalder M. From NASH to HCC: current concepts and future challenges. Nat Rev Gastroenterol Hepatol. 2019; 16:411–28. https://doi.org/10.1038/s41575-019-0145-7 [PubMed]
- 5. Calderaro J, Ziol M, Paradis V, Zucman-Rossi J. Molecular and histological correlations in liver cancer. J Hepatol. 2019; 71:616–30. https://doi.org/10.1016/j.jhep.2019.06.001 [PubMed]
- 6. Yang JD, Hainaut P, Gores GJ, Amadou A, Plymoth A, Roberts LR. A global view of hepatocellular carcinoma: trends, risk, prevention and management. Nat Rev Gastroenterol Hepatol. 2019; 16:589–604. https://doi.org/10.1038/s41575-019-0186-y [PubMed]
- 7. Sonntag R, Giebeler N, Nevzorova YA, Bangen JM, Fahrenkamp D, Lambertz D, Haas U, Hu W, Gassler N, Cubero FJ, Müller-Newen G, Abdallah AT, Weiskirchen R, et al. Cyclin E1 and cyclin-dependent kinase 2 are critical for initiation, but not for progression of hepatocellular carcinoma. Proc Natl Acad Sci U S A. 2018; 115:9282–87. https://doi.org/10.1073/pnas.1807155115 [PubMed]
- 8. Tomas-Loba A, Manieri E, Gonzalez-Teran B, Mora A, Leiva-Vega L, Santamans AM, Romero-Becerra R, Rodriguez E, Pintor-Chocano A, Feixas F, Lopez JA, Caballero B, Trakala M, et al. p38γ is essential for cell cycle progression and liver tumorigenesis. Nature. 2019; 568:557–60. https://doi.org/10.1038/s41586-019-1112-8 [PubMed]
- 9. Wang W, Smits R, Hao H, He C. Wnt/β-Catenin Signaling in Liver Cancers. Cancers (Basel). 2019; 11:926. https://doi.org/10.3390/cancers11070926 [PubMed]
- 10. Ye H, Zhang C, Wang BJ, Tan XH, Zhang WP, Teng Y, Yang X. Synergistic function of Kras mutation and HBx in initiation and progression of hepatocellular carcinoma in mice. Oncogene. 2014; 33:5133–38. https://doi.org/10.1038/onc.2013.468 [PubMed]
- 11. de Castro NP, Rangel MC, Nagaoka T, Salomon DS, Bianco C. Cripto-1: an embryonic gene that promotes tumorigenesis. Future Oncol. 2010; 6:1127–42. https://doi.org/10.2217/fon.10.68 [PubMed]
- 12. Klauzinska M, Castro NP, Rangel MC, Spike BT, Gray PC, Bertolette D, Cuttitta F, Salomon D. The multifaceted role of the embryonic gene Cripto-1 in cancer, stem cells and epithelial-mesenchymal transition. Semin Cancer Biol. 2014; 29:51–58. https://doi.org/10.1016/j.semcancer.2014.08.003 [PubMed]
- 13. Rangel MC, Karasawa H, Castro NP, Nagaoka T, Salomon DS, Bianco C. Role of Cripto-1 during epithelial-to-mesenchymal transition in development and cancer. Am J Pathol. 2012; 180:2188–200. https://doi.org/10.1016/j.ajpath.2012.02.031 [PubMed]
- 14. Nagaoka T, Karasawa H, Castro NP, Rangel MC, Salomon DS, Bianco C. An evolving web of signaling networks regulated by Cripto-1. Growth Factors. 2012; 30:13–21. https://doi.org/10.3109/08977194.2011.641962 [PubMed]
- 15. Strizzi L, Bianco C, Normanno N, Salomon D. Cripto-1: a multifunctional modulator during embryogenesis and oncogenesis. Oncogene. 2005; 24:5731–41. https://doi.org/10.1038/sj.onc.1208918 [PubMed]
- 16. Lin X, Zhao W, Jia J, Lin T, Xiao G, Wang S, Lin X, Liu Y, Chen L, Qin Y, Li J, Zhang T, Hao W, et al. Ectopic expression of Cripto-1 in transgenic mouse embryos causes hemorrhages, fatal cardiac defects and embryonic lethality. Sci Rep. 2016; 6:34501. https://doi.org/10.1038/srep34501 [PubMed]
- 17. Beachy PA, Karhadkar SS, Berman DM. Tissue repair and stem cell renewal in carcinogenesis. Nature. 2004; 432:324–31. https://doi.org/10.1038/nature03100 [PubMed]
- 18. Gidekel S, Pizov G, Bergman Y, Pikarsky E. Oct-3/4 is a dose-dependent oncogenic fate determinant. Cancer Cell. 2003; 4:361–70. https://doi.org/10.1016/s1535-6108(03)00270-8 [PubMed]
- 19. Hochedlinger K, Yamada Y, Beard C, Jaenisch R. Ectopic expression of Oct-4 blocks progenitor-cell differentiation and causes dysplasia in epithelial tissues. Cell. 2005; 121:465–77. https://doi.org/10.1016/j.cell.2005.02.018 [PubMed]
