Introduction
Lung cancer is the leading cause of cancer-related mortality worldwide, and non-small-cell lung cancer (NSCLC) represents the major histological subtype of the disease [1]. In recent years, studies have shown that triggering the antitumor immunity of CD8+ T cells in the tumor microenvironment (TME) by immunotherapy brings prominent and long-lasting clinical benefits for NSCLC [2]. However, only a small number of patients could benefit from such approaches alone [3]. The possible explanation is that TME is a complex ecosystem consisting of various cells and its formation and dynamic changes are regulated by multiple targets, multiple signaling pathways and multiple cells [4]. The tumor microenvironment (TME) is composed not only of tumor cells but also of stromal cells, inflammatory cells, vasculature, and extracellular matrices (ECM). Tumor cells escape immunosurveillance by increasing signaling through coinhibitory receptors of T cells or reducing the availability of antigen for presentation. In the TME, inflammatory cells secrete chemokines and cytokines to promote the growth and metastasis of tumor cells, genetic mutations, and angiogenesis [5]. The tumor vasculature excludes of effector lymphocytes from the tumor microenvironment by physical means. ECM enables cancer cells to subvert the immune component of the microenvironment via providing both a biochemical and biomechanical context. Stromal cells express numerous surface and secreted molecules such as IL-6, NO, PGE2 and TGFβ which directly suppress CD4+ and CD8+T cells and activate immunosuppressive myeloid cells. For this reason, combination strategy toward multi-target anticancer therapy is a promising field in development of immunotherapeutic drug to overcome this challenge [6].
Accumulated studies have shown that natural products can exert anti-tumor effects by regulating immune-related signaling pathways and targets [7–11]. At the same time, natural products have multi-target and multi-pathway characteristics compared with conventional therapeutic drugs, which can play a key role in the regulation of complex networks in TME [12–15]. Thus, natural products that promote these effects via a wide variety of mechanisms are gradually becoming a huge resource pool for the development of antitumor drugs [16–18]. For example, Epimedium, a famous herbal medicine, is widely used as a tonic, aphrodisiac and antirheumatic in China, Japan and Korea for more than 2000 years. Studies and clinical practices have demonstrated that Epimedium is a relatively non-toxic nutritional herbal product and has wide pharmacological actions, especially in immunological function modulation, anti-tumor [19–21]. Hitherto, more than 100 compounds have been isolated from Epimedium. However, it is unclear that which compound determine the pharmacological effect, and the mechanism(s) of how Epimedium promotes the antitumor immunity in NSCLC remains enigmatic.
In this work, we used a systems pharmacology approach to elucidate the action mechanism of polypharmacology molecules of Epimedium targeting TME for the treatment of NSCLC. We screened the polypharmacology molecules of Epimedium, predicted the targets of active compounds, constructed the networks, and linked the targets to the immune phenotype in lung cancer patients. These results indicated that polypharmacology molecules of Epimedium targeted several kinds of tumor-related factors and signaling pathways in the TME, such as inflammation, apoptosis, and migration. Additionally, we used in vivo and in vitro experiments to verify the antitumor efficacy of icaritin (ICT), since it is a major active ingredients of Epimedium and was predicted to have a good drug-like and pharmacological properties in targeting the NSCLC immune microenvironment. The LLC tumor-bearing mice model evidenced that tumor volume was significantly reduced and survival time was significantly extended after treatment with ICT. Furthermore, transcriptome sequencing and immunofluorescence showed that ICT could target the TME to upregulate the immune signaling pathway and T cell chemokines, thereby increasing the infiltration of CD8+ T cells to achieve antitumor immunity. In conclusion, systems pharmacology provided an important reference to insight into the action mechanism of polypharmacology molecules of Epimedium targeting TME against NSCLC, and offered a significant strategy for development and application of natural products for immunotherapy. As Figure 1 showed the process flow framework of this study.
Figure 1. The workflow of the systems pharmacology method.
Results
Polypharmacology molecules of Epimedium for the treatment of NSCLC
To elucidate the polypharmacology molecules of Epimedium targeting TME on NSCLC, we firstly achieved the compounds in Epimedium by searching literatures and Traditional Chinese Medicine Systems Pharmacology Database (TCMSP), which results in 130 compounds. Then, 16 ingredients of Epimedium with favorable pharmacokinetic characteristics were determined by ADME (Absorption, Distribution, Metabolism, and Excretion) screening conditions satisfied as previous study [22], i.e., oral bioavailability (OB) ≥ 30%, half-life (HL) ≥ 4 and drug-likeness (DL) ≥ 0.18 (Table 1). Among them, flavonoid plays an important role in the treatment and prevention of various cancers. For instance, ICT (MOL056 OB=45.41%, DL=0.44, HL=15.01) has various beneficial activities which inhibit tumor growth or target the TME [23–25]. Kaempferol (MOL044 OB=41.88%, DL=0.24, HL=14.74) induces cell cycle arrest in G1 and G2/M by inhibiting the activity of CDK2, CDK4 and Cdc2, leading to apoptosis [26]. Moreover, sitosterol (MOL042 OB=36.91%, DL=0.75, HL=5.37) has been shown antimigration in various lung cancer cell types [27–29]. Linoleate acetate (MOL008 OB = 42.10%, DL = 0.20, HL = 7.48) is used in the treatment of NSCLC to inhibit inflammatory factors COX, LOX and TNF-α in the TME, which were closely related to their immune activities [30, 31].
Table 1. Chemical information and network parameters of 16 active compounds from Epimedium.
| MOL-ID | Compounds | Structure | Categories | OB | DL | HL | Degree | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MOL056 | Icaritin | Flavonoids | 45.41 | 0.44 | 15.01 | 32 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MOL044 | Kaempferol | Flavonoids | 41.88 | 0.24 | 14.74 | 31 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MOL033 | Chrysoeriol | Flavonoids | 35.85 | 0.27 | 16.31 | 28 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MOL039 | 8-Isopentenyl-kaempferol | Flavonoids | 38.04 | 0.39 | 15.37 | 24 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MOL067 | Yinyanghuo C | Flavonoids | 45.67 | 0.50 | 15.74 | 24 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MOL069 | Yinyanghuo E | Flavonoids | 51.63 | 0.55 | 15.47 | 22 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MOL074 | 8-(3-methylbut-2-enyl)-2-phenyl-chromone | Flavonoids | 48.54 | 0.25 | 18.73 | 20 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MOL012 | (2S)-7-hydroxy-2-(4-hydroxyphenyl)-4-chromanone | Flavonoids | 32.76 | 0.18 | 17.89 | 19 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MOL065 | Yinyanghuo A | Flavonoids | 56.96 | 0.77 | 14.44 | 13 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MOL106 | Icariin | Flavonoids | 41.58 | 0.61 | 19.93 | 11 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MOL004 | 24-epicampesterol | Sterols | 37.58 | 0.71 | 4.50 | 18 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MOL010 | Poriferast-5-en-3beta-ol | Sterols | 36.91 | 0.75 | 5.07 | 17 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MOL042 | Sitosterol | Sterols | 36.91 | 0.75 | 5.37 | 17 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MOL008 | Linoleyl acetate | Lignans | 42.10 | 0.20 | 7.48 | 18 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MOL063 | C-Homoerythrinan, 1,6-didehydro 3,15,16-trimethoxy-, (3.beta.)- | Others | 39.14 | 0.49 | 6.58 | 18 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MOL077 | Anhydroicaritin-3-O-alpha-L-rhamnoside | Others | 41.58 | 0.61 | 16.23 | 12 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| OB, oral bioavailability; DL, drug-likeness; HL, Half-life. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Polypharmacology mechanism of Epimedium targeting TME to treat NSCLC
To clarify the potential action mechanism of the multi-target of candidate compounds targeting TME for the treatment of NSCLC. 389 candidate targets for the 16 potential active compounds were obtained by utilizing the similarity ensemble approach (SEA), weighted ensemble similarity (WES) and systematic drug targeting tool (SysDT) methods. After screening, we retrieved 89 potential targets (Table 2 and Supplementary Tables 1, 2). ESR1 and AR are the morphogenetic factors that regulate cell proliferation, differentiation, angiogenesis, and the development of some tumors [32, 33], which were predicted to link the Epimedium to cancer with highest scores. In line with our prediction, studies show that ICT can target ESR1 and AR to control the growth of advanced breast and prostate cancer [24, 34], which confirms the predictive accuracy of our approach. Except for ESR1 and AR, we also predicted many targets related to immune and inflammation with high scores, such as PPARG, PTGS2, PTGS1, ALOX5. In the subsequent study, we mainly focused on these targets. Related studies have shown that regulating the expression of these targets could promote apoptosis and inhibit inflammation in the TME. For example, the COX2 has a variety of biological activities such as inhibiting cell apoptosis, promoting cell proliferation, inhibiting immune surveillance and promoting angiogenesis in the TME, thus playing a key role in the occurrence and development of tumors [35–37]. Therefore, we described the relationship between targets and NSCLC from the following three aspects.
Table 2. The targets information of Epimedium.
| UniProt-ID | Protein names | Gene names | Degree | Species |
| P03372 | Estrogen receptor | ESR1 | 14 | homo sapiens |
| P10275 | Androgen receptor | AR | 14 | homo sapiens |
| Q92731 | Progesterone receptor | ESR2 | 12 | homo sapiens |
| P37231 | Peroxisome proliferator activated receptor gamma | PPARG | 11 | homo sapiens |
| P35354 | Prostaglandin G/H synthase 2 | PTGS2 | 11 | homo sapiens |
| Q9UNQ0 | ATP-binding cassette sub-family G member 2 | ABCG2 | 10 | homo sapiens |
| P49841 | Glycogen synthase kinase-3 beta | GSK3B | 10 | homo sapiens |
| Q16678 | Cytochrome P450 1B1 | CYP1B1 | 10 | homo sapiens |
| P23219 | Prostaglandin G/H synthase 1 | PTGS1 | 9 | homo sapiens |
| P11309 | Proto-oncogene serine/threonine-protein kinase Pim-1 | PIM1 | 9 | homo sapiens |
| P35228 | Nitric oxide synthase, inducible | NOS2 | 8 | homo sapiens |
| Q16539 | Mitogen-activated protein kinase 14 | MAPK14 | 8 | homo sapiens |
| P24941 | Cell division protein kinase 2 | CDK2 | 8 | homo sapiens |
| P47989 | Xanthine dehydrogenase/oxidase | XDH | 8 | homo sapiens |
| P16220 | Cyclic AMP-responsive element-binding protein 1, CREB-1, cAMP-responsive element-binding protein 1 | CREB1 | 7 | homo sapiens |
| P53355 | Death-associated protein kinase 1 | DAPK1 | 7 | homo sapiens |
| P60568 | Interleukin-2 | IL2 | 7 | homo sapiens |
| P51812 | Death-associated protein kinase 1 | RPS6KA3 | 7 | homo sapiens |
| P08183 | Multidrug resistance protein 1 | ABCB1 | 7 | homo sapiens |
| P15692 | Vascular endothelial growth factor A | VEGFA | 6 | homo sapiens |
| P15121 | Aldo-keto reductase family 1 member B1 | AKR1B1 | 6 | homo sapiens |
| P33527 | ATP-binding cassette sub-family C member 1 | ABCC1 | 6 | homo sapiens |
| P09917 | Arachidonate 5-lipoxygenase | ALOX5 | 5 | homo sapiens |
| P06401 | Progesterone receptor | PGR | 4 | homo sapiens |
| P00918 | Carbonic anhydrase 2 | CA2 | 4 | homo sapiens |
| P11511 | Aromatase | CYP19A1 | 4 | homo sapiens |
| P11473 | Vitamin D3 receptor | VDR | 4 | homo sapiens |
| P29474 | Nitric-oxide synthase, endothelial | NOS3 | 4 | homo sapiens |
| P07550 | Beta-2 adrenergic receptor | ADRB2 | 4 | homo sapiens |
| P02766 | Transthyretin | TTR | 4 | homo sapiens |
| Q12809 | Potassium voltage-gated channel subfamily H member 2 | KCNH2 | 4 | homo sapiens |
| Q08828 | Adenylate cyclase type 1 | ADCY1 | 3 | homo sapiens |
| P00915 | Carbonic anhydrase 1 | CA1 | 3 | homo sapiens |
| Q07973 | 1,25-dihydroxyvitamin D (3) 24-hydroxylase, mitochondrial | CYP24A1 | 3 | homo sapiens |
| O15439 | Multidrug resistance-associated protein 4 | ABCC4 | 3 | homo sapiens |
| P51449 | Nuclear receptor ROR-gamma | RORC | 3 | homo sapiens |
| Q12772 | Sterol regulatory element-binding protein 2 | SREBF2 | 3 | homo sapiens |
| P02774 | Vitamin D-binding protein | GC | 3 | homo sapiens |
| P48736 | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit, gamma isoform | PIK3CG | 3 | homo sapiens |
| P04798 | Cytochrome P450 1A1 | CYP1A1 | 3 | homo sapiens |
| O76074 | cGMP-specific 3',5'-cyclic phosphodiesterase | PDE5A | 3 | homo sapiens |
| P08709 | Coagulation factor VII | F7 | 3 | homo sapiens |
| Q9UBM7 | 7-dehydrocholesterol reductase | DHCR7 | 2 | homo sapiens |
| P04150 | Glucocorticoid receptor | NR3C1 | 2 | homo sapiens |
| P07451 | Carbonic anhydrase 3 | CA3 | 2 | homo sapiens |
| Q09472 | Histone acetyltransferase p300 | EP300 | 2 | homo sapiens |
| Q12791 | Calcium-activated potassium channel subunit alpha-1 | KCNMA1 | 2 | homo sapiens |
| P05091 | Aldehyde dehydrogenase, mitochondrial | ALDH2 | 2 | homo sapiens |
| Q04206 | Transcription factor p65 | RELA | 2 | homo sapiens |
| P27695 | DNA - (apurinic or apyrimidinic site) lyase | APEX1 | 2 | homo sapiens |
| Q9UHC3 | Acid-sensing ion channel 3 | ASIC3 | 2 | homo sapiens |
| P09382 | Galectin-1 | LGALS3 | 2 | homo sapiens |
| P14679 | Tyrosinase | TYR | 2 | homo sapiens |
| P05230 | Fibroblast growth factor 1 | FGF1 | 1 | homo sapiens |
| P21554 | Cannabinoid receptor 1 | CNR1 | 1 | homo sapiens |
| P34972 | Cannabinoid receptor 2 | CNR2 | 1 | homo sapiens |
| Q13822 | Ectonucleotide pyrophosphatase/phosphodiesterase family member 2 | ENPP2 | 1 | homo sapiens |
| P23141 | Liver carboxylesterase 1 | CES1 | 1 | homo sapiens |
| P15090 | Fatty acid-binding protein, adipocyte | FABP4 | 1 | homo sapiens |
| P17252 | Protein kinase C alpha type, PKC-A, PKC-alpha | PRKCA | 1 | homo sapiens |
| P47712 | Cytosolic phospholipase A2, cPLA2 | PLA2G4A | 1 | homo sapiens |
| Q53EL6 | Programmed cell death protein 4 | PDCD4 | 1 | homo sapiens |
| O95136 | Sphingosine 1-phosphate receptor 2, S1P receptor 2, S1P2 | S1PR2 | 1 | homo sapiens |
| P08842 | Steryl-sulfatase | STS | 1 | homo sapiens |
| O60603 | Toll-like receptor 2 | TLR2 | 1 | homo sapiens |
| Q8NER1 | Transient receptor potential cation channel subfamily V member 1 | TRPV1 | 1 | homo sapiens |
| P14780 | Matrix metalloproteinase-9 | MMP9 | 1 | homo sapiens |
| P05412 | Transcription factor AP-1 | JUN | 1 | homo sapiens |
| P00338 | L-lactate dehydrogenase A chain, LDH-A | LDHA | 1 | homo sapiens |
| Q9UDY8 | Mucosa-associated lymphoid tissue lymphoma translocation protein 1 | MALT1 | 1 | homo sapiens |
| Q16790 | Carbonic anhydrase 9 | CA9 | 1 | homo sapiens |
| P05177 | Cytochrome P450 1A2 | CYP1A2 | 1 | homo sapiens |
| P28702 | Retinoic acid receptor RXR-beta | RXRB | 1 | homo sapiens |
| P35367 | Histamine H1 receptor | HRH1 | 1 | homo sapiens |
| Q15788 | Nuclear receptor coactivator 1 | NCOA1 | 1 | homo sapiens |
| P15559 | NAD(P)H dehydrogenase [quinone] 1 | NQO1 | 1 | homo sapiens |
| P22460 | Potassium voltage-gated channel subfamily A member 5 | KCNA5 | 1 | homo sapiens |
| P08913 | Alpha-2A adrenergic receptor | ADRA2A | 1 | homo sapiens |
| P08908 | 5-hydroxytryptamine receptor 1A, 5-HT-1A, 5-HT1A | HTR1A | 1 | homo sapiens |
| P18089 | Alpha-2B adrenergic receptor | ADRA2B | 1 | homo sapiens |
| Q15822 | Neuronal acetylcholine receptor subunit alpha-2 | CHRNA2 | 1 | homo sapiens |
| P14416 | D (2) dopamine receptor | DRD2 | 1 | homo sapiens |
| P36544 | Neuronal acetylcholine receptor subunit alpha-7 | CHRNA7 | 1 | homo sapiens |
| P30532 | Neuronal acetylcholine receptor subunit alpha-5 | CHRNA5 | 1 | homo sapiens |
| Q9Y271 | Cysteinyl leukotriene receptor 1 | CYSLTR1 | 1 | homo sapiens |
| O43570 | Carbonic anhydrase 12 | CA12 | 1 | homo sapiens |
| P78527 | DNA-dependent protein kinase catalytic subunit, DNA-PK catalytic subunit, DNA-PKcs | PRKDC | 1 | homo sapiens |
| P02787 | Serotransferrin, Transferrin | TF | 1 | homo sapiens |
| Q9NYY3 | Serine/threonine-protein kinase PLK2 | PLK2 | 1 | homo sapiens |
Firstly, we analyzed the direct or indirect relationships between targets and diseases, as shown in Figure 2A. The radar chart analysis showed that the targets we screened were closely related to carcinoma (D002277), neoplasm metastasis (D009362), lung neoplasms (D008175), inflammation (D007249) and so on, and these targets simultaneously targeted two or more diseases. Moreover, these diseases were also affected by multiple targets simultaneously. For instance, lung neoplasms (D008175) were regulated by 23 targets such as iNOS, Bcl2 and PKC that have anti-inflammatory, pro-apoptotic effects in the TME. Then, we performed the Gene Ontology (GO) biological processes enrichment for these targets by using ClueGO and visualized them by Enrichment Map with the threshold of P-value ≤ 0.05 (Supplementary Table 3). We found that most of these targets were strongly associated with various biological processes in the TME, such as nuclear receptor activity, response to lipopolysaccharide, transcription initiation from RNA polymerase II promoter and so on (Supplementary Figure 1). Among these groups, regulation of inflammatory response, regulation of vasculature development and extrinsic apoptotic signaling pathway were all closely associated with NSCLC (Figure 2B). For example, by down-regulating angiogenic factors, the activation of vascular endothelial cells was inhibited, and the proliferation and migration of endothelial cells were prevented, thereby achieving the effect of treating tumors.
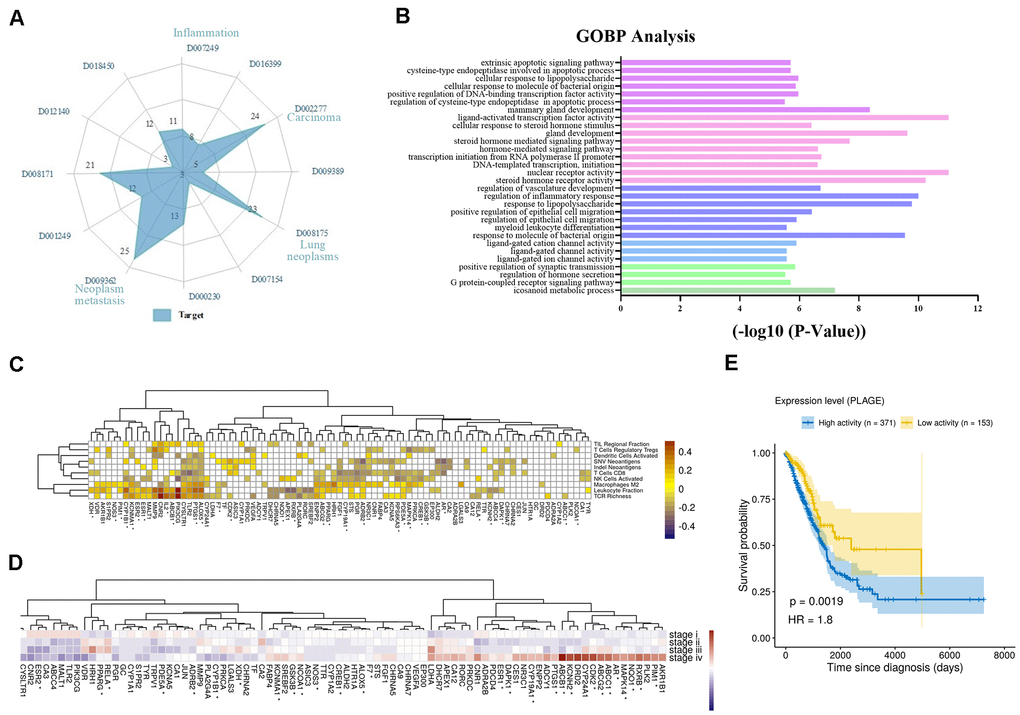
Figure 2. Analysis of multi-target of polypharmacology molecules of Epimedium treatment of NSCLC. (A) In an equi-angular spokes radar chart, each spoke represents a class of diseases, and the length of the spokes is the distribution data, which is proportional to the quantity of target proteins relative to the homologous disease. (B) The y-axis shows significantly enriched ‘Biological Process’ categories in GO of the target genes, and the x-axis shows the enrichment scores of these terms (P-value < 0.05). A specific color represents a class of biological processes. (C) The heatmap of Pearson correlation coefficients (PCCs) between gene expression level of Epimedium targets and immune phenotypes. The Benjamini-Hochberg (BH) adjusted p-values of PCCs <0.05 were shown as white color. Note that the genes labeled with an asterisk represents the targets for ICT. (D) Average expression of Epimedium targets among clinical staging (stages I–IV) in TCGA LUDA patients. For each gene, the expression was normalized by z-score transformation. Note that the genes labeled with an asterisk represents the targets for ICT. (E) Prognostic value of the Epimedium targets for overall survival of human LUAD patients comparing high and low activity. X-axis shows time for survival (days) and y-axis shows the probability of survival, where 1.0 corresponds to 100 percent.
In addition, we further analyzed the correlation between gene expression of Epimedium targets (rows) and scores of ten anti-tumor immunity related phenotypes (columns) within 533 LUDA patients in TCGA. The phenotypes include (i) the cell abundances of 6 major immune cell types (leukocyte, T cells CD8, activated NK cells, activated dendritic cells, Tregs, macrophages M2), which were deconvoluted from tumor transcriptome, (ii) the neoantigen number predicted from SNVs and Indels, (iii) TCR richness inferenced from tumor RNA-Seq and (iv) the TIL regional fraction estimated from the tissue image. These immune phenotypes were identified based on the methods established by llya Shmulevich et al. [38]. And the degree of association between targets and immune phenotypes is reflected by Pearson correlation coefficient. As shown in Figure 2C, we correlated targets with immune phenotypes, resulting in 315 significant associations (Benjamini–Hochberg adjusted p-values <0.05) that involved 82 of 89 targets. Leukocyte fraction correlated with CNR2, PI3KCG, TLR2, PTGS1, ALOX5 and so on. Among them, inactivation of CNR2 or activation of ALOX5 has been reported to enhance leukocyte migration [39]. T cells regulatory (Tregs) correlated with in PTGS1, ALOX5, CNR2 and others, which could be immune-evading mechanisms. T cells CD8 correlated with many targets, as well as PGR, XDH, MALT1, and ADCY1, which may relate to CD8+ T infiltration. Additionally, we observed that the expression of these targets is related to clinical progression as clinical staging correlated with increased/decreased target expression (Figure 2D). Further, we evaluated the overall expression levels of the target set among the LUAD patients using the PLAGE (Pathway Level Analysis of Gene Expression) approach. We observed that higher expression of these target genes in tumor samples was negatively associated with patient survival in LUAD (HR = 1.8, log-rank test P-value = 0.0019) (Figure 2E).
Finally, analysis of differentially expressed genes data from the Genomic Data Commons (GDC) Data Portal [40] indicated that most potential targets of Epimedium overlapped with differentially expressed genes (DEGs) of lung adenocarcinoma (LUAD) and lung squamous cell carcinoma (LUSC). As shown in Figure 3A, we found that 51.7% of the targets overlapped with the DEGs of LUSC, 41.6% overlapped with the DEGs of LUAD, and 33.7% overlapped with the DEGs of LUAD and LUSC, respectively. In addition, we built an expression heatmap of 89 potential targets of Epimedium active components and based on the correlation analysis of the expression matrix, we found that the degree of association among overlapping genes in the Venn diagram was significantly higher, such as DAPK1, MAPK14, CDK2, LDHA, CYP1A2, etc. These well-correlated targets are closely related to NSCLC. (Figure 3B and Supplementary Figure 2). Meanwhile, we found the correlation between the 54 genes overlapping with Epimedium and LUAD and LUSC was higher than the correlation between the random genes in LUAD and LUSC (Wasserstein Distance =0.0559) (Figure 3C). Taken together, these data suggested that the potential targets of polypharmacology molecules of Epimedium have the potential antitumor effect on the TME for the treatment of NSCLC.