- 20. Sun Y, Strizzi L, Raafat A, Hirota M, Bianco C, Feigenbaum L, Kenney N, Wechselberger C, Callahan R, Salomon DS. Overexpression of human Cripto-1 in transgenic mice delays mammary gland development and differentiation and induces mammary tumorigenesis. Am J Pathol. 2005; 167:585–97. https://doi.org/10.1016/S0002-9440(10)63000-3 [PubMed]
- 21. Strizzi L, Bianco C, Hirota M, Watanabe K, Mancino M, Hamada S, Raafat A, Lawson S, Ebert AD, D'Antonio A, Losito S, Normanno N, Salomon DS. Development of leiomyosarcoma of the uterus in MMTV-CR-1 transgenic mice. J Pathol. 2007; 211:36–44. https://doi.org/10.1002/path.2083 [PubMed]
- 22. Francescangeli F, Contavalli P, De Angelis ML, Baiocchi M, Gambara G, Pagliuca A, Fiorenzano A, Prezioso C, Boe A, Todaro M, Stassi G, Castro NP, Watanabe K, et al. Dynamic regulation of the cancer stem cell compartment by Cripto-1 in colorectal cancer. Cell Death Differ. 2015; 22:1700–13. https://doi.org/10.1038/cdd.2015.19 [PubMed]
- 23. Strizzi L, Margaryan NV, Gilgur A, Hardy KM, Normanno N, Salomon DS, Hendrix MJ. The significance of a Cripto-1 positive subpopulation of human melanoma cells exhibiting stem cell-like characteristics. Cell Cycle. 2013; 12:1450–56. https://doi.org/10.4161/cc.24601 [PubMed]
- 24. Liu Q, Cui X, Yu X, Bian BS, Qian F, Hu XG, Ji CD, Yang L, Ren Y, Cui W, Zhang X, Zhang P, Wang JM, et al. Cripto-1 acts as a functional marker of cancer stem-like cells and predicts prognosis of the patients in esophageal squamous cell carcinoma. Mol Cancer. 2017; 16:81. https://doi.org/10.1186/s12943-017-0650-7 [PubMed]
- 25. Lo RC, Leung CO, Chan KK, Ho DW, Wong CM, Lee TK, Ng IO. Cripto-1 contributes to stemness in hepatocellular carcinoma by stabilizing Dishevelled-3 and activating Wnt/β-catenin pathway. Cell Death Differ. 2018; 25:1426–41. https://doi.org/10.1038/s41418-018-0059-x [PubMed]
- 26. Wang JH, Wei W, Xu J, Guo ZX, Xiao CZ, Zhang YF, Jian PE, Wu XL, Shi M, Guo RP. Elevated expression of Cripto-1 correlates with poor prognosis in hepatocellular carcinoma. Oncotarget. 2015; 6:35116–28. https://doi.org/10.18632/oncotarget.5057 [PubMed]
- 27. Zhang Y, Xu H, Chi X, Fan Y, Shi Y, Niu J. High level of serum Cripto-1 in hepatocellular carcinoma, especially with hepatitis B virus infection. Medicine (Baltimore). 2018; 97:e11781. https://doi.org/10.1097/MD.0000000000011781 [PubMed]
- 28. Huang T, Guo YZ, Yue X, Zhang GP, Zhang Y, Kuang M, Peng BG, Li SQ. Cripto-1 promotes tumor invasion and predicts poor outcomes in hepatocellular carcinoma. Carcinogenesis. 2020; 41:571–81. https://doi.org/10.1093/carcin/bgz133 [PubMed]
- 29. Li XS, Wang JH, Yang XZ, Ma L, Shi YX, Song Y, Jiang P, Gao S, Dong Y, Lin JR, Jin C. Beneficial effects of Cripto-1 for transarterial chemoembolization in hepatocellular carcinoma. Aging (Albany NY). 2019; 11:2998–3011. https://doi.org/10.18632/aging.101951 [PubMed]
- 30. Baldassarre G, Tucci M, Lembo G, Pacifico FM, Dono R, Lago CT, Barra A, Bianco C, Viglietto G, Salomon D, Persico MG. A truncated form of teratocarcinoma-derived growth factor-1 (cripto-1) mRNA expressed in human colon carcinoma cell lines and tumors. Tumour Biol. 2001; 22:286–93. https://doi.org/10.1159/000050629 [PubMed]
- 31. Hamada S, Watanabe K, Hirota M, Bianco C, Strizzi L, Mancino M, Gonzales M, Salomon DS. beta-Catenin/TCF/LEF regulate expression of the short form human Cripto-1. Biochem Biophys Res Commun. 2007; 355:240–44. https://doi.org/10.1016/j.bbrc.2007.01.143 [PubMed]
- 32. Postic C, Shiota M, Niswender KD, Jetton TL, Chen Y, Moates JM, Shelton KD, Lindner J, Cherrington AD, Magnuson MA. Dual roles for glucokinase in glucose homeostasis as determined by liver and pancreatic beta cell-specific gene knock-outs using Cre recombinase. J Biol Chem. 1999; 274:305–15. https://doi.org/10.1074/jbc.274.1.305 [PubMed]