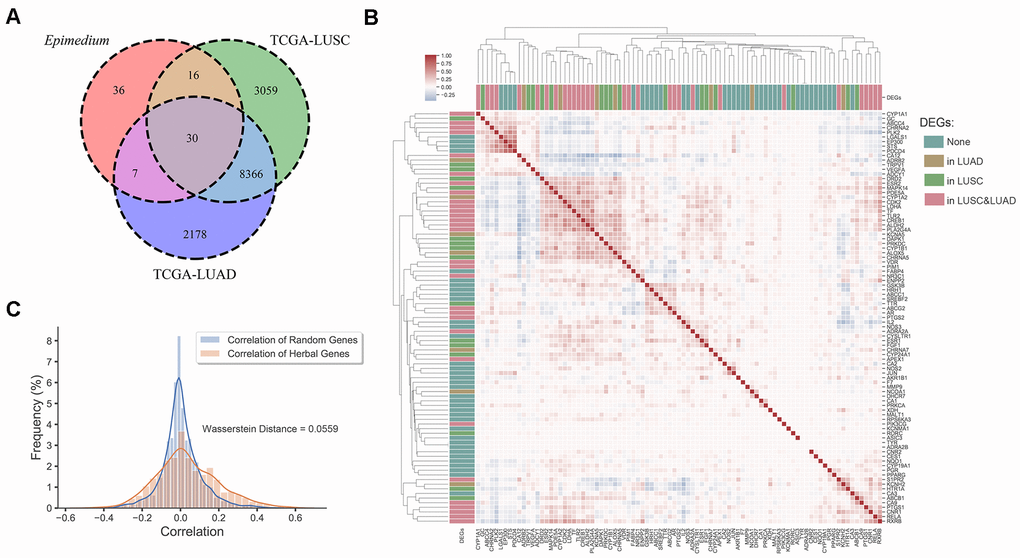
Figure 3. Differentially expression gene analysis. (A) Venn diagram of overlap genes in potential targets of Epimedium active ingredients and DEGs of LUSC and LUAD. Each region represents the number of genes. (B) Heatmap of the pairwise correlation of 89 potential genes of Epimedium active components. (C) Distribution diagram of random intergenic correlations and Epimedium intergenic correlation. Yellow indicates the correlation between the genes of Epimedium, and blue indicates the correlation between random genes (LUSC and LUAD).
Mechanism of polypharmacology molecules of Epimedium targeting TME for the treatment of NSCLC
To reveal action mechanism of the multi-compound and multi-target features of Epimedium targeting the TME in response to NSCLC, we analyzed the interaction of potential compounds, targets, pathways and diseases by compound-target (C-T) network, target-pathway (T-P) network and pathway-disease (P-D) chord diagram (Figure 4A–4C). For example, in the C-T network, ICT, kaempferol and sitosterol target proteins associated with inflammation, cancer cell apoptosis and migration in the TME [37, 41–43], such as COX2, IL-6, GSK3β, CDK2 and NOS3. For the T-P network, we found that about 22% of targets were involved in TNF, PI3K-Akt and VEGF signaling pathway, while P-D chord diagram showed that TNF, PI3K-Akt and VEGF signaling pathways were significantly enriched in Cancer and Neoplasm Metastasis diseases. Moreover, a "NSCLC-pathway" containing TNF, PI3K-Akt and VEGF signaling pathways was assembled and separated into two represent therapeutic modules (immunization module and tumor related module). (Figure 5).
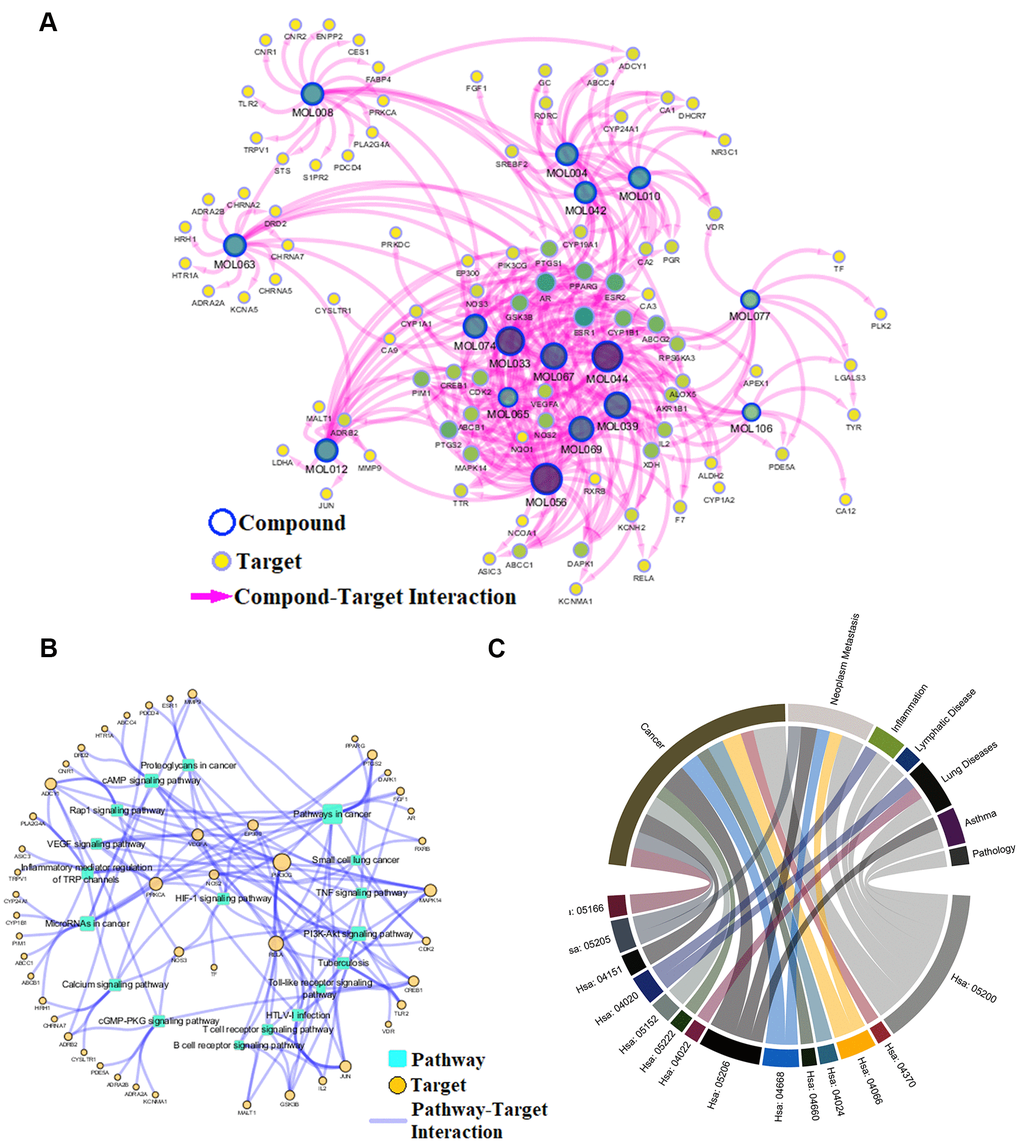
Figure 4. Network analysis. (A) C-T network. (B) T-P network. Yellow nodes represent potential targets; blue circles represent active ingredients of Epimedium; squares represent targets-related pathways; and edge represents the interaction between them. Node size is proportional to its degree. (C) P-D chord diagram. The circle represents the selected pathways and diseases, and the color of the bands within the circle reflects the relationship between the pathways and the diseases.
Immunization module
At present, inflammation has a close relationship with the onset of tumor, which not only promoted the occurrence and development of tumor, but also participated in various processes of tumor cells proliferation and migration in the TME. As shown in Figure 5, TNF-α, a member of the pro-inflammatory cytokine family, stimulated the recruitment of neutrophils and monocytes to the site of infection and promoted a sustained response to inflammation in the TME [44]. IL-6 is a cytokine that produces a broad range of cellular (macrophages and fibroblasts) and physiological responses upon activation, and that has been implicated in the tumorigenesis and tumor migration of epithelial cancer [45]. ICT (MOL056) could improve the TME by regulating TNF signaling pathway to inhibit inflammatory cells (neutrophils, monocytes, macrophages and fibroblasts) from expressing TNF-α and IL-6 and other inflammatory factors, thereby preventing the development of tumors. The results showed that the polypharmacology molecules of Epimedium played an anti-inflammatory role in the treatment of NSCLC by targeting the TNF signaling pathway.
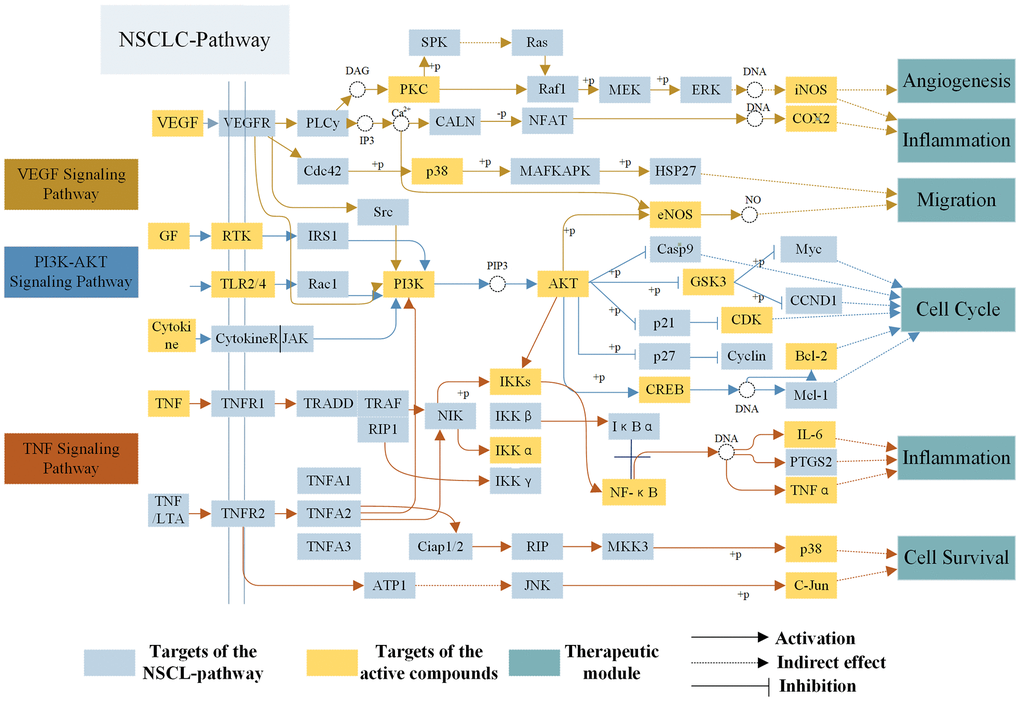
Figure 5. The NSCLC-pathway. Distribution of protein targets of Epimedium on the compressed ‘NSCLC-pathway’. Three pathways (VEGF, TNF, PI3K-AKT Signaling Pathway) form the compressed NSCLC pathway. Yellow rectangle remark targets on the NSCLC pathway, blue rectangle represent targets of active compounds and dark blue rectangle represent therapeutic modules. Solid arrows indicate activation, T-arrows indicate inhibition and dotted arrows indicate indirect effect.
In vivo, the anti-NSCLC effect of ICT
Based on the above systems pharmacology analysis, we selected ICT, a major active ingredient of Epimedium, possessing the highest degree in the C-T network and good pharmacological properties to verify the therapeutic effect of polypharmacology molecules of Epimedium on NSCLC. The LLC tumour models were constructed as stated in “Materials and methods” (Figure 6A). 5×105 LLC cells were injected subcutaneously in the left axilla of C57BL/6 mice. After three days, mice received daily intragastric administration of ICT or daily intraperitoneal administration of Taxol or daily intraperitoneal administration of diluent for 18 sequential days. Tumor volume and mice body weight were measured once every two days when diameter of tumor reached 5×5 mm after 6 days. At day 21, mice were sacrificed by cervical dislocation for molecule analysis. As shown in Figure 6B, 6C, we statistically analyzed the tumor volume and weight of mice and found that the ICT group and positive control group significantly inhibited the growth of tumor compared with the control group. Besides, as shown in Figure 6D, the ICT group and positive control group exhibited a significant increase in survival, compared with the control group. And no significant loss of mice weight was observed during the experiment (Supplementary Figure 3). These data suggested that ICT could inhibit tumor growth of LLC in mice and extend the life cycle of mice.
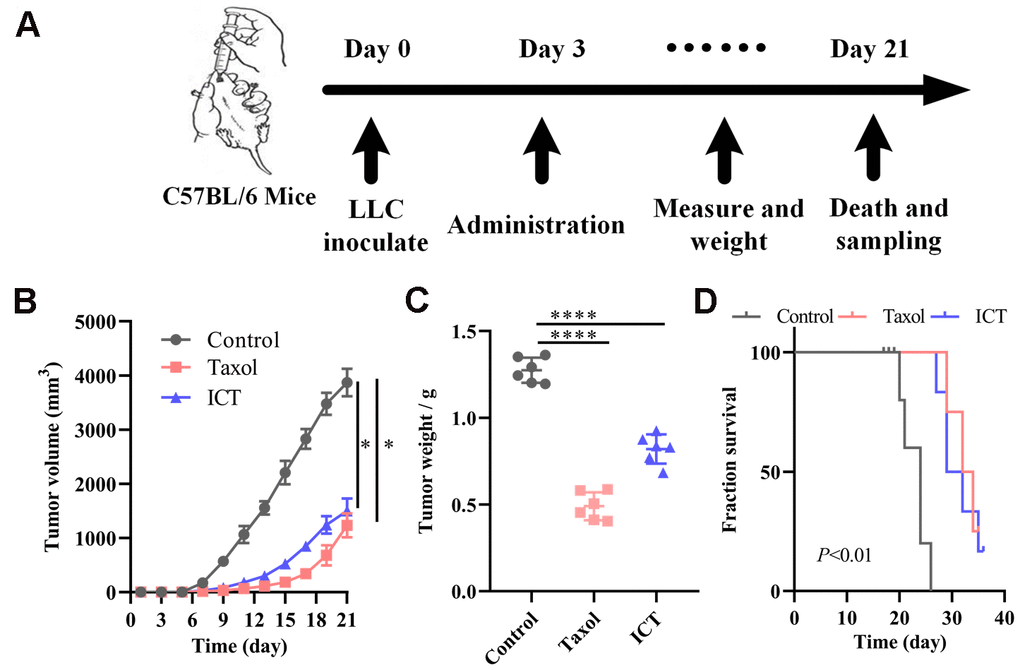
Figure 6. ICT treatment inhibited tumor growth in mice. (A) In vivo experimental treatment plan flow diagram. (B) The tumor volume in control, positive control, and ICT- treated mice was measured once every two days using digital calipers. (C) Weight of LLC tumors following sacrifice in mice subjected to daily administered for 21 days, and their controls. (D) Kaplan-Meier survival curves of mice which were inoculated with LLC cells. All data represent the means ± SEM of six mice. *P < 0.05, ****P < 0.0001 vs control group.
ICT altered the TME by increasing CD8+ T cell infiltration
To further explore the mechanisms of ICT in the treatment of NSCLC, we performed transcriptome sequencing for ICT-treated tumor (one sample) and control-treated tumor (one sample) (Supplementary Table 4). We used R LPEseq package to detect the differentially expressed genes (DEGs) with the thresholds of the p-adjust value ≤ 0.05 and |log2 fold change |≥1. LPEseq resulted 1779 upregulated and 596 downregulated DEGs in ICT-treated tumor, respectively. Surprisingly, we founded the 1779 upregulated DEGs in ICT-treated tumors were mainly enriched in immune system process, inflammatory response, NF−kappaB signaling and innate immune response, which indicated that ICT could target TME regulation immune pathway in murine tumor models (Figure 7A). A more detailed analysis revealed upregulation of genes involved in immune activation and immune cell homing, such as antigen presentation (MHC I and II), chemotaxis (CXCL9 and CXCL10) and immune cell homing (ICAM1) (Figure 7B). Then, we further confirmed that ICT treatment increased CXCL9 and CXCL10 secretion in the mouse LLC and the human H1975 lung cancer cell lines by RT-PCR (Figure 7C). Studies have shown that increased expression of T cell chemokines (such as CXCL9 and CXCL10) in the TME was closely related to immunotherapy [51].
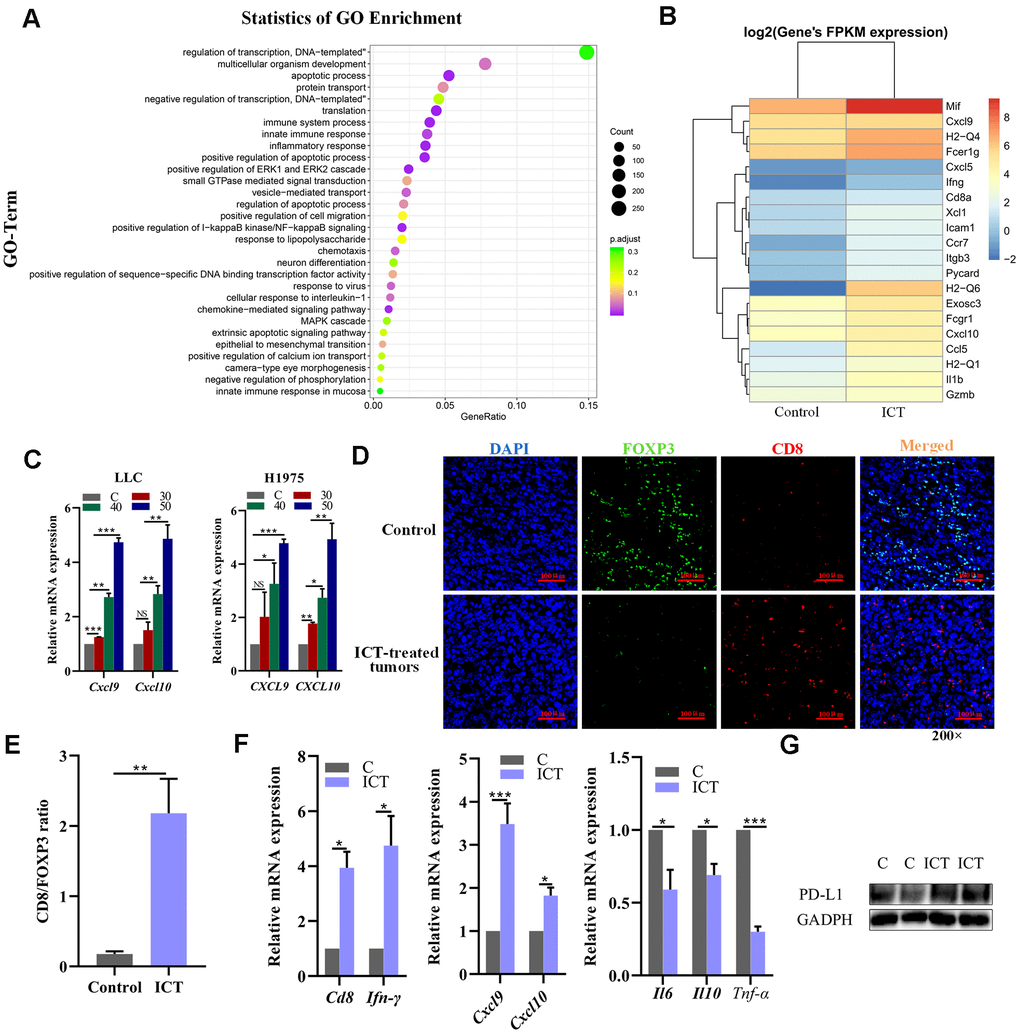
Figure 7. Analysis of increased CD8+ T cell infiltration in ICT-treated tumors. (A) GO enrichment analysis of differential genes between ICT-treated tumors and control group by ggplot2. The color represents the different adjusted P-values < 0.05, while the size of the circle represents the count. GeneRatio indicates the number of differential genes located in the GO/the total number of genes located in the GO, and the larger the GeneRatio, the higher the degree of GO enrichment. (B) The heatmap of the expression of different genes in different samples, where the abscissa is the sample and the ordinate are the gene. The different colors indicate different gene expression levels, and the color from blue to white to red indicates the expression level from low to high. (C) CXCL9 and CXCL10 mRNA expression in the mouse LLC and the human H1975 lung cancer lines were determined 24 h after treatment with ICT and the expression level increased at a dose-dependent manner. (D) Representative immunofluorescence staining images of CD8+ T cells and FoxP3+ regulatory T-cells infiltration in ICT-treated tumor and control group tissue sections. CD8+ T cells were stained red (CD8), FoxP3+ regulatory T-cells were stained green (FOXP3) and nuclei stained blue (DAPI). (E) Quantitative analysis of the ratio of CD8/FOXP3 in the figure (D). (F) T cell marker factors (CD8 and IFN-γ), chemokines (CXCL9 and CXCL10), and inflammatory factors (IL6, IL10 and TNF-α) mRNA expression were determined in ICT-treated tumor tissues by RT-PCR. *P < 0.05, **P < 0.005, ***P < 0.001, ****P < 0.0001 vs control group. (G) Protein expression of PD-L1 was determined in control (C) and ICT-treated tumor tissues by immunoblotting.
In addition, T cell chemokines promote increase of T cell infiltration and improved patient survival [52–54]. Therefore, we asked whether there were greater CD8+ T cells infiltration in ICT-treated tumors using immunofluorescence. As shown in Figure 7D, 7E, we observed a significant increase in the ratio of ICT-treated tumor-infiltrating CD8+ T cells to FoxP3+ regulatory T-cells compared to the control group, indicating that ICT activated the antitumor response of cytotoxic T lymphocytes. To substantiate this result, we further examined the expression levels of T cell marker factors and chemokines in tumor tissues using RT-PCR. We found that whole-tumor gene expression of T cell marker factors (CD8 and IFN-γ) and chemokines (CXCL9 and CXCL10) were significantly enhanced after ICT treatment, while inflammatory factors (IL6, IL10 and TNF-α) were reduced (Figure 7F). In the previously studies, the ICT has been used in the advanced hepatic cell carcinoma treatment via activating CD8+ T cells function in TME by downregulating PD-L1 expression [55]. To examine whether the PD-L1 pathway is blocked by ICT in LLC tumour tissues, we examined the PD-L1 expression by immunoblotting, but found that ICT upregulated the PD-L1 expression (Figure 7G), which may be induced by upregulated IFNG in TME [56]. The different effects of ICT on PD-L1 may be caused by tissular differences. These suggest that ICT triggers the antitumor immunity in lung cancer by regulating inflammatory signaling pathways and increases immune chemokines expression instead of downregulating PD-L1 pathway as in hepatic cell carcinoma.
Pro-apoptosis and anti-inflammatory effects of ICT on lung cancer cells
In view of effective effects of ICT on LLC tumor-bearing mice, we have further verified in vitro. Previously, it was reported ICT potently inhibited growth of prostate cancer PC-3 cells, breast cancer cells and hepatoma HepG2 cells [57, 58]. We decided to examine the effect of various concentrations of ICT on the survival rate of H1975 cells and RAW264.7 cells by CCK-8 analysis. As shown in Figure 8A, the IC50 values of ICT against H1975 cells and RAW264.7 cells were 53.92 μM and 55.45 μM, respectively. Then, according to the IC50 values, we selected the concentration of ICT at 50, 40, 30 μM and 60, 50, 40 μM for subsequent experiments on H1975 cells and RAW 264.7 cells, respectively.
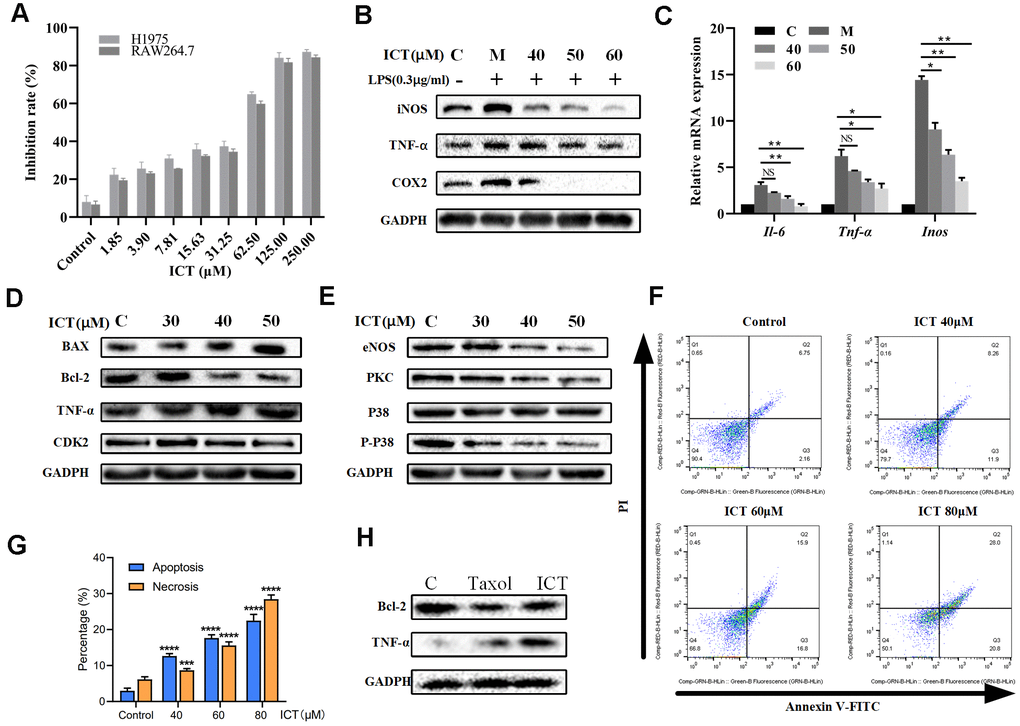
Figure 8. The effect of ICT on lung cancer cell lines. (A) Inhibition rates of ICT on H1975 and RAW264.7 cells. After incubated 24 h, H1975 and RAW264.7 cells viability was measured by CCK-8 assay after treated with control or ICT for 48h. (B) The expression of inflammatory factors (iNOS, TNF-α and COX-2) was measured by western blotting. (C) mRNA expression of inflammatory cytokines (IL-6, TNF-α and iNOS) was determined by RT-PCR. ICT reduced mRNA expression of IL-6, TNF-α and iNOS in LPS-induced RAW264.7 cells. *P < 0.05 vs LPS group, **P < 0.005 vs LPS group. (D) Western blotting showed the expression of classical apoptotic proteins (TNF-α, Bcl-2, Bax, CDK2) in H1975 cells. (E) Western blotting showed that ICT down-regulated the expression of migration proteins (eNOS, PKC, P38) in H1975 cells. (F) Apoptosis in H1975 cells was assessed after 24 h of treatment with ICT (0, 40, 60 and 80μM) by Annexin V-FITC/PI binding and measured by flow cytometry analysis. Numbers indicate the percentage of cells in each quadrant. The number of lower right quadrant and upper right quadrant represent the percentage of apoptosis and necrosis, respectively. (G) Bar plot represents the percentage of apoptosis and necrosis induced by ICT in flow cytometric image (F). *P < 0.05, **P < 0.005, ***P < 0.001, ****P < 0.0001 vs control group. (H) Western blotting showed that the expression of extrinsic protein TNF-α increased and the expression of internal protein Bcl-2 decreased in ICT-treated tumor tissues. *P < 0.05, **P < 0.005, ***P < 0.001, ****P < 0.0001 vs control group. GAPDH was used as the loading control.