- 33. Okada H, Kimura MT, Tan D, Fujiwara K, Igarashi J, Makuuchi M, Hui AM, Tsurumaru M, Nagase H. Frequent trefoil factor 3 (TFF3) overexpression and promoter hypomethylation in mouse and human hepatocellular carcinomas. Int J Oncol. 2005; 26:369–77. https://doi.org/10.3892/ijo.26.2.369 [PubMed]
- 34. Sun Q, Zhang Y, Liu F, Zhao X, Yang X. Identification of candidate biomarkers for hepatocellular carcinoma through pre-cancerous expression analysis in an HBx transgenic mouse. Cancer Biol Ther. 2007; 6:1532–38. https://doi.org/10.4161/cbt.6.10.4683 [PubMed]
- 35. Ochiai Y, Yamaguchi J, Kokuryo T, Yokoyama Y, Ebata T, Nagino M. Trefoil Factor Family 1 Inhibits the Development of Hepatocellular Carcinoma by Regulating β-Catenin Activation. Hepatology. 2020; 72:503–17. https://doi.org/10.1002/hep.31039 [PubMed]
- 36. Fang T, Cui M, Sun J, Ge C, Zhao F, Zhang L, Tian H, Zhang L, Chen T, Jiang G, Xie H, Cui Y, Yao M, et al. Orosomucoid 2 inhibits tumor metastasis and is upregulated by CCAAT/enhancer binding protein β in hepatocellular carcinomas. Oncotarget. 2015; 6:16106–19. https://doi.org/10.18632/oncotarget.3867 [PubMed]
- 37. Tobita H, Kajino K, Inami K, Kano S, Yasen M, Imamura O, Kinoshita Y, Hino O. Gene expression profile of DNA binding protein A transgenic mice. Int J Oncol. 2006; 29:673–79. https://doi.org/10.3892/ijo.29.3.673 [PubMed]
- 38. Deng H, Gao YB, Wang HF, Jin XL, Xiao JC. Expression of deleted in malignant brain tumours 1 (DMBT1) relates to the proliferation and malignant transformation of hepatic progenitor cells in hepatitis B virus-related liver diseases. Histopathology. 2012; 60:249–60. https://doi.org/10.1111/j.1365-2559.2011.04082.x [PubMed]
- 39. Yamashita T, Honda M, Takatori H, Nishino R, Minato H, Takamura H, Ohta T, Kaneko S. Activation of lipogenic pathway correlates with cell proliferation and poor prognosis in hepatocellular carcinoma. J Hepatol. 2009; 50:100–10. https://doi.org/10.1016/j.jhep.2008.07.036 [PubMed]
- 40. You Z, Li B, Xu J, Chen L, Ye H. Curcumin suppress the growth of hepatocellular carcinoma via down-regulating SREBF1. Oncol Res. 2018. [Epub ahead of print]. https://doi.org/10.3727/096504018X15219173841078 [PubMed]
- 41. Si T, Ning X, Zhao H, Zhang M, Huang P, Hu Z, Yang L, Lin L. microRNA-9-5p regulates the mitochondrial function of hepatocellular carcinoma cells through suppressing PDK4. Cancer Gene Ther. 2021; 28:706–18. https://doi.org/10.1038/s41417-020-00253-w [PubMed]
- 42. Qin YJ, Lin TY, Lin XL, Liu Y, Zhao WT, Li XY, Lian M, Chen HW, Li YL, Zhang XL, Xiao D, Jia JS, Sun Y. Loss of PDK4 expression promotes proliferation, tumorigenicity, motility and invasion of hepatocellular carcinoma cells. J Cancer. 2020; 11:4397–405. https://doi.org/10.7150/jca.43459 [PubMed]
- 43. Fekir K, Dubois-Pot-Schneider H, Désert R, Daniel Y, Glaise D, Rauch C, Morel F, Fromenty B, Musso O, Cabillic F, Corlu A. Retrodifferentiation of Human Tumor Hepatocytes to Stem Cells Leads to Metabolic Reprogramming and Chemoresistance. Cancer Res. 2019; 79:1869–83. https://doi.org/10.1158/0008-5472.CAN-18-2110 [PubMed]
- 44. Yang C, Wang S, Ruan H, Li B, Cheng Z, He J, Zuo Q, Yu C, Wang H, Lv Y, Gu D, Jin G, Yao M, et al. Downregulation of PDK4 Increases Lipogenesis and Associates with Poor Prognosis in Hepatocellular Carcinoma. J Cancer. 2019; 10:918–26. https://doi.org/10.7150/jca.27226 [PubMed]
- 45. Choiniere J, Wu J, Wang L. Pyruvate Dehydrogenase Kinase 4 Deficiency Results in Expedited Cellular Proliferation through E2F1-Mediated Increase of Cyclins. Mol Pharmacol. 2017; 91:189–96. https://doi.org/10.1124/mol.116.106757 [PubMed]
- 46. Kondoh N, Wakatsuki T, Ryo A, Hada A, Aihara T, Horiuchi S, Goseki N, Matsubara O, Takenaka K, Shichita M, Tanaka K, Shuda M, Yamamoto M. Identification and characterization of genes associated with human hepatocellular carcinogenesis. Cancer Res. 1999; 59:4990–96. [PubMed]