The NSCLC-pathway showed that Epimedium and ICT can target tumour-promoting inflammation signaling pathways. As a major component of the leukocyte infiltrate in tumors, macrophages are crucial drivers of tumour-promoting inflammation which impairs CD8+ T cells function [59]. Also, we found that over half of ICT targets (18/32) significantly correlate with abundance of macrophages in TCGA LUAD patients (Figure 2C). Thus, we speculated ICT might promote CD8+ T cell mediated antitumor immunity by suppressing inflammation derived from macrophages. To test this, RAW264.7, a typical macrophage-lineage cell line, was used to validate the anti-inflammatory effects of ICT on macrophages. The results presented in Figure 8B showed that level of TNF-α, iNOS and COX-2 in LPS stimulated culture were significantly higher than that in normal control, which implied the successful establishment of a model of inflammation. When RAW264.7 cells were treated with different concentrations of ICT, the LPS stimulation induced high expression of TNF-α, iNOS and COX-2 were remarkably inhibited. To further characterize the anti-inflammatory effect of ICT in modulating the TNF signaling pathway, we analyzed the expression of TNF-α, IL-6 and iNOS by RT-PCR, and found that ICT suppressed the LPS-induced mRNA expression of TNF-α, IL-6 and iNOS (Figure 8C). These results suggested that ICT mainly regulated the expression of inflammatory factors via the TNF signaling pathway to change the TME.
Moreover, to determine the proliferation-inhibition effect of ICT on H1975 cells, we measured the changes of extrinsic cell apoptosis protein TNF-α, anti-apoptotic protein Bcl-2, pro-apoptotic protein Bax and cell cycle regulatory proteins CDK2 in PI3K-AKT signaling pathway and angiogenic proteins eNOS, PKC and P38 in VEGF signaling pathway under the action of ICT. As shown in Figure 8D, in H1975, the expression of extrinsic cell apoptosis signal TNF-α is increased in a dose-dependent manner after ICT treatment. Note that, for internal signal, we also observed that ICT reduced the levels of Bcl-2 and CDK2 while increased Bax expression (Figure 8D). And eNOS, PKC and P38 also presented a decreasing trend after ICT treatment (Figure 8E). Thus, these results show that ICT triggers apoptosis through both of extrinsic and intrinsic apoptosis pathways in vitro. Furthermore, by flow cytometry analysis, compared with control group, ICT dramatically triggered apoptosis in human lung cancer H1975 cells (Figure 8F, 8G). According to the results of flowcytometry, we counted the percentage of apoptosis (lower right quadrant, Q3) and necrosis (upper right quadrant, Q2) induced by ICT. Compare with the control group, at relative low concentration (40 μM) ICT, ICT induced drastically increase of apoptosis (fold change = 4.2(12.65%/2.99%)), but not necrosis (fold change = 1.4 (8.69%/6.13%)) (Figure 8G). However, at high concentration (80 μM), ICT induced copious necrosis (28.46%) which overtaken apoptosis (22.47%). These results suggested that ICT induced apoptosis at the relative low concentration but necrosis at high concentration. Finally, we have detected the apoptosis related proteins in the ICT treated tumour tissue. Compared with the control, we found that the expression of extrinsic protein TNF-α increased and the expression of internal protein Bcl-2 decreased in ICT-treated tumor tissues (Figure 8H). To sum up, the experiments in vitro confirmed that ICT could exert the effect of anti-inflammation and pro-apoptosis by regulating the expression of critical proteins in our integrated NSCLC-pathway, achieving the anti-NSCLC effect.
Discussion
With the development of the field of tumor immunotherapy, immunotherapy that enhances the ability of cytotoxic T lymphocytes by targeting TME to kill tumor cells has been confirmed in a variety of human malignancies [60–62]. However, NSCLC patients are resistant to many targeted therapeutic drugs and have poor prognosis. Given the development of multi-target natural products that can target TME to regulate tumor biological networks and pathways, enhancing antitumoral responses of CD8+ T cells and making the efficacy more durable. Therefore, it is necessary to conduct additional studies for natural products to improve antitumor immune effect and optimize patient selection. In this paper, we used systems pharmacology approach to reveal the action mechanism of polypharmacology molecules of Epimedium targeting TME for NSCLC.
With the aid of favorable pharmacokinetic characteristics, 16 active ingredients of Epimedium were obtained. Among them, it was reported that some vital compounds of Epimedium such as ICT, kaempferol and sitosterol have antitumor activity [63–65]. And 89 potential targets were predicted by bioinformatic algorithms. Association analysis of diseases and target function enrichment analysis together displayed that the targets were closely related to neoplasms and processes of inflammation and immune.
In conclusion, the systems pharmacology models integrated the multiple methods including the active compounds screening, target predicting, network pharmacology analysis and onco-immune interacting to predict the natural products that target TME. Applying the systems pharmacology, we successfully revealed the ICT enhanced the infiltration of CD8+ T cells in TME via multiple pathways, which shown the accuracy and reliability of systems pharmacology. According to these results, we speculate ICT could be used to treat the NSCLC patients with low infiltration of CD8+ T cells, which provided the basis for the sponsors to set inclusion/exclusion criteria in future trial design. In addition, we consider that Epimedium and ICT might sensitize the cancers to anti-PD-1/PD-L1 or anti-CTLA4 via increasing the infiltration of CD8+ T cells, since immune checkpoint blockade therapies often fail in the patients with poor T cell infiltration. In the future, we believe that systems pharmacology is a potential framework for rationally designing combination therapies with immune checkpoint blockade through screening the lead compounds that target pathways determining the resistance of the immune checkpoint blockade.
Materials and Methods
Pharmacokinetic screening
All chemical components of Epimedium were retrieved from the literature and uploaded to the TCMSP (http://lsp.nwu.edu.cn/tcmsp.php) [66]. As a result, 130 ingredients were got for the Epimedium, which molecular structures were saved as mol2 format for subsequent analysis.
Based on the compounds obtained above, we used the in silico to screen potential active molecules with favorable pharmacokinetics properties that were potent anti-NSCLC. ADME-based screening criteria, i.e. OB ≥ 30%; DL ≥ 0.18; HL ≥ 4 h. The filtered active compounds must simultaneously satisfy three conditions.
Target fishing
In this work, we used SEA (http://sea.bkslab.org/) [67], WES algorithm [68] and SysDT [69] to understand the nature of the compound-target interaction. Among them, the WES computational model was introduced to detect drug direct targets of the active ingredients based on a large-scale of drug target relationships [68]. SysDT model, based on random forest (RF) and support vector machine (SVM), was utilized to explore potential targets of drugs, which could comprehensively determine the distribution of compound-target interaction. The compound-target interactions with SVM score ≥ 0.8 and the RF score ≥ 0.7 were selected for further research [69]. Then, we mapped targets to the Uniprot (http://www.uniprot.org), unifying their names and organisms. Normalized compound targets were mapped to the CTD database (http://ctdbase.org/) [70], Therapeutic Target Database (TTD, http://database.idrb.cqu.edu.cn/TTD/) [71], and Pharmacogenomics Knowledgebase (PharmGKB, https://www.pharmgkb.org/) [72] to obtain their corresponding diseases and screen out potential targets related to NSCLC.
Target-associated diseases analysis
To illustrate the relationship between predicted potential targets and treatment of NSCLC, we mapped these targets to the CTD to obtain their related diseases, and classified these diseases into 12 categories based on their direct or indirect effects. Then, we constructed a radar chart to analyze the degree of association between the target and the disease.
GOBP analysis
In order to further probe the meaningful functional annotation of the potential targets, we performed the GO enrichment analysis by linking the targets to ClueGO (Cytoscape plugin). Only GO terms with P-value ≤ 0.05 are selected. The Enrichment Map [73], a Cytoscape plug-in, was implemented as a freely available and open source for visualization and analysis software. We applied the GSEA (Gene-Set Enrichment Analysis) to analyze enrichment significance of gene-sets, and then used to find enriched GO gene-sets. Only gene-sets passing conservative significance thresholds (P-value < 0.005, False Discovery Rate (FDR) < 1%) were screened out for display in the Enrichment Map, illuminating the biology processes of the lists of genes.
Correlation between targets and clinical features in LUAD patients from TCGA
We correlated the targets to the LUAD clinical features including ten immune phenotypes, disease stage and overall survival time. The RNA-seq data and information of clinical stage and overall survival time of LUAD in TCGA were download by R TCGABiolinks package. The ten antitumor immunity-related phenotypes involve the proportion of major classes of immune cells (leukocyte, T cells CD8, activated NK cells, activated dendritic cells, Tregs, macrophages M2), TCR richness and the TIL regional fraction on tissue image, which are implicated in regulating tumor responses to host immunity. These data were obtained from the curated methods that were described in llya Shmulevich et al. [38]. We correlated the mRNA level (log2(TMP+1)) of targets with these phenotypes using Pearson's correlation coefficient (PCC). The p-values of PCC were adjusted by Benjamini–Hochberg methods. To correlate targets expression with overall survival, we evaluated the target set expression level for each of LUAD patient by PLAGE (Pathway Level Analysis of Gene Expression) approach, which translates gene expression levels into 'pathway activity' levels using singular value decomposition [74]. Kaplan-Meier plots summarized the results from correlation analysis between PLAGE expression level and patient survival. Patients were divided based on level of PLAGE activity into one of the two groups "low activity" (under cut off) or "high activity" (over cut off). The cut off was set to minimize the p-value of log rank test and guarantee the fraction of "low activity" or "high activity" group that should never go below 20%. To correlate targets with clinical staging, we normalized targets expression by z-score transformation and averaged the normalized expression by tumour stages (stages I–IV).
Differentially expression gene analysis
Publicly available gene expression profile of TCGA-LUAD and TCGA-LUSC were downloaded from the Genomic Data Commons (GDC) Data Portal and pre-processed via the TCGAbiolinks R package [40]. Then, the DEGs of LUAD and LUSC were filtered using the Limma pipeline in the TCGAbiolinks with the |log2FC| cutoff of 1 and FDR cutoff of 0.01. Finally, we plotted a Venn diagram and analyzed overlap ratio between the potential targets of the Epimedium active components and the DEGs of LUAD and LUSC. Furthermore, we further calculated pairwise correlation of 89 potential genes of Epimedium active components by the Pearson method and constructed a gene-gene correlation matrix and heatmap. Subsequently, we randomly acquired 53 genes and calculated the correlation between them. Meanwhile, we mapped the distribution diagram of random intergenic correlations and Epimedium intergenic correlation, and calculated Wasserstein distance between the two distribution diagrams.
Network construction
To directly reflect the relationship between compounds, targets, pathways and diseases, we constructed C-T network and T-P network by Cytoscape 3.7.0 [75], which is a popular bioinformatics package for biological network visualization and data integration. In the generated network, the nodes represent compounds, targets and pathways, and the interactions between them were represented by edges. The topological properties of these networks were analyzed using the Network Analysis plugin and CentiScaPe 1.2 of Cytoscape 3.7.0.
In addition, we collected genes of the Epimedium active components in the pathways and the diseases, and counted the number of common genes between them. The pathways and diseases with more than 3 overlapping genes were connected, and a chord diagram of P-D interactions was plotted by using the Circos (Circlize package in R v3.4) [76].
Pathway construction
To further explore how the active compounds exert their pharmacokinetic effects by modulating targets in pathways, an incorporated “NSCLC-pathway” was assembled based on the latest information on NSCLC pathology. In short, the targets were firstly mapped to DAVID and the pathways associated with NSCLC were obtained and screened from the KEGG database (http://www.genome.jp/kegg/) [77], and which were then manually integrated into the NSCLC-pathway based on pathological and clinical data.
Reagents
ICT was purchased from Shanghai Yuanye Bio-Technology Co., Ltd (Shang Hai, China). A stock solution of ICT (100mM) was dissolved in DMSO for in vitro experiments, stored at 4° C and further diluted to appropriate concentrations before use. The final concentration of DMSO was < 0.1% in culture and insured there was no effect on cell viability. Dulbecco’s modified Eagle’s medium (DMEM) and RPMI-1640 (Gibco BRL, USA), Dimethyl sulfoxide (DMSO) (Sigma, USA), CCK-8 (BestBio, Shanghai, China), fetal bovine serum (FBS) (Gibco BRL, USA), Qproteome™ Mammalian Protein Prep Kit (Qiagen, Germany), BCA Protein Assay Kit (Beyotime, Shanghai, China), FITC Annexin V apoptosis kit (BD Pharmingen, Franklin Lakes, NJ, USA), Trizol reagent (Takara BioCo., China), MuLV reverse transcriptase and Oligo-dT primers (Takara BioCo., China), SYBR green PCR Master Mix (Takara BioCo., China) were obtained as indicated. We used the following primary antibodies: GAPDH (1:1000); Bcl-2 (1:1000); Bax; GSK3β(1:8000); CDK2 (1:2000); iNOS (1:1000); eNOS (1:1000); PKC(1:50000); p38 (1:1000); p-p38 (1:1000); COX2 (1:1000); TNF-α (1:1000); and Goat Anti-Mouse IgG H&L (HRP) were obtained from Abcam.
Animal and drug treatment
Female C57BL/6 mice (5-6 weeks, 18–22g) were obtained from the Comparative Medicine Centre of Yangzhou University (Yangzhou, China). They were maintained under specific pathogen-free conditions at the Institute of Laboratory Animals, Jiangsu Kanion Pharmaceutical, Co., Ltd. and used under protocols approved by the respective Institute of pharmacology and toxicology institutional review board (IRB), IACUC No. (2019040401).
The animals were maintained in a pathogen-free environment at 25 ± 1° C and humidity of 55 ± 5% for 1 week, 12 h light/dark cycle and free access to food and water under a standard specific pathogen free (SPF) condition. LLC cells suspended in phosphate-buffered saline (PBS). Each mouse was subcutaneously inoculated with approximately 5×105 LLC cells to the left axilla. On the third day (after inoculation), mice were randomly divided into six groups: two groups of control (n = 6/group, intraperitoneal injection of diluent); two groups of positive control (n = 6/group, daily intraperitoneal administration of Taxol, 15mg/kg, Yuan ye, Shanghai) and two groups of ICT (n = 6/group, daily intragastric administration of ICT, 15mg/kg, Yuan ye, Shanghai). Among them, three groups (control group, positive control group and ICT group) were used for the analysis of tumor volume and weight. When the tumor reaches 5×5 mm, the size of the tumor is measured by digital caliper and body weight recorded once every two days until the tumor is larger than 20 mm×20 mm, then the mouse is killed and recorded as death. Tumor volume was measured along the longest orthogonal axes and calculated as volume = (length×width2)/2, where width was the shortest measurement. On the 21 days, the mice were sacrificed by cervical dislocation, and the tumor was taken out for weight, observation and analysis. While the other three groups (control group and medication group) were used for survival analysis.
Transcriptome sequencing
Total RNA was extracted using Trizol reagent (Invitrogen, CA, USA) following the manufacturer's procedure. The total RNA quantity and purity were analysis of Bioanalyzer 2100 and RNA 1000 Nano LabChip Kit (Agilent, CA, USA) with RIN number > 7.0. Poly(A) RNA is purified from total RNA (5 μg) using poly-T oligo-attached magnetic beads using two rounds of purification. Then the cleaved RNA fragments were reverse-transcribed to create the final cDNA library in accordance with the protocol for the mRNASeq sample preparation kit (Illumina, San Diego, USA). And then we performed the paired-end sequencing on an Illumina Novaseq™ 6000 at the (LC Sciences, USA) following the vendor’s recommended protocol.
We aligned reads of sample ICT-treated tumors and sample control to the UCSC (http://genome.ucsc.edu/) homo sapiens reference genome using HISAT package, which initially remove a portion of the reads based on quality information accompanying each read and then maps the reads to the reference genome. The mapped reads of each sample were assembled using StringTie. Then, all transcriptomes from Samples were merged to reconstruct a comprehensive transcriptome using perl scripts. After the final transcriptome was generated, StringTie and edgeR was used to estimate the expression levels of all transcripts. StringTie was used to perform expression level for mRNAs by calculating FPKM (Fragments Per Kilobase of exon model per Million mapped reads). The differentially expressed mRNAs and genes were detect threshold of |log2(fold change)| >1 and with statistical significance (q-value < 0.05) by R LPEseq package which is developed to implement the analysis of differential expression with a small number of or non-replicated samples [47]. As a result, we identified 2,375 differentially expressed genes (DEGs), including 1779 upregulated and 596 downregulated DEGs in ICT-treated tumor. And the sequencing data were uploaded to our website of Traditional Chinese Medicine Systems Pharmacology Database (TCMSP, https://tcmspw.com/). Finally, the GO enrichment analysis chart of differentially expressed genes was created by clusterProfiler.
Immunofluorescence
Analysis of CD8+ T cells infiltration in ICT-treated tumor tissues by immunofluorescence. Firstly, the tumor tissues were formaldehyde-fixed and paraffin-embedded following standard procedures. Then, paraffin tissue sections of 4 μm thickness were prepared using a paraffin slicing machine (LEICA RM2126) and the sections were roasted overnight at 37° C, dewaxed and rehydrated. They were then sealed with goat serum for 30 min, washed 3-4 times in PBS, and incubated overnight at 4° C with CD8 Polyclonal Antibody (PA5-88265, Thermo Scientific) and FOXP3 Monoclonal Antibody (11-5773-82, Thermo Scientific). After washing 3-4 times with PBS, the Goat anti-Rabbit IgG (H+L) Highly Cross-Adsorbed Secondary Antibody (A-21245, Thermo Scientific) was added dropwise, incubated at 37° C for 1 h, stained with DAPI for 5-10 minutes, washed 3-4 times with PBS, and air-dried. Observed under a confocal microscopy (Nikon A1R).
Cells and cell culture
Human NSCLC H1975, RAW264.7 and Lewis lung carcinoma (LLC) cells were obtained from Chinese Academy of Sciences Shanghai cell bank. H1975 cells were cultured in RPMI-1640 media complemented with 10% fetal bovine serum (FBS). RAW264.7 and LLC cells were maintained in DMEM with 10% FBS. The all experiments cells were maintained in a humidified atmosphere of 5% CO2 at 37° C. The culture medium was substituted fresh medium every 2-3 days and amplified to new culture when the cells reached approximately 80-90% confluency.
Cell viability assays
The effects of ICT on H1975 cell viability were determined by measuring the metabolic activity (CCK-8 assay). H1975 cells in the logarithmic phase were seeded in 96-well-plates at a density of 8000 per well and cultured for 24 h. After incubated 24 h, cells were treated with various concentrations of ICT (250, 125, 62.5, 31.25, 15.63, 7.81, 3.9, 1.85, 0 μM) for 48 h. RAW264.7 and H1975 cells have the same experimental protocol. Then, 10 μl of CCK-8 assay was added to each well for 4 h incubation at 37° C and 5% CO2. And the absorbance was detected by a microplate reader (Molecular Devices, California, USA) at 450 nm. Cells treated with medium containing 1% DMSO was regarded as negative control. Three reduplicate wells were used for each treatment and the experiments were performed three times.
Inflammation model
RAW264.7 cells (2×106) were cultured in 150 mm culture dish 24 h and treated with the various concentrations of ICT for 2 h. It was then incubated with 0.3 μg/mL LPS for 18 h. The cells were collected at the end of the culture for western blotting assays, which were used as a detection of inflammatory mediators.
Western blotting
The expression of protein was analyzed using western blotting. Firstly, H1975 or RAW264.7 cells (2×106) were seeded in 150 mm culture dish overnight and treated with the various concentrations of ICT for 24 h. The cells were harvested and scraped, collected by centrifugation and lysed in Qproteome™ Mammalian Protein Prep Kit. The protein concentration was determined using BCA Protein Assay Kit. Secondly, equalized amounts of proteins from each sample separated by SDS-PAGE and transferred onto a PVDF (Millipore, Bedford, MA, USA) membrane. The membrane was washed three times with TBST buffer and blocked with 5% skim milk for 2 h. After washing three times, the membrane incubated with primary antibodies against Bcl-2, Bax, CDK2, eNOS, PKC, p38, p-p38, iNOS, COX2, TNF-α, and GAPDH at 4° C overnight. Finally, the membrane was washed three times again and incubated with a horseradish peroxidase conjugated secondary goat anti-mouse IgG H&L (HRP) for 1 h at room temperature and then washed twice in TBST, once with TBS, afterwards, detected by using the ChemiDoc™ XRS+ Imaging System (Bio-Rad) and labelling were visualized by ImageLab sofware (Bio-Rad). GAPDH was used as the loading control. All of the western blotting experiments were performed at least three times.
Flow cytometry assays
Cell apoptosis was determined using a FITC Annexin V apoptosis kit according to the manufacturer’s instructions. Following treatment with various concentrations (80, 60, 40 μM) of ICT for 24 h, the cell suspension was prepared using trypsin and centrifuged at 1,000 rpm for 3 min then rinsed with ice-cold PBS. Cells were then resuspended in binding buffer (10 mmol/L HEPES, pH 7.4, 140 mmol/L NaCl and 2.5 mmol/L CaCl2) at a concentration of 1×105 cells/ml. Cells were stained with annexin V-FITC and propidium (PI) for 15 min in the dark before analysis by a flow cytometer (Beckman Coulter Inc, Miami, FL, USA). Annexin V+/PI- and V+/PI+ cells were considered as apoptotic cells.
Quantitative real-time PCR
Gene expression of TNF-α, iNOS, IL-6, IL-10, IFN-γ, CD8, CXCL9, CXCL10 was detected by RT-PCR. Total RNA of cells and tissues was extracted using Trizol reagent according to the manufacturer's instructions. The primer sequences of the selected genes were designed by the Primer3 software (http://frodo.wi.mit.edu/cgi-bin/primer3/primer3_www.cgi) and listed in below: TNF-α, 5′-GCGACGTGGAACTGGCAGAAG-3′ (forward) and 5′-TCCATGCCGTTGGCCAGGAGG-3′ (reverse); iNOS, 5′-TGGAGTCACAGAAGGAGTGGCTAAC-3′ (forward) and 5′-TCTGAC CACAGTGAGGAATGTCCAC-3′ (reverse); IL-6, 5′-TGGAGTCACAGAAGGAGTGGCTAAG-3′ (forward) and 5′-TCTGACCACAGTGAGGAATGTCCAC-3′ (reverse); IL-10, 5′-ACGGCGCTGTCATCGATT-3′ (forward), 5′-GGCATTCTTCACCTGCTCCA-3′ (reverse); IFN-γ, 5′-ATGAACGCTACACACTGCATC-3′ (forward), 5′-CCATCCTTTTGCCAGTTCCTC-3′ (reverse); CD8, 5′-GTCTATATGGCTTCATCCCACA-3′ (forward), 5′-GTTCAGGGTGAGAACGTACTTA-3′ (reverse); CXCL9, 5′-GGCTCGCAGGGATGATTTCAAGAPDH-3′ (forward), 5′-CCAAGTGCTGCCGTCATTTTC-3′ (reverse); CXCL10, 5′-GGAGTTCGAGGAACCCTAGTG-3′ (forward), 5′-GGGATTTGTAGTGGATCGTGC-3′ (reverse). Reverse transcription was undertaken using MuLV reverse transcriptase and Oligo-dT primers. RT-PCR analysis was performed with the SYBR green PCR Master Mix. The relative expression levels of the genes were quantified as cycle time (Ct) values normalized with GAPDH of the same sample.
Statistical analysis
All data were expressed as the mean ± SD and represent the results of three separate experiments performed in triplicate unless otherwise stated. Student’s t-test was used to evaluate the difference between two groups, and a one-way ANOVA with a post hoc test was used for comparison among three or more groups. In the figures, the standard symbols were used: *P < 0.05, **P < 0.005, ***P < 0.001, ****P < 0.0001 vs control group. P < 0.05 was considered to be statistically significant.
Supplementary Materials
Author Contributions
Conceived and designed: Chao Huang, Zhihua Li, and Chunli Zheng; conducted experiments: Zhihua Li, Jinglin Zhu, Chao Huang, Xia Zhang, Yuanyuan Hao, Ruijie Yang, Ruifei Huang; performed data analysis: Chao Huang, Xuetong Chen, Jun Zhou, Zhenzhong Wang; wrote or contributed to the writing of the manuscript: Zhihua Li, Chao Huang; contributed to manuscript revision: Yonghua Wang, Chunli Zheng, Wei Xiao.
Acknowledgments
We thank Dr. Haojie Yang for proofreading the language. The author would like to thank Jiangsu Kanion Pharmaceutical, Co., Ltd. for its experimental support in this study.