- 47. Hsieh HC, Chen YT, Li JM, Chou TY, Chang MF, Huang SC, Tseng TL, Liu CC, Chen SF. Protein profilings in mouse liver regeneration after partial hepatectomy using iTRAQ technology. J Proteome Res. 2009; 8:1004–13. https://doi.org/10.1021/pr800696m [PubMed]
- 48. Ma D, Wang J, Liu L, Chen M, Wang Z. Acteoside as a potential therapeutic option for primary hepatocellular carcinoma: a preclinical study. BMC Cancer. 2020; 20:936. https://doi.org/10.1186/s12885-020-07447-3 [PubMed]
- 49. Barcena C, Aran G, Perea L, Sanjurjo L, Tellez E, Oncins A, Masnou H, Serra I, Garcia-Gallo M, Kremer L, Sala M, Armengol C, Sancho-Bru P, et al. CD5L is a pleiotropic player in liver fibrosis controlling damage, fibrosis and immune cell content. EBioMedicine. 2019; 43:513–24. https://doi.org/10.1016/j.ebiom.2019.04.052 [PubMed]
- 50. Aran G, Sanjurjo L, Bárcena C, Simon-Coma M, Téllez É, Vázquez-Vitali M, Garrido M, Guerra L, Díaz E, Ojanguren I, Elortza F, Planas R, Sala M, et al. CD5L is upregulated in hepatocellular carcinoma and promotes liver cancer cell proliferation and antiapoptotic responses by binding to HSPA5 (GRP78). FASEB J. 2018; 32:3878–91. https://doi.org/10.1096/fj.201700941RR [PubMed]
- 51. Sanjurjo L, Aran G, Téllez É, Amézaga N, Armengol C, López D, Prats C, Sarrias MR. CD5L Promotes M2 Macrophage Polarization through Autophagy-Mediated Upregulation of ID3. Front Immunol. 2018; 9:480. https://doi.org/10.3389/fimmu.2018.00480 [PubMed]
- 52. Nakamura N, Hatano E, Iguchi K, Sato M, Kawaguchi H, Ohtsu I, Sakurai T, Aizawa N, Iijima H, Nishiguchi S, Tomono T, Okuda Y, Wada S, et al. Elevated levels of circulating ITIH4 are associated with hepatocellular carcinoma with nonalcoholic fatty liver disease: from pig model to human study. BMC Cancer. 2019; 19:621. https://doi.org/10.1186/s12885-019-5825-8 [PubMed]
- 53. Li X, Li B, Li B, Guo T, Sun Z, Li X, Chen L, Chen W, Chen P, Mao Y, Zeng Y. ITIH4: Effective Serum Marker, Early Warning and Diagnosis, Hepatocellular Carcinoma. Pathol Oncol Res. 2018; 24:663–70. https://doi.org/10.1007/s12253-017-0285-4 [PubMed]
- 54. Lee EJ, Yang SH, Kim KJ, Cha H, Lee SJ, Kim JH, Song J, Chun KH, Seong J. Inter-alpha Inhibitor H4 as a Potential Biomarker Predicting the Treatment Outcomes in Patients with Hepatocellular Carcinoma. Cancer Res Treat. 2018; 50:646–57. https://doi.org/10.4143/crt.2016.550 [PubMed]
- 55. Kannan S, De Santis M, Lohmeyer M, Riese DJ
2nd , Smith GH, Hynes N, Seno M, Brandt R, Bianco C, Persico G, Kenney N, Normanno N, Martinez-Lacaci I, et al. Cripto enhances the tyrosine phosphorylation of Shc and activates mitogen-activated protein kinase (MAPK) in mammary epithelial cells. J Biol Chem. 1997; 272:3330–35. https://doi.org/10.1074/jbc.272.6.3330 [PubMed] - 56. Zhang JG, Zhao J, Xin Y. Significance and relationship between Cripto-1 and p-STAT3 expression in gastric cancer and precancerous lesions. World J Gastroenterol. 2010; 16:571–77. https://doi.org/10.3748/wjg.v16.i5.571 [PubMed]
- 57. Schmitz KJ, Wohlschlaeger J, Lang H, Sotiropoulos GC, Malago M, Steveling K, Reis H, Cicinnati VR, Schmid KW, Baba HA. Activation of the ERK and AKT signalling pathway predicts poor prognosis in hepatocellular carcinoma and ERK activation in cancer tissue is associated with hepatitis C virus infection. J Hepatol. 2008; 48:83–90. https://doi.org/10.1016/j.jhep.2007.08.018 [PubMed]
- 58. Wang SN, Lee KT, Tsai CJ, Chen YJ, Yeh YT. Phosphorylated p38 and JNK MAPK proteins in hepatocellular carcinoma. Eur J Clin Invest. 2012; 42:1295–301. https://doi.org/10.1111/eci.12003 [PubMed]
- 59. Yang SF, Wang SN, Wu CF, Yeh YT, Chai CY, Chunag SC, Sheen MC, Lee KT. Altered p-STAT3 (tyr705) expression is associated with histological grading and intratumour microvessel density in hepatocellular carcinoma. J Clin Pathol. 2007; 60:642–48. https://doi.org/10.1136/jcp.2006.036970 [PubMed]