Conflicts of Interest
The authors declare that they have no conflicts of interest.
Funding
This work was supported by grants from Key Laboratory of Resource Biology and Biotechnology in Western China, National Natural Science Foundation of China (Grant No. 81803960 and U1603285) and the 64th batch of postdoctoral general programs in China (Grant No. 2018M643723). It also was supported by the Key Project of Natural Science Fund of Science and Technology Department of Shaanxi Province (Grant No. 2018JZ3007).
References
- 1. Rotow J, Bivona TG. Understanding and targeting resistance mechanisms in NSCLC. Nat Rev Cancer. 2017; 17:637–58. https://doi.org/10.1038/nrc.2017.84 [PubMed]
- 2. Anagnostou VK, Brahmer JR. Cancer immunotherapy: a future paradigm shift in the treatment of non-small cell lung cancer. Clin Cancer Res. 2015; 21:976–84. https://doi.org/10.1158/1078-0432.CCR-14-1187 [PubMed]
- 3. Rizvi NA, Hellmann MD, Snyder A, Kvistborg P, Makarov V, Havel JJ, Lee W, Yuan J, Wong P, Ho TS, Miller ML, Rekhtman N, Moreira AL, et al. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science. 2015; 348:124–28. https://doi.org/10.1126/science.aaa1348 [PubMed]
- 4. Lambrechts D, Wauters E, Boeckx B, Aibar S, Nittner D, Burton O, Bassez A, Decaluwé H, Pircher A, Van den Eynde K, Weynand B, Verbeken E, De Leyn P, et al. Phenotype molding of stromal cells in the lung tumor microenvironment. Nat Med. 2018; 24:1277–89. https://doi.org/10.1038/s41591-018-0096-5 [PubMed]
- 5. Tang H, Qiao J, Fu YX. Immunotherapy and tumor microenvironment. Cancer Lett. 2016; 370:85–90. https://doi.org/10.1016/j.canlet.2015.10.009 [PubMed]
- 6. Sharma P, Allison JP. Immune checkpoint targeting in cancer therapy: toward combination strategies with curative potential. Cell. 2015; 161:205–14. https://doi.org/10.1016/j.cell.2015.03.030 [PubMed]
- 7. Taniguchi K, Karin M. NF-κB, inflammation, immunity and cancer: coming of age. Nat Rev Immunol. 2018; 18:309–24. https://doi.org/10.1038/nri.2017.142 [PubMed]
- 8. Radwan FF, Perez JM, Haque A. Apoptotic and immune restoration effects of ganoderic acids define a new prospective for complementary treatment of cancer. J Clin Cell Immunol. 2011; S3:4. https://doi.org/10.4172/2155-9899.S3-004 [PubMed]
- 9. Kim BH, Na KM, Oh I, Song IH, Lee YS, Shin J, Kim TY. Kurarinone regulates immune responses through regulation of the JAK/STAT and TCR-mediated signaling pathways. Biochem Pharmacol. 2013; 85:1134–44. https://doi.org/10.1016/j.bcp.2013.01.005 [PubMed]
- 10. Pan G, Xie Z, Huang S, Tai Y, Cai Q, Jiang W, Sun J, Yuan Y. Immune-enhancing effects of polysaccharides extracted from Lilium lancifolium Thunb. Int Immunopharmacol. 2017; 52:119–26. https://doi.org/10.1016/j.intimp.2017.08.030 [PubMed]
- 11. Chen Y, Lin Y, Li Y, Li C. Total flavonoids of Hedyotis diffusa willd inhibit inflammatory responses in LPS-activated macrophages via suppression of the NF-κB and MAPK signaling pathways. Exp Ther Med. 2016; 11:1116–22. https://doi.org/10.3892/etm.2015.2963 [PubMed]
- 12. Pan J, Yang C, Jiang Z, Huang J. Trametes robiniophila Murr: a traditional Chinese medicine with potent anti-tumor effects. Cancer Manag Res. 2019; 11:1541–49. https://doi.org/10.2147/CMAR.S193174 [PubMed]
- 13. Apaya MK, Chang MT, Shyur LF. Phytomedicine polypharmacology: cancer therapy through modulating the tumor microenvironment and oxylipin dynamics. Pharmacol Ther. 2016; 162:58–68. https://doi.org/10.1016/j.pharmthera.2016.03.001 [PubMed]
- 14. Guo Q, Li J, Lin H. Effect and Molecular Mechanisms of Traditional Chinese Medicine on Regulating Tumor Immunosuppressive Microenvironment. BioMed Res Int. 2015; 2015:261620. https://doi.org/10.1155/2015/261620 [PubMed]
- 15. Huang Y, Cai T, Xia X, Cai Y, Wu XY. Research advances in the intervention of inflammation and cancer by active ingredients of traditional Chinese medicine. J Pharm Pharm Sci. 2016; 19:114–26. https://doi.org/10.18433/J3SG7K [PubMed]
- 16. Wang P, Yang HL, Yang YJ, Wang L, Lee SC. Overcome cancer cell drug resistance using natural products. Evid Based Complement Alternat Med. 2015; 2015:767136. https://doi.org/10.1155/2015/767136 [PubMed]
- 17. Safe S, Kasiappan R. Natural products as mechanism-based anticancer agents: Sp transcription factors as targets. Phytother Res. 2016; 30:1723–32. https://doi.org/10.1002/ptr.5669 [PubMed]
- 18. Demir S, Aliyazicioglu Y, Turan I, Misir S, Mentese A, Yaman SO, Akbulut K, Kilinc K, Deger O. Antiproliferative and proapoptotic activity of Turkish propolis on human lung cancer cell line. Nutr Cancer. 2016; 68:165–72. https://doi.org/10.1080/01635581.2016.1115096 [PubMed]
- 19. Sheng MY, Wang LJ, Tian XJ. Karyomorphology of eighteen species of genus Epimedium (Berberidaceae) and its phylogenetic implications. Genet Resour Crop Evol. 2010; 57:1165–76. https://doi.org/10.1007/s10722-010-9556-6
- 20. Miura Y, Oyama M, Iguchi K, Ito T, Baba M, Shikama Y, Usui S, Hirano K, Iinuma M, Mikamo H. Anti-androgenic activity of icarisid II from epimedium herb in prostate cancer LNCaP cells. J Nutr Sci Vitaminol (Tokyo). 2015; 61:201–04. https://doi.org/10.3177/jnsv.61.201 [PubMed]
- 21. Yap SP, Shen P, Butler MS, Gong Y, Loy CJ, Yong EL. New estrogenic prenylflavone from epimedium brevicornum inhibits the growth of breast cancer cells. Planta Med. 2005; 71:114–19. https://doi.org/10.1055/s-2005-837776 [PubMed]
- 22. Huang C, Zheng C, Li Y, Wang Y, Lu A, Yang L. Systems pharmacology in drug discovery and therapeutic insight for herbal medicines. Brief Bioinform. 2014; 15:710–33. https://doi.org/10.1093/bib/bbt035 [PubMed]
- 23. Zhu S, Wang Z, Li Z, Peng H, Luo Y, Deng M, Li R, Dai C, Xu Y, Liu S, Zhang G. Icaritin suppresses multiple myeloma, by inhibiting IL-6/JAK2/STAT3. Oncotarget. 2015; 6:10460–72. https://doi.org/10.18632/oncotarget.3399 [PubMed]
- 24. Sun F, Indran IR, Zhang ZW, Tan MH, Li Y, Lim ZL, Hua R, Yang C, Soon FF, Li J, Xu HE, Cheung E, Yong EL. A novel prostate cancer therapeutic strategy using icaritin-activated arylhydrocarbon-receptor to co-target androgen receptor and its splice variants. Carcinogenesis. 2015; 36:757–68. https://doi.org/10.1093/carcin/bgv040 [PubMed]
- 25. Pan C, Fujiwara Y, Horlad H, Shiraishi D, Iriki T, Tsuboki J, Ikeda T, Komohara Y. Flavonoid compounds contained in Epimedii herba inhibit tumor progression by suppressing STAT3 activation in the tumor microenvironment. Front Pharmacol. 2020; 11:262. https://doi.org/10.3389/fphar.2020.00262 [PubMed]
- 26. Cho HJ, Park JH. Kaempferol induces cell cycle arrest in HT-29 human colon cancer cells. J Cancer Prev. 2013; 18:257–63. https://doi.org/10.15430/jcp.2013.18.3.257 [PubMed]
- 27. Shin EJ, Choi HK, Sung MJ, Park JH, Chung MY, Chung S, Hwang JT. Anti-tumour effects of beta-sitosterol are mediated by AMPK/PTEN/HSP90 axis in AGS human gastric adenocarcinoma cells and xenograft mouse models. Biochem Pharmacol. 2018; 152:60–70. https://doi.org/10.1016/j.bcp.2018.03.010 [PubMed]
- 28. Sundstrøm T, Varughese J, Prestegarden L, Azuaje F, Røsland GV, Skaftnesmo KO, Ingham ES, Even L, Tam S, Tepper C, Petersen K, Ferrara KW, Tronstad KJ, et al. BMET-34 Drug repurposing discovers beta-sitosterol as an effective therapeutic agent against melanoma brain metastases in vivo. Neuro-oncol. 2015 (Suppl 5); 17:v52. https://doi.org/10.1093/neuonc/nov208.34
- 29. She LC, Liu CM, Chen CT, Li HT, Li WJ, Chen CY. The anti-cancer and anti-metastasis effects of phytochemical constituents from Leucaena leucocephala. Biomed Res. 2017; 28:2893–97.
- 30. Lee KW, Lee HJ, Cho HY, Kim YJ. Role of the conjugated linoleic acid in the prevention of cancer. Crit Rev Food Sci Nutr. 2005; 45:135–44. https://doi.org/10.1080/10408690490911800 [PubMed]
- 31. Rached W, Zeghada FZ, Bennaceur M, Barros L, Calhelha RC, Heleno S, Alves MJ, Carvalho AM, Marouf A, Ferreira IC. Phytochemical analysis and assessment of antioxidant, antimicrobial, anti-inflammatory and cytotoxic properties of Tetraclinis articulata (Vahl) Masters leaves. Ind Crops Prod. 2018; 112:460–66. https://doi.org/10.1016/j.indcrop.2017.12.037
- 32. Slattery ML, Sweeney C, Herrick J, Wolff R, Baumgartner K, Giuliano A, Byers T. ESR1, AR, body size, and breast cancer risk in Hispanic and non-Hispanic white women living in the Southwestern United States. Breast Cancer Res Treat. 2007; 105:327–35. https://doi.org/10.1007/s10549-006-9453-z [PubMed]
- 33. Pesek M, Kopeckova M, Benesova L, Meszarosova A, Mukensnabl P, Bruha F, Minarik M. Clinical significance of hypermethylation status in NSCLC: evaluation of a 30-gene panel in patients with advanced disease. Anticancer Res. 2011; 31:4647–52. [PubMed]
- 34. Thiebaut C, Konan HP, Guerquin MJ, Chesnel A, Livera G, Le Romancer M, Dumond H. The role of ERα36 in development and tumor Malignancy. Int J Mol Sci. 2020; 21:4116. https://doi.org/10.3390/ijms21114116 [PubMed]
- 35. Li J, Kong X, Zhang J, Luo Q, Li X, Fang L. MiRNA-26b inhibits proliferation by targeting PTGS2 in breast cancer. Cancer Cell Int. 2013; 13:7. https://doi.org/10.1186/1475-2867-13-7 [PubMed]
- 36. Markosyan N, Li J, Sun YH, Richman LP, Lin JH, Yan F, Quinones L, Sela Y, Yamazoe T, Gordon N, Tobias JW, Byrne KT, Rech AJ, et al. Tumor cell-intrinsic EPHA2 suppresses anti-tumor immunity by regulating PTGS2 (COX-2). J Clin Invest. 2019; 129:3594–609. https://doi.org/10.1172/JCI127755 [PubMed]
- 37. Xu L, Stevens J, Hilton MB, Seaman S, Conrads TP, Veenstra TD, Logsdon D, Morris H, Swing DA, Patel NL, Kalen J, Haines DC, Zudaire E, St Croix B. COX-2 inhibition potentiates antiangiogenic cancer therapy and prevents metastasis in preclinical models. Sci Transl Med. 2014; 6:242ra84. https://doi.org/10.1126/scitranslmed.3008455 [PubMed]
- 38. Thorsson V, Gibbs DL, Brown SD, Wolf D, Bortone DS, Ou Yang TH, Porta-Pardo E, Gao GF, Plaisier CL, Eddy JA, Ziv E, Culhane AC, Paull EO, et al, and Cancer Genome Atlas Research Network. The immune landscape of cancer. Immunity. 2018; 48:812–30.e14. https://doi.org/10.1016/j.immuni.2018.03.023 [PubMed]
- 39. Liu YJ, Fan HB, Jin Y, Ren CG, Jia XE, Wang L, Chen Y, Dong M, Zhu KY, Dong ZW, Ye BX, Zhong Z, Deng M, et al. Cannabinoid receptor 2 suppresses leukocyte inflammatory migration by modulating the JNK/c-Jun/Alox5 pathway. J Biol Chem. 2013; 288:13551–62. https://doi.org/10.1074/jbc.M113.453811 [PubMed]
- 40. Colaprico A, Silva TC, Olsen C, Garofano L, Cava C, Garolini D, Sabedot TS, Malta TM, Pagnotta SM, Castiglioni I, Ceccarelli M, Bontempi G, Noushmehr H. TCGAbiolinks: an R/Bioconductor package for integrative analysis of TCGA data. Nucleic Acids Res. 2016; 44:e71. https://doi.org/10.1093/nar/gkv1507 [PubMed]
- 41. Haura EB, Livingston S, Coppola D. Autocrine interleukin-6/interleukin-6 receptor stimulation in non-small-cell lung cancer. Clin Lung Cancer. 2006; 7:273–75. https://doi.org/10.3816/CLC.2006.n.006 [PubMed]
- 42. Zeng J, Liu D, Qiu Z, Huang Y, Chen B, Wang L, Xu H, Huang N, Liu L, Li W. GSK3β overexpression indicates poor prognosis and its inhibition reduces cell proliferation and survival of non-small cell lung cancer cells. PLoS One. 2014; 9:e91231. https://doi.org/10.1371/journal.pone.0091231 [PubMed]
- 43. Yang TY, Chang GC, Chen KC, Hung HW, Hsu KH, Sheu GT, Hsu SL. Sustained activation of ERK and Cdk2/cyclin-A signaling pathway by pemetrexed leading to S-phase arrest and apoptosis in human non-small cell lung cancer A549 cells. Eur J Pharmacol. 2011; 663:17–26. https://doi.org/10.1016/j.ejphar.2011.04.057 [PubMed]
- 44. Annibaldi A, Meier P. Checkpoints in TNF-induced cell death: implications in inflammation and cancer. Trends Mol Med. 2018; 24:49–65. https://doi.org/10.1016/j.molmed.2017.11.002 [PubMed]
- 45. Shintani Y, Fujiwara A, Kimura T, Kawamura T, Funaki S, Minami M, Okumura M. IL-6 secreted from cancer-associated fibroblasts mediates chemoresistance in NSCLC by increasing epithelial-mesenchymal transition signaling. J Thorac Oncol. 2016; 11:1482–92. https://doi.org/10.1016/j.jtho.2016.05.025 [PubMed]
- 46. Yaren A, Oztop I, Kargi A, Ulukus C, Onen A, Sanli A, Binicier O, Yilmaz U, Alakavuklar M. Bax, bcl-2 and c-kit expression in non-small-cell lung cancer and their effects on prognosis. Int J Clin Pract. 2006; 60:675–82. https://doi.org/10.1111/j.1368-5031.2006.00742.x [PubMed]
- 47. Tong JS, Zhang QH, Huang X, Fu XQ, Qi ST, Wang YP, Hou Y, Sheng J, Sun QY. Icaritin causes sustained ERK1/2 activation and induces apoptosis in human endometrial cancer cells. PLoS One. 2011; 6:e16781. https://doi.org/10.1371/journal.pone.0016781 [PubMed]
- 48. Wang Y, Chen Y, Cheng X, Zhang K, Wang H, Liu B, Wang J. Design, synthesis and biological evaluation of pyrimidine derivatives as novel CDK2 inhibitors that induce apoptosis and cell cycle arrest in breast cancer cells. Bioorg Med Chem. 2018; 26:3491–501. https://doi.org/10.1016/j.bmc.2018.05.024 [PubMed]
- 49. Zhang W, Huai Y, Miao Z, Chen C, Shahen M, Rahman SU, Alagawany M, El-Hack ME, Zhao H, Qian A. Systems pharmacology approach to investigate the molecular mechanisms of herb rhodiola rosea L. Radix. Drug Dev Ind Pharm. 2019; 45:456–64. https://doi.org/10.1080/03639045.2018.1546316 [PubMed]
- 50. Yang J, Xu J, Danniel M, Wang X, Wang W, Zeng L, Shen L. The interaction between XBP1 and eNOS contributes to endothelial cell migration. Exp Cell Res. 2018; 363:262–70. https://doi.org/10.1016/j.yexcr.2018.01.016 [PubMed]
- 51. Zheng H, Zhao W, Yan C, Watson CC, Massengill M, Xie M, Massengill C, Noyes DR, Martinez GV, Afzal R, Chen Z, Ren X, Antonia SJ, et al. HDAC inhibitors enhance t-cell chemokine expression and augment response to PD-1 immunotherapy in lung adenocarcinoma. Clin Cancer Res. 2016; 22:4119–32. https://doi.org/10.1158/1078-0432.CCR-15-2584 [PubMed]
- 52. Hopewell EL, Zhao W, Fulp WJ, Bronk CC, Lopez AS, Massengill M, Antonia S, Celis E, Haura EB, Enkemann SA, Chen DT, Beg AA. Lung tumor NF-κB signaling promotes T cell-mediated immune surveillance. J Clin Invest. 2013; 123:2509–22. https://doi.org/10.1172/JCI67250 [PubMed]
- 53. Zhang L, Conejo-Garcia JR, Katsaros D, Gimotty PA, Massobrio M, Regnani G, Makrigiannakis A, Gray H, Schlienger K, Liebman MN, Rubin SC, Coukos G. Intratumoral T cells, recurrence, and survival in epithelial ovarian cancer. N Engl J Med. 2003; 348:203–13. https://doi.org/10.1056/NEJMoa020177 [PubMed]
- 54. Pagès F, Berger A, Camus M, Sanchez-Cabo F, Costes A, Molidor R, Mlecnik B, Kirilovsky A, Nilsson M, Damotte D, Meatchi T, Bruneval P, Cugnenc PH, et al. Effector memory T cells, early metastasis, and survival in colorectal cancer. N Engl J Med. 2005; 353:2654–66. https://doi.org/10.1056/NEJMoa051424 [PubMed]
- 55. Fan Y, Li S, Ding X, Yue J, Jiang J, Zhao H, Hao R, Qiu W, Liu K, Li Y, Wang S, Zheng L, Ye B, et al. First-in-class immune-modulating small molecule icaritin in advanced hepatocellular carcinoma: preliminary results of safety, durable survival and immune biomarkers. BMC Cancer. 2019; 19:279. https://doi.org/10.1186/s12885-019-5471-1 [PubMed]
- 56. Garcia-Diaz A, Shin DS, Moreno BH, Saco J, Escuin-Ordinas H, Rodriguez GA, Zaretsky JM, Sun L, Hugo W, Wang X, Parisi G, Saus CP, Torrejon DY, et al. Interferon Receptor Signaling Pathways Regulating PD-L1 and PD-L2 Expression. Cell Rep. 2017; 19:1189–201. https://doi.org/10.1016/j.celrep.2017.04.031 [PubMed]
- 57. Huang X, Zhu D, Lou Y. A novel anticancer agent, icaritin, induced cell growth inhibition, G1 arrest and mitochondrial transmembrane potential drop in human prostate carcinoma PC-3 cells. Eur J Pharmacol. 2007; 564:26–36. https://doi.org/10.1016/j.ejphar.2007.02.039 [PubMed]
- 58. Guo Y, Zhang X, Meng J, Wang ZY. An anticancer agent icaritin induces sustained activation of the extracellular signal-regulated kinase (ERK) pathway and inhibits growth of breast cancer cells. Eur J Pharmacol. 2011; 658:114–22. https://doi.org/10.1016/j.ejphar.2011.02.005 [PubMed]
- 59. Larionova I, Cherdyntseva N, Liu T, Patysheva M, Rakina M, Kzhyshkowska J. Interaction of tumor-associated macrophages and cancer chemotherapy. Oncoimmunology. 2019; 8:1596004. https://doi.org/10.1080/2162402X.2019.1596004 [PubMed]
- 60. Schumacher TN, Schreiber RD. Neoantigens in cancer immunotherapy. Science. 2015; 348:69–74. https://doi.org/10.1126/science.aaa4971 [PubMed]
- 61. Lasek W, Zagożdżon R, Jakobisiak M. Interleukin 12: still a promising candidate for tumor immunotherapy? Cancer Immunol Immunother. 2014; 63:419–35. https://doi.org/10.1007/s00262-014-1523-1 [PubMed]
- 62. Mellman I, Coukos G, Dranoff G. Cancer immunotherapy comes of age. Nature. 2011; 480:480–89. https://doi.org/10.1038/nature10673 [PubMed]
- 63. Song J, Zhong R, Huang H, Zhang Z, Ding D, Yan H, Sun E, Jia X. Combined treatment with epimedium koreanum nakai extract and gefitinib overcomes drug resistance caused by T790M mutation in non-small cell lung cancer cells. Nutr Cancer. 2014; 66:682–89. https://doi.org/10.1080/01635581.2014.895392 [PubMed]
- 64. Kang GY, Lee ER, Kim JH, Jung JW, Lim J, Kim SK, Cho SG, Kim KP. Downregulation of PLK-1 expression in kaempferol-induced apoptosis of MCF-7 cells. Eur J Pharmacol. 2009; 611:17–21. https://doi.org/10.1016/j.ejphar.2009.03.068 [PubMed]
- 65. Alvarez-Sala A, Attanzio A, Tesoriere L, Garcia-Llatas G, Barberá R, Cilla A. Apoptotic effect of a phytosterol-ingredient and its main phytosterol (β-sitosterol) in human cancer cell lines. Int J Food Sci Nutr. 2019; 70:323–34. https://doi.org/10.1080/09637486.2018.1511689 [PubMed]
- 66. Ru J, Li P, Wang J, Zhou W, Li B, Huang C, Li P, Guo Z, Tao W, Yang Y, Xu X, Li Y, Wang Y, Yang L. TCMSP: a database of systems pharmacology for drug discovery from herbal medicines. J Cheminform. 2014; 6:13. https://doi.org/10.1186/1758-2946-6-13 [PubMed]
- 67. Keiser MJ, Roth BL, Armbruster BN, Ernsberger P, Irwin JJ, Shoichet BK. Relating protein pharmacology by ligand chemistry. Nat Biotechnol. 2007; 25:197–206. https://doi.org/10.1038/nbt1284 [PubMed]
- 68. Zheng C, Guo Z, Huang C, Wu Z, Li Y, Chen X, Fu Y, Ru J, Ali Shar P, Wang Y, Wang Y. Large-scale direct targeting for drug repositioning and discovery. Sci Rep. 2015; 5:11970. https://doi.org/10.1038/srep11970 [PubMed]
- 69. Yu H, Chen J, Xu X, Li Y, Zhao H, Fang Y, Li X, Zhou W, Wang W, Wang Y. A systematic prediction of multiple drug-target interactions from chemical, genomic, and pharmacological data. PLoS One. 2012; 7:e37608. https://doi.org/10.1371/journal.pone.0037608 [PubMed]
- 70. Davis AP, King BL, Mockus S, Murphy CG, Saraceni-Richards C, Rosenstein M, Wiegers T, Mattingly CJ. The comparative toxicogenomics database: update 2011. Nucleic Acids Res. 2011; 39:D1067–72. https://doi.org/10.1093/nar/gkq813 [PubMed]
- 71. Zhu F, Shi Z, Qin C, Tao L, Liu X, Xu F, Zhang L, Song Y, Liu X, Zhang J, Han B, Zhang P, Chen Y. Therapeutic target database update 2012: a resource for facilitating target-oriented drug discovery. Nucleic Acids Res. 2012; 40:D1128–36. https://doi.org/10.1093/nar/gkr797 [PubMed]
- 72. Thorn CF, Klein TE, Altman RB. PharmGKB: the pharmacogenomics knowledge base. Methods Mol Biol. 2013; 1015:311–20. https://doi.org/10.1007/978-1-62703-435-7_20 [PubMed]
- 73. Merico D, Isserlin R, Stueker O, Emili A, Bader GD. Enrichment map: a network-based method for gene-set enrichment visualization and interpretation. PLoS One. 2010; 5:e13984. https://doi.org/10.1371/journal.pone.0013984 [PubMed]
- 74. Tomfohr J, Lu J, Kepler TB. Pathway level analysis of gene expression using singular value decomposition. BMC Bioinformatics. 2005; 6:225. https://doi.org/10.1186/1471-2105-6-225 [PubMed]
- 75. Liu J, Jiang M, Li Z, Zhang X, Li X, Hao Y, Su X, Zhu J, Zheng C, Xiao W, Wang Y. A Novel systems pharmacology method to investigate molecular mechanisms of Scutellaria barbata D. Don for non-small cell lung cancer. Front Pharmacol. 2018; 9:1473. https://doi.org/10.3389/fphar.2018.01473 [PubMed]
- 76. Das J, Verma D, Gustafsson M, Lerm M. Identification of DNA methylation patterns predisposing for an efficient response to BCG vaccination in healthy BCG-naïve subjects. Epigenetics. 2019; 14:589–601. https://doi.org/10.1080/15592294.2019.1603963 [PubMed]
- 77. Kanehisa M, Furumichi M, Tanabe M, Sato Y, Morishima K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 2017; 45:D353–61. https://doi.org/10.1093/nar/gkw1092 [PubMed]