- 60. Hajjou M, Norel R, Carver R, Marion P, Cullen J, Rogler LE, Rogler CE. cDNA microarray analysis of HBV transgenic mouse liver identifies genes in lipid biosynthetic and growth control pathways affected by HBV. J Med Virol. 2005; 77:57–65. https://doi.org/10.1002/jmv.20427 [PubMed]
- 61. Patil MA, Chua MS, Pan KH, Lin R, Lih CJ, Cheung ST, Ho C, Li R, Fan ST, Cohen SN, Chen X, So S. An integrated data analysis approach to characterize genes highly expressed in hepatocellular carcinoma. Oncogene. 2005; 24:3737–47. https://doi.org/10.1038/sj.onc.1208479 [PubMed]
- 62. Segal E, Friedman N, Kaminski N, Regev A, Koller D. From signatures to models: understanding cancer using microarrays. Nat Genet. 2005; 37:S38–45. https://doi.org/10.1038/ng1561 [PubMed]
- 63. Okabe H, Satoh S, Kato T, Kitahara O, Yanagawa R, Yamaoka Y, Tsunoda T, Furukawa Y, Nakamura Y. Genome-wide analysis of gene expression in human hepatocellular carcinomas using cDNA microarray: identification of genes involved in viral carcinogenesis and tumor progression. Cancer Res. 2001; 61:2129–37. [PubMed]
- 64. De Ponti A, Wiechert L, Schneller D, Pusterla T, Longerich T, Hogg N, Vogel A, Schirmacher P, Hess J, Angel P. A pro-tumorigenic function of S100A8/A9 in carcinogen-induced hepatocellular carcinoma. Cancer Lett. 2015; 369:396–404. https://doi.org/10.1016/j.canlet.2015.09.005 [PubMed]
- 65. Nemeth J, Stein I, Haag D, Riehl A, Longerich T, Horwitz E, Breuhahn K, Gebhardt C, Schirmacher P, Hahn M, Ben-Neriah Y, Pikarsky E, Angel P, et al. S100A8 and S100A9 are novel nuclear factor kappa B target genes during malignant progression of murine and human liver carcinogenesis. Hepatology. 2009; 50:1251–62. https://doi.org/10.1002/hep.23099 [PubMed]
- 66. Nishimura J, Dewa Y, Okamura T, Muguruma M, Jin M, Saegusa Y, Umemura T, Mitsumori K. Possible involvement of oxidative stress in fenofibrate-induced hepatocarcinogenesis in rats. Arch Toxicol. 2008; 82:641–54. https://doi.org/10.1007/s00204-007-0278-2 [PubMed]
- 67. Frau M, Simile MM, Tomasi ML, Demartis MI, Daino L, Seddaiu MA, Brozzetti S, Feo CF, Massarelli G, Solinas G, Feo F, Lee JS, Pascale RM. An expression signature of phenotypic resistance to hepatocellular carcinoma identified by cross-species gene expression analysis. Cell Oncol (Dordr). 2012; 35:163–73. https://doi.org/10.1007/s13402-011-0067-z [PubMed]
- 68. Markowitz J, Carson WE
3rd . Review of S100A9 biology and its role in cancer. Biochim Biophys Acta. 2013; 1835:100–09. https://doi.org/10.1016/j.bbcan.2012.10.003 [PubMed] - 69. Duan L, Wu R, Zhang X, Wang D, You Y, Zhang Y, Zhou L, Chen W. HBx-induced S100A9 in NF-κB dependent manner promotes growth and metastasis of hepatocellular carcinoma cells. Cell Death Dis. 2018; 9:629. https://doi.org/10.1038/s41419-018-0512-2 [PubMed]
- 70. Jeoung NH. Pyruvate Dehydrogenase Kinases: Therapeutic Targets for Diabetes and Cancers. Diabetes Metab J. 2015; 39:188–97. https://doi.org/10.4093/dmj.2015.39.3.188 [PubMed]
- 71. Zhang SL, Hu X, Zhang W, Yao H, Tam KY. Development of pyruvate dehydrogenase kinase inhibitors in medicinal chemistry with particular emphasis as anticancer agents. Drug Discov Today. 2015; 20:1112–19. https://doi.org/10.1016/j.drudis.2015.03.012 [PubMed]
- 72. Saunier E, Benelli C, Bortoli S. The pyruvate dehydrogenase complex in cancer: An old metabolic gatekeeper regulated by new pathways and pharmacological agents. Int J Cancer. 2016; 138:809–17. https://doi.org/10.1002/ijc.29564 [PubMed]
- 73. Stacpoole PW. Therapeutic Targeting of the Pyruvate Dehydrogenase Complex/Pyruvate Dehydrogenase Kinase (PDC/PDK) Axis in Cancer. J Natl Cancer Inst. 2017; 109:djx071. https://doi.org/10.1093/jnci/djx071 [PubMed]
- 74. Sugden MC, Holness MJ. Recent advances in mechanisms regulating glucose oxidation at the level of the pyruvate dehydrogenase complex by PDKs. Am J Physiol Endocrinol Metab. 2003; 284:E855–62. https://doi.org/10.1152/ajpendo.00526.2002 [PubMed]
- 75. Zhao L, Tang M, Hu Z, Yan B, Pi W, Li Z, Zhang J, Zhang L, Jiang W, Li G, Qiu Y, Hu F, Liu F, et al. miR-504 mediated down-regulation of nuclear respiratory factor 1 leads to radio-resistance in nasopharyngeal carcinoma. Oncotarget. 2015; 6:15995–6018. https://doi.org/10.18632/oncotarget.4138 [PubMed]
- 76. Blouin JM, Penot G, Collinet M, Nacfer M, Forest C, Laurent-Puig P, Coumoul X, Barouki R, Benelli C, Bortoli S. Butyrate elicits a metabolic switch in human colon cancer cells by targeting the pyruvate dehydrogenase complex. Int J Cancer. 2011; 128:2591–601. https://doi.org/10.1002/ijc.25599 [PubMed]
- 77. Grassian AR, Metallo CM, Coloff JL, Stephanopoulos G, Brugge JS. Erk regulation of pyruvate dehydrogenase flux through PDK4 modulates cell proliferation. Genes Dev. 2011; 25:1716–33. https://doi.org/10.1101/gad.16771811 [PubMed]
- 78. Sun Y, Daemen A, Hatzivassiliou G, Arnott D, Wilson C, Zhuang G, Gao M, Liu P, Boudreau A, Johnson L, Settleman J. Metabolic and transcriptional profiling reveals pyruvate dehydrogenase kinase 4 as a mediator of epithelial-mesenchymal transition and drug resistance in tumor cells. Cancer Metab. 2014; 2:20. https://doi.org/10.1186/2049-3002-2-20 [PubMed]
- 79. Wang Z, Li Z, Ye Y, Xie L, Li W. Oxidative Stress and Liver Cancer: Etiology and Therapeutic Targets. Oxid Med Cell Longev. 2016; 2016:7891574. https://doi.org/10.1155/2016/7891574 [PubMed]
- 80. Perl A, Hanczko R, Telarico T, Oaks Z, Landas S. Oxidative stress, inflammation and carcinogenesis are controlled through the pentose phosphate pathway by transaldolase. Trends Mol Med. 2011; 17:395–403. https://doi.org/10.1016/j.molmed.2011.01.014 [PubMed]
- 81. Fu Y, Chung FL. Oxidative stress and hepatocarcinogenesis. Hepatoma Res. 2018; 4:39. https://doi.org/10.20517/2394-5079.2018.29 [PubMed]
- 82. Bartucci R, Salvati A, Olinga P, Boersma YL. Vanin 1: Its Physiological Function and Role in Diseases. Int J Mol Sci. 2019; 20:3891. https://doi.org/10.3390/ijms20163891 [PubMed]
- 83. Berruyer C, Martin FM, Castellano R, Macone A, Malergue F, Garrido-Urbani S, Millet V, Imbert J, Duprè S, Pitari G, Naquet P, Galland F. Vanin-1-/- mice exhibit a glutathione-mediated tissue resistance to oxidative stress. Mol Cell Biol. 2004; 24:7214–24. https://doi.org/10.1128/MCB.24.16.7214-7224.2004 [PubMed]
- 84. Lin ZY, Chuang WL. Pharmacologic concentrations of melatonin have diverse influence on differential expressions of angiogenic chemokine genes in different hepatocellular carcinoma cell lines. Biomed Pharmacother. 2010; 64:659–62. https://doi.org/10.1016/j.biopha.2010.09.006 [PubMed]
- 85. Zhang B, Lo C, Shen L, Sood R, Jones C, Cusmano-Ozog K, Park-Snyder S, Wong W, Jeng M, Cowan T, Engleman EG, Zehnder JL. The role of vanin-1 and oxidative stress-related pathways in distinguishing acute and chronic pediatric ITP. Blood. 2011; 117:4569–79. https://doi.org/10.1182/blood-2010-09-304931 [PubMed]
- 86. Higgs MR, Chouteau P, Lerat H. 'Liver let die': oxidative DNA damage and hepatotropic viruses. J Gen Virol. 2014; 95:991–1004. https://doi.org/10.1099/vir.0.059485-0 [PubMed]
- 87. Yang SF, Chang CW, Wei RJ, Shiue YL, Wang SN, Yeh YT. Involvement of DNA damage response pathways in hepatocellular carcinoma. Biomed Res Int. 2014; 2014:153867. https://doi.org/10.1155/2014/153867 [PubMed]
- 88. Eckerdt F, Yuan J, Saxena K, Martin B, Kappel S, Lindenau C, Kramer A, Naumann S, Daum S, Fischer G, Dikic I, Kaufmann M, Strebhardt K. Polo-like kinase 1-mediated phosphorylation stabilizes Pin1 by inhibiting its ubiquitination in human cells. J Biol Chem. 2005; 280:36575–83. https://doi.org/10.1074/jbc.m504548200 [PubMed]
- 89. Helmke C, Becker S, Strebhardt K. The role of Plk3 in oncogenesis. Oncogene. 2016; 35:135–47. https://doi.org/10.1038/onc.2015.105 [PubMed]
- 90. Herzog CR. Chk2 meets Plk3 in damage control. Cell Cycle. 2002; 1:408–09. https://doi.org/10.4161/cc.1.6.268 [PubMed]
- 91. Strebhardt K. Multifaceted polo-like kinases: drug targets and antitargets for cancer therapy. Nat Rev Drug Discov. 2010; 9:643–60. https://doi.org/10.1038/nrd3184 [PubMed]
- 92. Xu D, Dai W, Li C. Polo-like kinase 3, hypoxic responses, and tumorigenesis. Cell Cycle. 2017; 16:2032–36. https://doi.org/10.1080/15384101.2017.1373224 [PubMed]
- 93. Nakagawa H, Maeda S. Inflammation- and stress-related signaling pathways in hepatocarcinogenesis. World J Gastroenterol. 2012; 18:4071–81. https://doi.org/10.3748/wjg.v18.i31.4071 [PubMed]
- 94. Sun B, Karin M. Obesity, inflammation, and liver cancer. J Hepatol. 2012; 56:704–13. https://doi.org/10.1016/j.jhep.2011.09.020 [PubMed]
- 95. Ringelhan M, Pfister D, O'Connor T, Pikarsky E, Heikenwalder M. The immunology of hepatocellular carcinoma. Nat Immunol. 2018; 19:222–32. https://doi.org/10.1038/s41590-018-0044-z [PubMed]
- 96. Sun B, Karin M. Inflammation and liver tumorigenesis. Front Med. 2013; 7:242–54. https://doi.org/10.1007/s11684-013-0256-4 [PubMed]
- 97. Li Y, Qu P, Wu L, Li B, Du H, Yan C. Api6/AIM/Spα/CD5L overexpression in alveolar type II epithelial cells induces spontaneous lung adenocarcinoma. Cancer Res. 2011; 71:5488–99. https://doi.org/10.1158/0008-5472.CAN-10-4225 [PubMed]
- 98. Li X, Ni R, Chen J, Liu Z, Xiao M, Jiang F, Lu C. The presence of IGHG1 in human pancreatic carcinomas is associated with immune evasion mechanisms. Pancreas. 2011; 40:753–61. https://doi.org/10.1097/MPA.0b013e318213d51b [PubMed]
- 99. Baghdadi M, Jinushi M. The impact of the TIM gene family on tumor immunity and immunosuppression. Cell Mol Immunol. 2014; 11:41–48. https://doi.org/10.1038/cmi.2013.57 [PubMed]
- 100. Zhang Q, Wang H, Wu X, Liu B, Liu W, Wang R, Liang X, Ma C, Gao L. TIM-4 promotes the growth of non-small-cell lung cancer in a RGD motif-dependent manner. Br J Cancer. 2015; 113:1484–92. https://doi.org/10.1038/bjc.2015.323 [PubMed]
- 101. Pineiro M, Alava MA, Gonzalez-Ramon N, Osada J, Lasierra P, Larrad L, Pineiro A, Lampreave F. ITIH4 serum concentration increases during acute-phase processes in human patients and is up-regulated by interleukin-6 in hepatocarcinoma HepG2 cells. Biochem Biophys Res Commun. 1999; 263:224–29. https://doi.org/10.1006/bbrc.1999.1349 [PubMed]
- 102. Lin X, Jia J, Du T, Li W, Wang X, Wei J, Lin X, Zeng H, Yao L, Chen X, Zhuang J, Weng J, Liu Y, et al. Overexpression of miR-155 in the liver of transgenic mice alters the expression profiling of hepatic genes associated with lipid metabolism. PLoS One. 2015; 10:e0118417. https://doi.org/10.1371/journal.pone.0118417 [PubMed]
- 103. Lin X, Chen L, Li H, Liu Y, Guan Y, Li X, Jia Z, Lin X, Jia J, Sun Y, Xiao D. miR-155 accelerates proliferation of mouse hepatocytes during liver regeneration by directly targeting SOCS1. Am J Physiol Gastrointest Liver Physiol. 2018; 315:G443–53. https://doi.org/10.1152/ajpgi.00072.2018 [PubMed]
- 104. Rong XX, Wei F, Lin XL, Qin YJ, Chen L, Wang HY, Shen HF, Jia LT, Xie RY, Lin TY, Hao WC, Yang J, Yang S, et al. Recognition and killing of cancer stem-like cell population in hepatocellular carcinoma cells by cytokine-induced killer cells via NKG2d-ligands recognition. Oncoimmunology. 2016; 5:e1086060. https://doi.org/10.1080/2162402x.2015.1086060 [PubMed]
- 105. Shi JW, Zhang TT, Liu W, Yang J, Lin XL, Jia JS, Shen HF, Wang SC, Li J, Zhao WT, Gu WW, Sun Y, Xiao D. Direct conversion of pig fibroblasts to chondrocyte-like cells by c-Myc. Cell Death Discov. 2019; 5:55. https://doi.org/10.1038/s41420-018-0136-4 [PubMed]
- 106. Li X, Zhao Z, Zhang X, Yang S, Lin X, Yang X, Lin X, Shi J, Wang S, Zhao W, Li J, Gao F, Liu M, et al. Klf4 reduces stemness phenotype, triggers mesenchymal-epithelial transition (MET)-like molecular changes, and prevents tumor progression in nasopharygeal carcinoma. Oncotarget. 2017; 8:93924–41. https://doi.org/10.18632/oncotarget.21370 [PubMed]
- 107. Liu YM, Liu W, Jia JS, Chen BZ, Chen HW, Liu Y, Bie YN, Gu P, Sun Y, Xiao D, Gu WW. Abnormalities of hair structure and skin histology derived from CRISPR/Cas9-based knockout of phospholipase C-delta 1 in mice. J Transl Med. 2018; 16:141. https://doi.org/10.1186/s12967-018-1512-9 [PubMed]
- 108. Sun Y, Xiao D, Li HA, Jiang JF, Li Q, Zhang RS, Chen XG. Phenotypic changes of human cells in human-rat liver during partial hepatectomy-induced regeneration. World J Gastroenterol. 2009; 15:3611–20. https://doi.org/10.3748/wjg.15.3611 [PubMed]
- 109. Wang SC, Lin XL, Wang HY, Qin YJ, Chen L, Li J, Jia JS, Shen HF, Yang S, Xie RY, Wei F, Gao F, Rong XX, et al. Hes1 triggers epithelial-mesenchymal transition (EMT)-like cellular marker alterations and promotes invasion and metastasis of nasopharyngeal carcinoma by activating the PTEN/AKT pathway. Oncotarget. 2015; 6:36713–30. https://doi.org/10.18632/oncotarget.5457 [PubMed]
- 110. Gray PC, Shani G, Aung K, Kelber J, Vale W. Cripto binds transforming growth factor beta (TGF-beta) and inhibits TGF-beta signaling. Mol Cell Biol. 2006; 26:9268–78. https://doi.org/10.1128/MCB.01168-06 [PubMed]
- 111. Shi JW, Liu W, Zhang TT, Wang SC, Lin XL, Li J, Jia JS, Sheng HF, Yao ZF, Zhao WT, Zhao ZL, Xie RY, Yang S, et al. The enforced expression of c-Myc in pig fibroblasts triggers mesenchymal-epithelial transition (MET) via F-actin reorganization and RhoA/Rock pathway inactivation. Cell Cycle. 2013; 12:1119–27. https://doi.org/10.4161/cc.24164 [PubMed]
- 112. Watanabe K, Hamada S, Bianco C, Mancino M, Nagaoka T, Gonzales M, Bailly V, Strizzi L, Salomon DS. Requirement of glycosylphosphatidylinositol anchor of Cripto-1 for trans activity as a Nodal co-receptor. J Biol Chem. 2007; 282:35772–86. https://doi.org/10.1074/jbc.m707351200 [PubMed]
- 113. Zheng L, Njauw CN, Martins-Green M. A hCXCR1 transgenic mouse model containing a conditional color-switching system for imaging of hCXCL8/IL-8 functions in vivo. J Leukoc Biol. 2007; 82:1247–56. https://doi.org/10.1189/jlb.0307141 [PubMed]
- 114. Zheng L, Njauw CN, Martins-Green M. A one-plasmid conditional color-switching transgenic system for multimodal bioimaging. Transgenic Res. 2008; 17:741–47. https://doi.org/10.1007/s11248-007-9160-5 [PubMed]
- 115. Nagy A, Gertsenstein M, Vintersten K, Behringer R. Manipulating the mouse embryo : a laboratory manual. Cold Spring Harbor Laboratory Press. 2003.
- 116. Lin X, Jia J, Qin Y, Lin X, Li W, Xiao G, Li Y, Xie R, Huang H, Zhong L, Wu Q, Wang W, Huang W, et al. Simple and rapid determination of homozygous transgenic mice via in vivo fluorescence imaging. Oncotarget. 2015; 6:39073–87. https://doi.org/10.18632/oncotarget.5535 [PubMed]
- 117. Wei F, Rong XX, Xie RY, Jia LT, Wang HY, Qin YJ, Chen L, Shen HF, Lin XL, Yang J, Yang S, Hao WC, Chen Y, et al. Cytokine-induced killer cells efficiently kill stem-like cancer cells of nasopharyngeal carcinoma via the NKG2D-ligands recognition. Oncotarget. 2015; 6:35023–39. https://doi.org/10.18632/oncotarget.5280 [PubMed]
- 118. Jia J, Lin X, Lin X, Lin T, Chen B, Hao W, Cheng Y, Liu Y, Dian M, Yao K, Xiao D, Gu W. R/L, a double reporter mouse line that expresses luciferase gene upon Cre-mediated excision, followed by inactivation of mRFP expression. Genome. 2016; 59:816–26. https://doi.org/10.1139/gen-2016-0090 [PubMed]